- Mef2
-
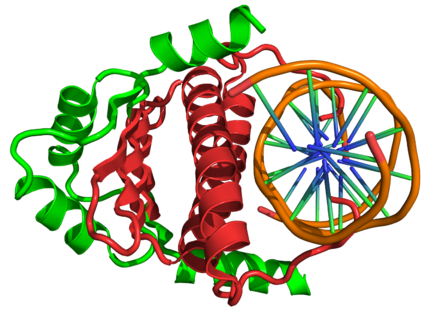 Dimeric structure of the MADS (red) and MEF2 (green) domains of the human MEF2B transcription factor complexed with DNA (orange) based on the PDB 1N6J crystallographic coordinates.
Dimeric structure of the MADS (red) and MEF2 (green) domains of the human MEF2B transcription factor complexed with DNA (orange) based on the PDB 1N6J crystallographic coordinates.
 The domain organization and sequence comparison of Mef2 proteins from representative species.[1] The amino acid numbering shown is of the human MEF2A sequence and the per cent sequence identities are all relative to hMEF2A. The three domain, MADS (red), MEF2 (green), and transactivation domains (TAD; cyan) are each highlighted in a different color.
The domain organization and sequence comparison of Mef2 proteins from representative species.[1] The amino acid numbering shown is of the human MEF2A sequence and the per cent sequence identities are all relative to hMEF2A. The three domain, MADS (red), MEF2 (green), and transactivation domains (TAD; cyan) are each highlighted in a different color.
In the field of molecular biology, myocyte enhancer factor-2 (Mef2) proteins are a family of transcription factors which through control of gene expression are important regulators of cellular differentiation and consequently play a critical role in embryonic development.[1] In adult organisms, Mef2 proteins mediate the stress response in some tissues.[1] Mef2 proteins contain both MADS-box and Mef2 DNA-binding domains.
Contents
Discovery
Mef2 was originally identified as a transcription factor complex through promoter analysis of the muscle creatine kinase (mck) gene to identify nuclear factors interacting with the mck enhancer region during muscle differentiation.[2]
Species distribution
The Mef2 gene is widely expressed in all branches of eukaryotes from yeast to humans. While drosophila has a single Mef2 gene, vertebrates have four versions of the Mef2 gene (human versions are denoted as MEF2A, MEF2B, MEF2C, and MEF2D), all expressed in distinct but overlapping patterns during embryogenesis through adulthood.[3]
Sequence and structure
All of the mammalian Mef2 genes share approximately 50% overall amino acid identity and about 95% similarity throughout the highly conserved N-terminal MADS-box and Mef2 domains, however their sequences diverge in their C-terminal transactivation domain (see figure to the right).[4]
The MADS-box serves as the minimal DNA-binding domain, however an adjacent 29-amino acid extension called the Mef2 domain is required for high affinity DNA-binding and dimerization. Through an interaction with the MADS-box, Mef2 transcription factors have the ability to homo- and heterodimerize,[5] and a classic nuclear localization sequence (NLS) in the C-terminus of Mef2A, -C, and – D ensures nuclear localization of the protein.[6] Interestingly, D-Mef2 and human MEF2B lack this conserved NLS but are still found in the nucleus.[7]
Function
Development
In drosophila, Mef2 regulates muscle development.[8] Vertebrate skeletal muscle differentiation in conjunction with bHLH transcription factors is also regulated by Mef2.[9][10]
Loss of Mef2c in neural crest cells results in results in craniofacial defects in the developing embryo and neonatal death caused by blocking of the upper airway passages.[11][12] Mef2c upregulates the expression of the homeodomain transcription factors DLX5 and DLX6, two transcription factors that are necessary for craniofacial development.[11][12]
Stress response
In adult tissues, Mef2 proteins regulate the stress-response during cardiac hypertrophy[13] and tissue remodeling in cardiac and skeletal muscle.[14]
References
- ^ a b c Potthoff MJ, Olson EN (December 2007). "MEF2: a central regulator of diverse developmental programs". Development 134 (23): 4131–40. doi:10.1242/dev.008367. PMID 17959722.
- ^ Lilly B, Galewsky S, Firulli AB, Schulz RA, Olson EN (June 1994). "D-MEF2: a MADS box transcription factor expressed in differentiating mesoderm and muscle cell lineages during Drosophila embryogenesis". Proc. Natl. Acad. Sci. U.S.A. 91 (12): 5662–6. doi:10.1073/pnas.91.12.5662. PMC 44056. PMID 8202544. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=44056.
- ^ McKinsey TA, Zhang CL, Olson EN (2002). "MEF2: a calcium-dependent regulator of cell division, differentiation and death". Trends Biochem Sci. 27 (1): 40–7. doi:10.1016/S0968-0004(01)02031-X. PMID 11796223.
- ^ Black BL, Olson EN (1998). "Transcriptional control of muscle development by myocyte enhancer factor-2 (MEF2) proteins". Annu Rev Cell Dev Biol 14: 167–96. doi:10.1146/annurev.cellbio.14.1.167. PMID 9891782.
- ^ Molkentin JD, Olson EN (1996). "Combinatorial control of muscle development by basic helix-loop-helix and MADS-box transcription factors". Proc Natl Acad Sci USA 93 (18): 9366–73. doi:10.1073/pnas.93.18.9366. PMC 38433. PMID 8790335. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=38433.
- ^ Borghi S, Molinari S, Razzini G, Parise F, Battini R, Ferrari S (2001). "The nuclear localization domain of the Mef2 family of transcription factors shows member-specific features and mediates the nuclear import of histone deacetylase 4". J Cell Sci 114 (Pt 24): 4477–83. PMID 11792813.
- ^ Yu YT (1996). "Distinct domains of myocyte enhancer binding factor-2A determining nuclear localization and cell type-specific transcriptional activity". J Biol Chem 271 (40): 24675–83. doi:10.1074/jbc.271.40.24675. PMID 8798735.
- ^ Gossett LA, Kelvin DJ, Sternberg EA, Olson EN (1 November 1989). "A new myocyte-specific enhancer-binding factor that recognizes a conserved element associated with multiple muscle-specific genes". Mol. Cell. Biol. 9 (11): 5022–33. PMC 363654. PMID 2601707. http://mcb.asm.org/cgi/pmidlookup?view=long&pmid=2601707.
- ^ Molkentin JD, Black BL, Martin JF, Olson EN (December 1995). "Cooperative activation of muscle gene expression by MEF2 and myogenic bHLH proteins". Cell 83 (7): 1125–36. doi:10.1016/0092-8674(95)90139-6. PMID 8548800.
- ^ Wang DZ, Valdez MR, McAnally J, Richardson J, Olson EN (November 2001). "The Mef2c gene is a direct transcriptional target of myogenic bHLH and MEF2 proteins during skeletal muscle development". Development 128 (22): 4623–33. PMID 11714687. http://dev.biologists.org/cgi/pmidlookup?view=long&pmid=11714687.
- ^ a b Verzi MP, Agarwal P, Brown C, McCulley DJ, Schwarz JJ, Black BL (April 2007). "The transcription factor MEF2C is required for craniofacial development". Dev. Cell 12 (4): 645–52. doi:10.1016/j.devcel.2007.03.007. PMC 1920108. PMID 17420000. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1920108.
- ^ a b Miller CT, Swartz ME, Khuu PA, Walker MB, Eberhart JK, Kimmel CB (August 2007). "mef2ca is required in cranial neural crest to effect Endothelin1 signaling in zebrafish". Dev. Biol. 308 (1): 144–57. doi:10.1016/j.ydbio.2007.05.018. PMC 2148033. PMID 17574232. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2148033.
- ^ Zhang CL, McKinsey TA, Chang S, Antos CL, Hill JA, Olson EN (August 2002). "Class II histone deacetylases act as signal-responsive repressors of cardiac hypertrophy". Cell 110 (4): 479–88. doi:10.1016/S0092-8674(02)00861-9. PMID 12202037. http://linkinghub.elsevier.com/retrieve/pii/S0092867402008619.
- ^ Potthoff MJ, Wu H, Arnold MA, Shelton JM, Backs J, McAnally J, Richardson JA, Bassel-Duby R, Olson EN (September 2007). "Histone deacetylase degradation andMEF2 activation promote the formation of slow-twitch myofibers". J. Clin. Invest. 117 (9): 2459–67. doi:10.1172/JCI31960. PMC 1957540. PMID 17786239. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1957540.
External links
- MeSH MEF2+protein,+C+elegans
- MeSH Mef2+protein,+Drosophila
- MeSH Mef2+protein,+zebrafish
- MeSH SMP1+protein,+Arabidopsis
- MeSH SMP1+protein,+S+cerevisiae
Transcription factors and intracellular receptors (1) Basic domains (1.1) Basic leucine zipper (bZIP)Activating transcription factor (AATF, 1, 2, 3, 4, 5, 6, 7) · AP-1 (c-Fos, FOSB, FOSL1, FOSL2, JDP2, c-Jun, JUNB, JUND) · BACH (1, 2) · BATF · BLZF1 · C/EBP (α, β, γ, δ, ε, ζ) · CREB (1, 3, L1) · CREM · DBP · DDIT3 · GABPA · HLF · MAF (B, F, G, K) · NFE (2, L1, L2, L3) · NFIL3 · NRL · NRF (1, 2, 3) · XBP1(1.2) Basic helix-loop-helix (bHLH)ATOH1 · AhR · AHRR · ARNT · ASCL1 · BHLHB2 · BMAL (ARNTL, ARNTL2) · CLOCK · EPAS1 · FIGLA · HAND (1, 2) · HES (5, 6) · HEY (1, 2, L) · HES1 · HIF (1A, 3A) · ID (1, 2, 3, 4) · LYL1 · MESP2 · MXD4 · MYCL1 · MYCN · Myogenic regulatory factors (MyoD, Myogenin, MYF5, MYF6) · Neurogenins (1, 2, 3) · NeuroD (1, 2) · NPAS (1, 2, 3) · OLIG (1, 2) · Pho4 · Scleraxis · SIM (1, 2) · TAL (1, 2) · Twist · USF1(1.3) bHLH-ZIP(1.4) NF-1(1.5) RF-X(1.6) Basic helix-span-helix (bHSH)(2) Zinc finger DNA-binding domains (2.1) Nuclear receptor (Cys4)subfamily 1 (Thyroid hormone (α, β), CAR, FXR, LXR (α, β), PPAR (α, β/δ, γ), PXR, RAR (α, β, γ), ROR (α, β, γ), Rev-ErbA (α, β), VDR)
subfamily 2 (COUP-TF (I, II), Ear-2, HNF4 (α, γ), PNR, RXR (α, β, γ), Testicular receptor (2, 4), TLX)
subfamily 3 (Steroid hormone (Androgen, Estrogen (α, β), Glucocorticoid, Mineralocorticoid, Progesterone), Estrogen related (α, β, γ))
subfamily 4 NUR (NGFIB, NOR1, NURR1) · subfamily 5 (LRH-1, SF1) · subfamily 6 (GCNF) · subfamily 0 (DAX1, SHP)(2.2) Other Cys4(2.3) Cys2His2General transcription factors (TFIIA, TFIIB, TFIID, TFIIE (1, 2), TFIIF (1, 2), TFIIH (1, 2, 4, 2I, 3A, 3C1, 3C2))
ATBF1 · BCL (6, 11A, 11B) · CTCF · E4F1 · EGR (1, 2, 3, 4) · ERV3 · GFI1 · GLI-Krüppel family (1, 2, 3, REST, S2, YY1) · HIC (1, 2) · HIVEP (1, 2, 3) · IKZF (1, 2, 3) · ILF (2, 3) · KLF (2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 17) · MTF1 · MYT1 · OSR1 · PRDM9 · SALL (1, 2, 3, 4) · SP (1, 2, 4, 7, 8) · TSHZ3 · WT1 · Zbtb7 (7A, 7B) · ZBTB (16, 17, 20, 32, 33, 40) · zinc finger (3, 7, 9, 10, 19, 22, 24, 33B, 34, 35, 41, 43, 44, 51, 74, 143, 146, 148, 165, 202, 217, 219, 238, 239, 259, 267, 268, 281, 295, 300, 318, 330, 346, 350, 365, 366, 384, 423, 451, 452, 471, 593, 638, 644, 649, 655)(2.4) Cys6(2.5) Alternating composition(3) Helix-turn-helix domains (3.1) HomeodomainARX · CDX (1, 2) · CRX · CUTL1 · DBX (1, 2) · DLX (3, 4, 5) · EMX2 · EN (1, 2) · FHL (1, 2, 3) · HESX1 · HHEX · HLX · Homeobox (A1, A2, A3, A4, A5, A7, A9, A10, A11, A13, B1, B2, B3, B4, B5, B6, B7, B8, B9, B13, C4, C5, C6, C8, C9, C10, C11, C13, D1, D3, D4, D8, D9, D10, D11, D12, D13) · HOPX · IRX (1, 2, 3, 4, 5, 6, MKX) · LMX (1A, 1B) · MEIS (1, 2) · MEOX2 · MNX1 · MSX (1, 2) · NANOG · NKX (2-1, 2-2, 2-3, 2-5, 3-1, 3-2, 6-1, 6-2) · NOBOX · PBX (1, 2, 3) · PHF (1, 3, 6, 8, 10, 16, 17, 20, 21A) · PHOX (2A, 2B) · PITX (1, 2, 3) · POU domain (PIT-1, BRN-3: A, B, C, Octamer transcription factor: 1, 2, 3/4, 6, 7, 11) · OTX (1, 2) · PDX1 · SATB2 · SHOX2 · VAX1 · ZEB (1, 2)(3.2) Paired box(3.3) Fork head / winged helix(3.4) Heat Shock Factors(3.5) Tryptophan clusters(3.6) TEA domain(4) β-Scaffold factors with minor groove contacts (4.1) Rel homology region(4.2) STAT(4.3) p53(4.4) MADS box(4.6) TATA binding proteins(4.7) High-mobility group(4.10) Cold-shock domainCSDA, YBX1(4.11) Runt(0) Other transcription factors (0.2) HMGI(Y)(0.3) Pocket domain(0.6) Miscellaneoussee also transcription factor/coregulator deficiencies
B bsyn: dna (repl, cycl, reco, repr) · tscr (fact, tcrg, nucl, rnat, rept, ptts) · tltn (risu, pttl, nexn) · dnab, rnab/runp · stru (domn, 1°, 2°, 3°, 4°)Categories:- Genes
- Transcription factors
Wikimedia Foundation. 2010.
