- TP63
-
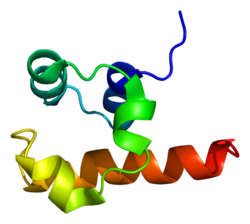
Tumor protein p63 also known as transformation-related protein 63 is a protein that in humans is encoded by the TP63 gene.[1][2][3][4]
TP63 also known as the p63 gene was discovered 20 years after the discovery of the p53 tumor suppressor gene and along with p73 constitutes the p53 gene family based on their structural similarity.[5] Despite being discovered significantly later than p53, phylogenetic analysis of p53, p63 and p73, suggest that p63 was the original member of the family from which p53 and p73 evolved.[6]
Contents
Function
Tumor protein p63 is a member of the p53 family of transcription factors. p63 -/- mice have several developmental defects which include the lack of limbs and other tissues, such as teeth and mammary glands, which develop as a result of interactions between mesenchyme and epithelium. TP63 encodes for two main isoforms by alternative promoters (TAp63 and ΔNp63). ΔNp63 is involved in multiple functions during skin development and in adult stem/progenitor cell regulation.[7] In contrast, TAp63 has been mostly restricted to its apoptotic function and more recently as the guardian of oocyte integrity.[8] Recently, two new functions have been attributed to TAp63 in heart development[9] and premature aging.[10]
Clinical significance
Mutations in the TP63 gene are associated with ectrodactyly-ectodermal dysplasia-cleft syndrome, Hay–Wells syndrome, cleft lip/palate syndrome 3 (EEC3); split-hand/foot malformation 4 (SHFM4); ankyloblepharon-ectodermal defects-cleft lip/palate; ADULT syndrome (acro-dermato-ungual-lacrimal-tooth); limb-mammary syndrome; Rap-Hodgkin syndrome (RHS); and orofacial cleft 8.
Diagnostic utility
p63 immunostaining has utility for differentiating prostatic adenocarcinoma (the most common type of prostate cancer) and benign prostatic tissue;[11] normal prostatic glands stain with p63 (as they have basal cells), while the malignant glands in prostatic adenocarcinoma (which lacks these cells) do not.[12] P63 is also helpful in distinguishing poorly differentiated squamous cell carcinoma from small cell carcinoma or adenocarcinoma. P63 should be strongly stained in poorly differentiated squamous cell, but negative in small cell or adenocarcinoma.[13]
Interactions
TP63 has been shown to interact with HNRPAB.[14]
Regulation
There is some evidence that the expression of p63 is regulated by the microRNA miR-203.[15][16]
See also
- AMACR - another marker for prostate adenocarcinoma
References
- ^ Yang A, Kaghad M, Wang Y, Gillett E, Fleming MD, Dötsch V, Andrews NC, Caput D, McKeon F (September 1998). "p63, a p53 homolog at 3q27-29, encodes multiple products with transactivating, death-inducing, and dominant-negative activities". Mol. Cell 2 (3): 305–16. doi:10.1016/S1097-2765(00)80275-0. PMID 9774969.
- ^ Osada M, Ohba M, Kawahara C, Ishioka C, Kanamaru R, Katoh I, Ikawa Y, Nimura Y, Nakagawara A, Obinata M, Ikawa S (July 1998). "Cloning and functional analysis of human p51, which structurally and functionally resembles p53". Nat. Med. 4 (7): 839–43. doi:10.1038/nm0798-839. PMID 9662378.
- ^ Zeng X, Zhu Y, Lu H (February 2001). "NBP is the p53 homolog p63". Carcinogenesis 22 (2): 215–9. doi:10.1093/carcin/22.2.215. PMID 11181441.
- ^ Tan M, Bian J, Guan K, Sun Y (February 2001). "p53CP is p51/p63, the third member of the p53 gene family: partial purification and characterization". Carcinogenesis 22 (2): 295–300. doi:10.1093/carcin/22.2.295. PMID 11181451.
- ^ Wu G, Nomoto S, Hoque MO, Dracheva T, Osada M, Lee CC, Dong SM, Guo Z, Benoit N, Cohen Y, Rechthand P, Califano J, Moon CS, Ratovitski E, Jen J, Sidransky D, Trink B (May 2003). "DeltaNp63alpha and TAp63alpha regulate transcription of genes with distinct biological functions in cancer and development". Cancer Res. 63 (10): 2351–7. PMID 12750249.
- ^ Skipper M (January 2007). "Dedicated protection for the female germline". Nature Reviews Molecular Cell Biology 8 (1): 4–5. doi:10.1038/nrm2091.
- ^ Crum CP, McKeon FD (2010). "p63 in epithelial survival, germ cell surveillance, and neoplasia". Annu Rev Pathol 5: 349–71. doi:10.1146/annurev-pathol-121808-102117. PMID 20078223.
- ^ Deutsch GB, Zielonka EM, Coutandin D, Weber TA, Schäfer B, Hannewald J, Luh LM, Durst FG, Ibrahim M, Hoffmann J, Niesen FH, Sentürk A, Kunkel H, Brutschy B, Schleiff E, Knapp S, Acker-Palmer A, Grez M, McKeon F, Dötsch V (February 2011). "DNA damage in oocytes induces a switch of the quality control factor TAp63α from dimer to tetramer". Cell 144 (4): 566–76. doi:10.1016/j.cell.2011.01.013. PMC 3087504. PMID 21335238. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3087504.
- ^ Rouleau M, Medawar A, Hamon L, Shivtiel S, Wolchinsky Z, Zhou H, De Rosa L, Candi E, de la Forest Divonne S, Mikkola ML, van Bokhoven H, Missero C, Melino G, Pucéat M, Aberdam D (September 2011). "TAp63 is Important for Cardiac Differentiation of Embryonic Stem Cells and Heart Development". Stem Cells 29: 1612–1683. doi:10.1002/stem.723. PMID 21898690. http://minus.com/lNWTY9g2q5yYG.
- ^ Su X, Paris M, Gi YJ, Tsai KY, Cho MS, Lin YL, Biernaskie JA, Sinha S, Prives C, Pevny LH, Miller FD, Flores ER (July 2009). "TAp63 prevents premature aging by promoting adult stem cell maintenance". Cell Stem Cell 5 (1): 64–75. doi:10.1016/j.stem.2009.04.003. PMID 19570515.
- ^ Shiran MS, Tan GC, Sabariah AR, Rampal L, Phang KS (March 2007). "p63 as a complementary basal cell specific marker to high molecular weight-cytokeratin in distinguishing prostatic carcinoma from benign prostatic lesions". Med. J. Malaysia 62 (1): 36–9. PMID 17682568.
- ^ Herawi M, Epstein JI (June 2007). "Immunohistochemical antibody cocktail staining (p63/HMWCK/AMACR) of ductal adenocarcinoma and Gleason pattern 4 cribriform and noncribriform acinar adenocarcinomas of the prostate". Am. J. Surg. Pathol. 31 (6): 889–94. doi:10.1097/01.pas.0000213447.16526.7f. PMID 17527076.
- ^ Zhang H, Liu J, Cagle PT, Allen TC, Laga AC, Zander DS (January 2005). "Distinction of pulmonary small cell carcinoma from poorly differentiated squamous cell carcinoma: an immunohistochemical approach". Mod. Pathol. 18 (1): 111–8. doi:10.1038/modpathol.3800251. PMID 15309021.
- ^ Fomenkov A, Huang YP, Topaloglu O, Brechman A, Osada M, Fomenkova T et al. (2003). "P63 alpha mutations lead to aberrant splicing of keratinocyte growth factor receptor in the Hay-Wells syndrome.". J Biol Chem 278 (26): 23906–14. doi:10.1074/jbc.M300746200. PMID 12692135.
- ^ Yi R, Poy MN, Stoffel M, Fuchs E (March 2008). "A skin microRNA promotes differentiation by repressing 'stemness'". Nature 452 (7184): 225–9. doi:10.1038/nature06642. PMID 18311128.
- ^ Aberdam D, Candi E, Knight RA, Melino G (December 2008). "miRNAs, 'stemness' and skin". Trends Biochem. Sci. 33 (12): 583–91. doi:10.1016/j.tibs.2008.09.002. PMID 18848452. http://min.us/lGvvAc29SY8ux.
Further reading
- Little NA, Jochemsen AG (2002). "p63.". Int. J. Biochem. Cell Biol. 34 (1): 6–9. doi:10.1016/S1357-2725(01)00086-3. PMID 11733180.
- van Bokhoven H, McKeon F (2002). "Mutations in the p53 homolog p63: allele-specific developmental syndromes in humans.". Trends in molecular medicine 8 (3): 133–9. doi:10.1016/S1471-4914(01)02260-2. PMID 11879774.
- van Bokhoven H, Brunner HG (2002). "Splitting p63.". Am. J. Hum. Genet. 71 (1): 1–13. doi:10.1086/341450. PMC 384966. PMID 12037717. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=384966.
- Brunner HG, Hamel BC, Bokhoven Hv H (2003). "P63 gene mutations and human developmental syndromes.". Am. J. Med. Genet. 112 (3): 284–90. doi:10.1002/ajmg.10778. PMID 12357472.
- Jacobs WB, Walsh GS, Miller FD (2005). "Neuronal survival and p73/p63/p53: a family affair.". The Neuroscientist : a review journal bringing neurobiology, neurology and psychiatry 10 (5): 443–55. doi:10.1177/1073858404263456. PMID 15359011.
- Zusman I (2005). "The soluble p51 protein in cancer diagnosis, prevention and therapy.". In Vivo 19 (3): 591–8. PMID 15875781.
- Morasso MI, Radoja N (2005). "Dlx genes, p63, and ectodermal dysplasias.". Birth Defects Res. C Embryo Today 75 (3): 163–71. doi:10.1002/bdrc.20047. PMC 1317295. PMID 16187309. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1317295.
- Barbieri CE, Pietenpol JA (2006). "p63 and epithelial biology.". Exp. Cell Res. 312 (6): 695–706. doi:10.1016/j.yexcr.2005.11.028. PMID 16406339.
- Shalom-Feuerstein R, Lena AM, Zhou H, De La Forest Divonne S, Van Bokhoven H, Candi E, Melino G, Aberdam D. ΔNp63 is an ectodermal gatekeeper of epidermal morphogenesis. Cell Death Differ. 2011 May;18(5):887-96. Link
External links
- MeSH TP73L+protein,+human
- GeneReviews/NCBI/NIH/UW entry on Ankyloblepharon-Ectodermal Defects-Cleft Lip/Palate Syndrome or AEC Syndrome, Hay-Wells Syndrome. Includes: Rapp-Hodgkin Syndrome
- OMIM entries on AEC
- http://insertech.simdif.com [3]
PDB gallery Neoplasm: Tumor suppressor genes/proteins and Oncogenes/Proto-oncogenes Ligand Receptor TSP: CDH1TSP: PTCH1TSP: TGF beta receptor 2Intracellular signaling P+Ps ONCO: Beta-catenin · TSP: APCHippo signaling pathwayOther/unknownNucleus TSP: VHL · ONCO: CBL - MDM2Mitochondria Other/ungrouped M: NEO
tsoc, mrkr
tumr, epon, para
drug (L1i/1e/V03)
Transcription factors and intracellular receptors (1) Basic domains (1.1) Basic leucine zipper (bZIP)Activating transcription factor (AATF, 1, 2, 3, 4, 5, 6, 7) · AP-1 (c-Fos, FOSB, FOSL1, FOSL2, JDP2, c-Jun, JUNB, JUND) · BACH (1, 2) · BATF · BLZF1 · C/EBP (α, β, γ, δ, ε, ζ) · CREB (1, 3, L1) · CREM · DBP · DDIT3 · GABPA · HLF · MAF (B, F, G, K) · NFE (2, L1, L2, L3) · NFIL3 · NRL · NRF (1, 2, 3) · XBP1(1.2) Basic helix-loop-helix (bHLH)ATOH1 · AhR · AHRR · ARNT · ASCL1 · BHLHB2 · BMAL (ARNTL, ARNTL2) · CLOCK · EPAS1 · FIGLA · HAND (1, 2) · HES (5, 6) · HEY (1, 2, L) · HES1 · HIF (1A, 3A) · ID (1, 2, 3, 4) · LYL1 · MESP2 · MXD4 · MYCL1 · MYCN · Myogenic regulatory factors (MyoD, Myogenin, MYF5, MYF6) · Neurogenins (1, 2, 3) · NeuroD (1, 2) · NPAS (1, 2, 3) · OLIG (1, 2) · Pho4 · Scleraxis · SIM (1, 2) · TAL (1, 2) · Twist · USF1(1.3) bHLH-ZIP(1.4) NF-1(1.5) RF-X(1.6) Basic helix-span-helix (bHSH)(2) Zinc finger DNA-binding domains (2.1) Nuclear receptor (Cys4)subfamily 1 (Thyroid hormone (α, β), CAR, FXR, LXR (α, β), PPAR (α, β/δ, γ), PXR, RAR (α, β, γ), ROR (α, β, γ), Rev-ErbA (α, β), VDR)
subfamily 2 (COUP-TF (I, II), Ear-2, HNF4 (α, γ), PNR, RXR (α, β, γ), Testicular receptor (2, 4), TLX)
subfamily 3 (Steroid hormone (Androgen, Estrogen (α, β), Glucocorticoid, Mineralocorticoid, Progesterone), Estrogen related (α, β, γ))
subfamily 4 NUR (NGFIB, NOR1, NURR1) · subfamily 5 (LRH-1, SF1) · subfamily 6 (GCNF) · subfamily 0 (DAX1, SHP)(2.2) Other Cys4(2.3) Cys2His2General transcription factors (TFIIA, TFIIB, TFIID, TFIIE (1, 2), TFIIF (1, 2), TFIIH (1, 2, 4, 2I, 3A, 3C1, 3C2))
ATBF1 · BCL (6, 11A, 11B) · CTCF · E4F1 · EGR (1, 2, 3, 4) · ERV3 · GFI1 · GLI-Krüppel family (1, 2, 3, REST, S2, YY1) · HIC (1, 2) · HIVEP (1, 2, 3) · IKZF (1, 2, 3) · ILF (2, 3) · KLF (2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 17) · MTF1 · MYT1 · OSR1 · PRDM9 · SALL (1, 2, 3, 4) · SP (1, 2, 4, 7, 8) · TSHZ3 · WT1 · Zbtb7 (7A, 7B) · ZBTB (16, 17, 20, 32, 33, 40) · zinc finger (3, 7, 9, 10, 19, 22, 24, 33B, 34, 35, 41, 43, 44, 51, 74, 143, 146, 148, 165, 202, 217, 219, 238, 239, 259, 267, 268, 281, 295, 300, 318, 330, 346, 350, 365, 366, 384, 423, 451, 452, 471, 593, 638, 644, 649, 655)(2.4) Cys6(2.5) Alternating composition(3) Helix-turn-helix domains (3.1) HomeodomainARX · CDX (1, 2) · CRX · CUTL1 · DBX (1, 2) · DLX (3, 4, 5) · EMX2 · EN (1, 2) · FHL (1, 2, 3) · HESX1 · HHEX · HLX · Homeobox (A1, A2, A3, A4, A5, A7, A9, A10, A11, A13, B1, B2, B3, B4, B5, B6, B7, B8, B9, B13, C4, C5, C6, C8, C9, C10, C11, C12, C13, D1, D3, D4, D8, D9, D10, D11, D12, D13) · HOPX · IRX (1, 2, 3, 4, 5, 6, MKX) · LMX (1A, 1B) · MEIS (1, 2) · MEOX2 · MNX1 · MSX (1, 2) · NANOG · NKX (2-1, 2-2, 2-3, 2-5, 3-1, 3-2, 6-1, 6-2) · NOBOX · PBX (1, 2, 3) · PHF (1, 3, 6, 8, 10, 16, 17, 20, 21A) · PHOX (2A, 2B) · PITX (1, 2, 3) · POU domain (PIT-1, BRN-3: A, B, C, Octamer transcription factor: 1, 2, 3/4, 6, 7, 11) · OTX (1, 2) · PDX1 · SATB2 · SHOX2 · VAX1 · ZEB (1, 2)(3.2) Paired box(3.3) Fork head / winged helix(3.4) Heat Shock Factors(3.5) Tryptophan clusters(3.6) TEA domain(4) β-Scaffold factors with minor groove contacts (4.1) Rel homology region(4.2) STAT(4.3) p53(4.4) MADS box(4.6) TATA binding proteins(4.7) High-mobility group(4.10) Cold-shock domainCSDA, YBX1(4.11) Runt(0) Other transcription factors (0.2) HMGI(Y)(0.3) Pocket domain(0.6) MiscellaneousCell cycle proteins Cyclin CDK CDK inhibitor P53 p63 p73 family Phases and
checkpointsOther cellular phasesCategories:- Human proteins
- Chromosome 3 gene stubs
Wikimedia Foundation. 2010.