- CYP3A4
-
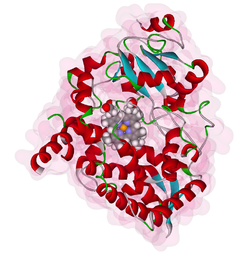
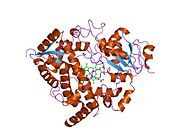
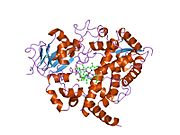
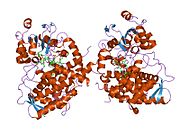
Cytochrome P450 3A4 (abbreviated CYP3A4) (EC 1.14.13.97), a member of the cytochrome P450 mixed-function oxidase system, is one of the most important enzymes involved in the metabolism of xenobiotics in the body. CYP3A4 is involved in the oxidation of the largest range of substrates of all the CYPs. As a result, CYP3A4 is present in the largest quantity of all the CYPs in the liver. In humans, the CYP3A4 protein is encoded by the CYP3A4 gene.[1] This gene is part of a cluster of cytochrome P450 genes on chromosome 7q21.1.[2]
Contents
Function
CYP3A4 is a member of the cytochrome P450 superfamily of enzymes. The cytochrome P450 proteins are monooxygenases that catalyze many reactions involved in drug metabolism and synthesis of cholesterol, steroids, and other lipids. This protein localizes to the endoplasmic reticulum, and its expression is induced by glucocorticoids and some pharmacological agents. This enzyme is involved in the metabolism of approximately half the drugs that are used today, including acetaminophen, codeine, ciclosporin, diazepam, and erythromycin. The enzyme also metabolizes some steroids and carcinogens.[3] Most drugs undergo deactivation by CYP3A4, either directly or by facilitated excretion from the body. Also, many substances are bioactivated by CYP3A4 to form their active compounds, and many protoxins being toxicated into their toxic forms (for examples - see table below).
Tissue distribution
Fetuses do not express CYP3A4 in their liver tissue, but rather CYP3A7 (EC 1.14.14.1), which acts on a similar range of substrates. CYP3A4 is absent in fetal liver but increases to approximately 40% of adult levels in the fourth month of life and 72% at 12 months.[4][5]
Although CYP3A4 is predominantly found in the liver, it is also present in other organs and tissues of the body, where it may play an important role in metabolism. CYP3A4 in the intestine plays an important role in the metabolism of certain drugs. Often this allows prodrugs to be activated and absorbed - as in the case of the histamine H1-receptor antagonist terfenadine.
Recently CYP3A4 has also been identified in the brain, however its role in the central nervous system is still unknown.[6]
Variability
While over 28 single nucleotide polymorphisms (SNPs) have been identified in the CYP3A4 gene, it has been found that this does not translate into significant interindividual variability in vivo. It can be supposed that this may be due to the induction of CYP3A4 on exposure to substrates.
Variability in CYP3A4 function can be determined noninvasively by the erythromycin breath test (ERMBT). The ERMBT estimates in vivo CYP3A4 activity by measuring the radiolabelled carbon dioxide exhaled after an intravenous dose of (14C-N-methyl)-erythromycin.[7]
Induction
CYP3A4 is induced by a wide variety of ligands. These ligands bind to the pregnane X receptor (PXR). The activated PXR complex forms a heterodimer with the retinoid X receptor (RXR), which binds to the XREM region of the CYP3A4 gene. XREM is a regulatory region of the CYP3A4 gene, and binding causes a cooperative interaction with proximal promoter regions of the gene, resulting in increased transcription and expression of CYP3A4.
Turnover
Estimates of the turnover rate of human CYP3A4 vary widely. For hepatic CYP3A4, in vivo methods yield estimates of enzyme half-life mainly in the range of 70 to 140 hours, whereas in vitro methods give estimates from 26 to 79 hours.[8] Turnover of gut CYP3A4 is likely to be a function of the rate of enterocyte renewal; an indirect approach based on recovery of activity following exposure to grapefruit juice yields measurements in the 12 to 33 hour range.[8]
CYP3A4 ligands
Following is a table of selected substrates, inducers and inhibitors of CYP3A4. Where classes of agents are listed, there may be exceptions within the class.
Inhibitors of CYP3A4 can be classified by their potency, such as:
- Strong inhibitor being one that causes at least a 5-fold increase in the plasma AUC values, or more than 80% decrease in clearance.[9]
- Moderate inhibitor being one that causes at least a 2-fold increase in the plasma AUC values, or 50-80% decrease in clearance.[9]
- Weak inhibitor being one that causes at least a 1.25-fold but less than 2-fold increase in the plasma AUC values, or 20-50% decrease in clearance.[9]
Selected inducers, inhibitors and substrates of CYP3A4[10] Substrates Inhibitors Inducers - some immunosuppressants
- many chemotherapeutic
- azole antifungals
- macrolides
- (not azithromycin)[9]
- dapsone[9] (in leprosy)
- tricyclic antidepressants
- SSRIs
- citalopram[11]
- norfluoxetine[11]
- sertraline[11]
- some other antidepressants
- buspirone[11][9] (anxiolytic)
- antipsychotics
- opiate (mainly analgesics)
- benzodiazepines
- some hypnotics
- donepezil[11] (acetylcholinesterase inhibitor)
- statins
- (not pravastatin)[9]
- (not rosuvastatin)[9]
- calcium channel blockers
- amiodarone[11] (class III antiarrhythmic)
- quinidine[9] (class I antiarrhythmic)
- PDE5 inhibitors
- kinins[11] (vasodilators, smooth muscle contractors)
- sex hormones agonists and antagonists
- finasteride[11][9] (antiandrogen)
- estradiol[9] (estrogen)
- progesterone[9]
- ethinylestradiol[11] (hormonal contraceptive)
- testosterone[9] (androgen)
- toremifene[11] (SERM)
- H1-receptor antagonists
- non-nucleoside reverse transcriptase inhibitors
- some glucocorticoids
- budesonide[11]
- hydrocortisone[9]
- dexamethasone[9]
- cisapride[11][9] (5-HT4 receptor agonist)
- aprepitant[9] (antiemetic)
- caffeine[9] (stimulant)
- cocaine[9] (stimulant)
- cilostazol[9] (phosphodiesterase inhibitor)
- dextromethorphan[9] (antitussive)
- domperidone[9] (antidopaminergic)
- eplerenone[9] (aldosterone antagonist)
- lidocaine[9] (local anesthetic, antiarrhythmic)
- ondansetron[9] (5-HT3 antagonist)
- propranolol[9] (beta blocker)
- salmeterol[9] (beta agonist)
- warfarin[15] (anticoagulant)
- clopidogrel, becoming bioactivated[16] (antiplatelet)
- esomeprazole[11] (proton pump inhibitor)
- nateglinide[9] (antidiabetic)
strong: - protease inhibitors
- some macrolide antibiotics[17]
- chloramphenicol (antibiotic)[18]
- some azole antifungals
- nefazodone[11][9] (antidepressant)
- amentoflavone[19] (in Ginkgo biloba and St. John’s Wort)
moderate
- aprepitant[9] (antiemetic)
- some calcium channel blockers
- some macrolide antibiotics
- some azole antifungals[17]
- bergamottin (constituent of grapefruit juice)[9]
weak:
- cimetidine[9] (H2-receptor antagonist)
- buprenorphine (analgesic)[20]
- cafestol (in unfiltered coffee)[21]
unspecified potency:
- amiodarone[9] (antiarrhythmic)
- ciprofloxacin[9] (antibiotic)
- dithiocarbamate[9] (functional group)
- voriconazole[9] (antifungal)
- imatinib[9] (anticancer)
- mifepristone[9] (abortifacient)
- norfloxacin[9] (antibiotic)
- some non-nucleoside reverse transcriptase inhibitors[22]
- gestodene[9] (hormonal contraceptive)
- mibefradil[9] (in angina pectoris)
- SSRIs
- fluoxetine/norfluoxetine[9]
- fluvoxamine[9]
- star fruit[9][23]
- milk thistle[24]
- ginko biloba[19][25]
- anticonvulsants, mood stabilizers
- barbiturates[17]
- St Johns Wort[11][9]
- some bactericidals
- some non-nucleoside reverse transcriptase inhibitors [22]
- some hypoglycemics
- glucocorticoids[9] (blood glucose increase, immunosuppressive)
- modafinil[9] (stimulant)
See also
References
- ^ Hashimoto H, Toide K, Kitamura R, Fujita M, Tagawa S, Itoh S, Kamataki T (December 1993). "Gene structure of CYP3A4, an adult-specific form of cytochrome P450 in human livers, and its transcriptional control". Eur. J. Biochem. 218 (2): 585–95. doi:10.1111/j.1432-1033.1993.tb18412.x. PMID 8269949.
- ^ Inoue K, Inazawa J, Nakagawa H, Shimada T, Yamazaki H, Guengerich FP, Abe T (June 1992). "Assignment of the human cytochrome P-450 nifedipine oxidase gene (CYP3A4) to chromosome 7 at band q22.1 by fluorescence in situ hybridization". Jpn. J. Hum. Genet. 37 (2): 133–8. doi:10.1007/BF01899734. PMID 1391968.
- ^ "Entrez Gene: cytochrome P450". http://www.ncbi.nlm.nih.gov/sites/entrez?Db=gene&Cmd=ShowDetailView&TermToSearch=1576.
- ^ Johnson TN, Rostami-Hodjegan A, Tucker GT (2006). "Prediction of the clearance of eleven drugs and associated variability in neonates, infants and children". Clin Pharmacokinet 45 (9): 931–56. doi:10.2165/00003088-200645090-00005. PMID 16928154.
- ^ Johnson TN, Tucker GT, Rostami-Hodjegan A (May 2008). "Development of CYP2D6 and CYP3A4 in the first year of life". Clin. Pharmacol. Ther. 83 (5): 670–1. doi:10.1038/sj.clpt.6100327. PMID 18043691.
- ^ Robertson G, Field J, Goodwin B, Bierach S, Tran M, Lehnert A, Liddle C (2003). "Transgenic mouse models of human CYP3A4 gene regulation". Mol Pharmacol 64 (1): 42–50. doi:10.1124/mol.64.1.42. PMID 12815159. http://molpharm.aspetjournals.org/cgi/content/full/64/1/42.
- ^ Watkins P (1994). "Noninvasive tests of CYP3A enzymes". Pharmacogenetics 4 (4): 171–84. doi:10.1097/00008571-199408000-00001. PMID 7987401.
- ^ a b Yang, J.; Liao, M.; Shou, M.; Jamei, M.; Yeo, K. R.; Tucker, G. T.; Rostami-Hodjegan, A. (2008-06-01). "Cytochrome P450 turnover: regulation of synthesis and degradation, methods for determining rates, and implications for the prediction of drug interactions". Current Drug Metabolism (Bentham) 9 (5): 384–393. doi:10.2174/138920008784746382. PMID 18537575. http://www.ingentaconnect.com/content/ben/cdm/2008/00000009/00000005/art00004. Retrieved 2010-12-20.
- ^ a b c d e f g h i j k l m n o p q r s t u v w x y z aa ab ac ad ae af ag ah ai aj ak al am an ao ap aq ar as at au av aw ax ay az ba bb bc bd be bf bg bh bi bj bk bl bm bn bo bp bq br bs bt bu bv bw bx by bz ca cb cc cd ce cf cg ch ci cj ck cl cm cn co cp cq cr cs ct cu cv cw cx cy cz da db dc dd de df dg dh di dj dk dl dm dn do dp dq Flockhart DA (2007). "Drug Interactions: Cytochrome P450 Drug Interaction Table". Indiana University School of Medicine. http://medicine.iupui.edu/flockhart/table.htm. Retrieved on December 25, 2008.
- ^ Where classes of agents are listed, there may be exceptions within the class
- ^ a b c d e f g h i j k l m n o p q r s t u v w x y z aa ab ac ad ae af ag ah ai aj ak al am an ao ap aq ar as at au av aw ax ay az ba bb bc bd be bf bg bh bi bj bk bl bm bn bo FASS (drug formulary): Swedish environmental classification of pharmaceuticals Facts for prescribers (Fakta för förskrivare). Retrieved July 2011
- ^ "Erlotinib". http://www.drugs.com/ppa/erlotinib.html. "Metabolized primarily by CYP3A4 and, to a lesser degree, by CYP1A2 and the extrahepatic isoform CYP1A1"
- ^ Druglib.com
- ^ Matsumoto S, Yamazoe Y (February 2001). "Involvement of multiple human cytochromes P450 in the liver microsomal metabolism of astemizole and a comparison with terfenadine". British journal of clinical pharmacology 51 (2): 133–42. PMC 2014443. PMID 11259984. http://www.blackwell-synergy.com/openurl?genre=article&sid=nlm:pubmed&issn=0306-5251&date=2001&volume=51&issue=2&spage=133.
- ^ Daly, A. K.; King, B. P. (2003). "Pharmacogenetics of oral anticoagulants". Pharmacogenetics 13 (5): 247–252. doi:10.1097/01.fpc.0000054071.64000.bd. PMID 12724615.
- ^ Lau, WC; Waskell, LA; Watkins, PB; Neer, CJ; Horowitz, K; Hopp, AS; Tait, AR; Carville, DG et al. (2003). "Atorvastatin reduces the ability of clopidogrel to inhibit platelet aggregation: a new drug-drug interaction". Circulation 107 (1): 32–7. PMID 12515739.
- ^ a b c d e f Rod Flower; Humphrey P. Rang; Maureen M. Dale; Ritter, James M. (2007). Rang & Dale's pharmacology. Edinburgh: Churchill Livingstone. ISBN 0-443-06911-5.
- ^ Park JY, Kim KA, Kim SL (November 2003). "Chloramphenicol Is a Potent Inhibitor of Cytochrome P450 Isoforms CYP2C19 and CYP3A4 in Human Liver Microsomes". Antimicrob. Agents Chemother. 47 (11): 3464–9. doi:10.1128/AAC.47.11.3464-3469.2003. PMC 253795. PMID 14576103. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=253795.
- ^ a b http://www.ncbi.nlm.nih.gov/pubmed/19883715
- ^ Zhang W, Ramamoorthy Y, Tyndale RF, Sellers EM (June 2003). "Interaction of buprenorphine and its metabolite norbuprenorphine with cytochromes P450 in vitro". Drug Metab. Dispos. 31 (6): 768–72. doi:10.1124/dmd.31.6.768. PMID 12756210.
- ^ http://www.aapsj.org/abstracts/AM_2009/AAPS2009-001235.PDF
- ^ a b Non-nucleoside reverse transcriptase inhibitors have been shown to both induce and inhibit CYP3A4.
- ^ Hidaka M, Fujita K, Ogikubo T, et al. (June 2004). "Potent inhibition by star fruit of human cytochrome P450 3A (CYP3A) activity". Drug Metab. Dispos. 32 (6): 581–3. doi:10.1124/dmd.32.6.581. PMID 15155547.
- ^ HCVadvocate.org
- ^ Ginko Biloba has been shown to contain the potent inhibitor amentoflavone.
External links
This article incorporates text from the United States National Library of Medicine, which is in the public domain.
PDB gallery Oxidoreductases: dioxygenases, including steroid hydroxylases (EC 1.14) 1.14.11: 2-oxoglutarate 1.14.13: NADH or NADPH Flavin-containing monooxygenase (FMO1, FMO2, FMO3, FMO4, FMO5) - Nitric oxide synthase (NOS1, NOS2, NOS3) - Cholesterol 7 alpha-hydroxylase - Methane monooxygenase - 3A4 - Lanosterol 14 alpha-demethylase1.14.14: reduced flavin or flavoprotein 1.14.15: reduced iron-sulfur protein 1.14.16: reduced pteridine (BH4 dependent) 1.14.17: reduced ascorbate 1.14.18-19: other 1.14.99 - miscellaneous Cytochromes, oxygenases: cytochrome P450 (EC 1.14) CYP1 CYP2 CYP3 (CYP3A) CYP4 CYP5-20 CYP21-51 Categories:- Human proteins
- Cytochrome P450
- EC 1.14.14
Wikimedia Foundation. 2010.