- Voltage-gated potassium channel
-
Ion channel (eukariotic) 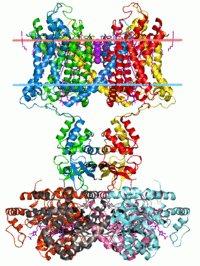
Potassium channel, structure in a membrane-like environment. Calculated hydrocarbon boundaries of the lipid bilayer are indicated by red and blue dots. Identifiers Symbol Ion_trans Pfam PF00520 InterPro IPR005821 SCOP 1bl8 TCDB 1.A.1 OPM family 8 OPM protein 2a79 Available protein structures: Pfam structures PDB RCSB PDB; PDBe PDBsum structure summary Ion channel (bacterial) 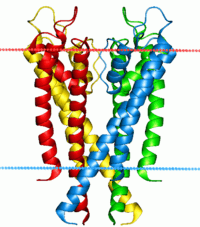
Potassium channel KcsA. Calculated hydrocarbon boundaries of the lipid bilayer are indicated by red and blue dots. Identifiers Symbol Ion_trans_2 Pfam PF07885 InterPro IPR013099 SCOP 1bl8 OPM protein 1r3j Available protein structures: Pfam structures PDB RCSB PDB; PDBe PDBsum structure summary Voltage-gated potassium channels are transmembrane channels specific for potassium and sensitive to voltage changes in the cell's membrane potential. During action potentials, they play a crucial role in returning the depolarized cell to a resting state.
Contents
Classification
Alpha subunits
Alpha subunits form the actual conductance pore. Based on sequence homology of the hydrophobic transmembrane cores, the alpha subunits of voltage-gated potassium channels are grouped into 12 classes, labeled Kvα1-12.[1] The following is a list of the 40 known human voltage-gated potassium channel alpha subunits grouped first according to function and then subgrouped according to the Kv sequence homology classification scheme:
Delayed rectifier
slowly inactivating or non-inactivating
- Kvα1.x - Shaker-related: Kv1.1 (KCNA1), Kv1.2 (KCNA2), Kv1.3 (KCNA3), Kv1.5 (KCNA5), Kv1.6 (KCNA6), Kv1.7 (KCNA7), Kv1.8 (KCNA10)
- Kvα2.x - Shab-related: Kv2.1 (KCNB1), Kv2.2 (KCNB2)
- Kvα3.x - Shaw-related: Kv3.1 (KCNC1), Kv3.2 (KCNC2)
- Kvα7.x: Kv7.1 (KCNQ1) - KvLQT1, Kv7.2 (KCNQ2), Kv7.3 (KCNQ3), Kv7.4 (KCNQ4), Kv7.5 (KCNQ5)
- Kvα10.x: Kv10.1 (KCNH1)
A-type potassium channel
rapidly inactivating
- Kvα1.x - Shaker-related: Kv1.4 (KCNA4)
- Kvα3.x - Shaw-related: Kv3.3 (KCNC3), Kv3.4 (KCNC4)
- Kvα4.x - Shal-related: Kv4.1 (KCND1), Kv4.2 (KCND2), Kv4.3 (KCND3)
Outward-rectifying
- Kvα10.x: Kv10.2 (KCNH5)
Inward-rectifying
Passes current more easily into the inwards direction (Into the cell).
Slowly activating
Modifier/silencer
Unable to form functional channels as homotetramers but instead heterotetramerize with Kvα2 family members to form conductive channels.
- Kvα5.x: Kv5.1 (KCNF1)
- Kvα6.x: Kv6.1 (KCNG1), Kv6.2 (KCNG2), Kv6.3 (KCNG3), Kv6.4 (KCNG4)
- Kvα8.x: Kv8.1 (KCNV1), Kv8.2 (KCNV2)
- Kvα9.x: Kv9.1 (KCNS1), Kv9.2 (KCNS2), Kv9.3 (KCNS3)
Beta subunits
Beta subunits are auxiliary proteins which associate with alpha subunits, sometimes in a α4β4 stoichiometry.[2] These subunits do not conduct current on their own but rather modulate the activity of Kv channels.[3]
- Kvβ1 (KCNAB1)
- Kvβ2 (KCNAB2)
- Kvβ3 (KCNAB3)
- minK[4] (KCNE1)
- MiRP1[5] (KCNE2)
- MiRP2 (KCNE3)
- MiRP3 (KCNE4)
- KCNE1-like (KCNE1L)
- KCNIP1 (KCNIP1)
- KCNIP2 (KCNIP2)
- KCNIP3 (KCNIP3)
- KCNIP4 (KCNIP4)
Proteins minK and MiRP1 are putative hERG beta subunits.[6]
Animal research
The voltage-gated K+ channels that provide the outward currents of action potentials have similarities to bacterial K+ channels.
These channels have been studied by X-ray diffraction, allowing determination of structural features at atomic resolution.
The function of these channels is explored by electrophysiological studies.
Genetic approaches include screening for behavioral changes in animals with mutations in K+ channel genes. Such genetic methods allowed the genetic identification of the "Shaker" K+ channel gene in Drosophila before ion channel gene sequences were well known.
Study of the altered properties of voltage-gated K+ channel proteins produced by mutated genes has helped reveal the functional roles of K+ channel protein domains and even individual amino acids within their structures.
Structure
Typically, vertebrate voltage-gated K+ channels are tetramers of four identical subunits arranged as a ring, each contributing to the wall of the trans-membrane K+ pore. Each subunit is composed of six membrane spanning hydrophobic α-helical sequences. The high resolution crystallographic structure of the rat Kvα1.2/β2 channel has recently been solved (Protein Databank Accession Number 2A79),[7] and then refined in a lipid membrane-like environment (PDB 2r9r).
Selectivity
Main article: Potassium_channel#Selectivity_filterVoltage-gated K+ channels are selective for K+ over other cations such as Na+. There is a selectivity filter at the narrowest part of the transmembrane pore.
Channel mutation studies have revealed the parts of the subunits that are essential for ion selectivity. They include the amino acid sequence (Thr-Val-Gly-Tyr-Gly) or (Thr-Val-Gly-Phe-Gly) typical to the selectivity filter of voltage-gated K+ channels. As K+ passes through the pore, interactions between potassium ions and water molecules are prevented and the K+ interacts with specific atomic components of the Thr-Val-Gly-X-Gly sequences from the four channel subunits[1].
It seems illogical at first that a channel should be able to allow potassium ions but not the smaller sodium ions through. However in an aqueous environment, potassium and sodium cations are solvated by water molecules. When moving through the selectivity filter of the potassium channel, these solvated water molecules are replaced by backbone carbonyl groups of the channel protein. The diameter of the selectivity filter is ideal for the potassium cation, but too big for the smaller sodium cation. Hence the potassium cations are well "solvated" by the protein carbonyl groups, but these same carbonyl groups are too far apart to adequately solvate the sodium cation. Hence the passage of potassium cations through this selectivity filter is strongly favored over sodium cations.
Open and closed conformations
Attempts continue to relate the structure of the mammalian voltage-gated K+ channel to its ability to respond to the voltage that exists across the membrane.[8] Specific domains of the channel subunits have been identified that are important for voltage-sensing and converting between the open conformation of the channel and closed conformations. There are at least two closed conformations; in one, the channel can open if the membrane potential becomes positive inside. Voltage-gated K+ channels inactivate after opening, entering a distinctive, second closed conformation. In the inactivated conformation, the channel cannot open, even if the transmembrane voltage is favorable. The amino terminal domain of the K+ channel or an auxiliary protein can mediate "N-type" inactivation. The former has been described as a "ball and chain" model where the N-terminus of the protein forms a ball which is tethered to the rest of the protein through a loop (the chain). The tethered ball is transiently sucked into the inner porehole, preventing ion movement through the channel.[9][10]
See also
References
- ^ Gutman GA, Chandy KG, Grissmer S, Lazdunski M, McKinnon D, Pardo LA, Robertson GA, Rudy B, Sanguinetti MC, Stuhmer W, Wang X (2005). "International Union of Pharmacology. LIII. Nomenclature and molecular relationships of voltage-gated potassium channels.". Pharmacol Rev 57 (4): 473–508. doi:10.1124/pr.57.4.10. PMID 16382104.
- ^ Pongs O, Leicher T, Berger M, Roeper J, Bahring R, Wray D, Giese KP, Silva AJ, Storm JF (1999). "Functional and molecular aspects of voltage-gated K+ channel beta subunits". Ann N Y Acad Sci 868 (Apr 30): 344–55. doi:10.1111/j.1749-6632.1999.tb11296.x. PMID 10414304.
- ^ Li Y, Um SY, McDonald TV (2006). "Voltage-gated potassium channels: regulation by accessory subunits". Neuroscientist 12 (3): 199–210. doi:10.1177/1073858406287717. PMID 16684966.
- ^ Zhang M, Jiang M, Tseng GN (2001). "minK-related peptide 1 associates with Kv4.2 and modulates its gating function: potential role as beta subunit of cardiac transient outward channel?". Circ Res 88 (10): 1012–9. doi:10.1161/hh1001.090839. PMID 11375270.
- ^ McCrossan ZA, Abbott GW (2004). "The MinK-related peptides". Neuropharmacology 47 (6): 787–821. doi:10.1016/j.neuropharm.2004.06.018. PMID 15527815.
- ^ Anantharam A, Abbott GW (2005). "Does hERG coassemble with a beta subunit? Evidence for roles of MinK and MiRP1". Novartis Found Symp 266 (42): 112–7, 155–8. doi:10.1002/047002142X.fmatter. PMID 16050264.
- ^ Long SB, Campbell EB, Mackinnon R (2005). "Crystal structure of a mammalian voltage-dependent Shaker family K+ channel". Science 309 (5736): 897–903. doi:10.1126/science.1116269. PMID 16002581.
- ^ Lee S, Lee A, Chen J, MacKinnon R (2005). "Structure of the KvAP voltage-dependent K+ channel and its dependence on the lipid membrane.". Proc Natl Acad Sci USA 102 (43): 15441–6. doi:10.1073/pnas.0507651102. PMC 1253646. PMID 16223877. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1253646.
- ^ Armstrong CM, Bezanilla F (April 1973). "Currents related to movement of the gating particles of the sodium channels". Nature 242 (5398): 459–61. doi:10.1038/242459a0. PMID 4700900.
- ^ Murrell-Lagnado RD, Aldrich RW (December 1993). "Energetics of Shaker K channels block by inactivation peptides". J. Gen. Physiol. 102 (6): 977–1003. doi:10.1085/jgp.102.6.977. PMC 2229186. PMID 8133246. http://www.jgp.org/cgi/pmidlookup?view=long&pmid=8133246.
External links
- MeSH Voltage-Gated+Potassium+Channels
- "Voltage-Gated Potassium Channels". IUPHAR Database of Receptors and Ion Channels. International Union of Basic and Clinical Pharmacology. http://www.iuphar-db.org/IC/FamilyMenuForward?familyId=16.
- Li B, Gallin W (2004). "VKCDB: voltage-gated potassium channel database.". BMC Bioinformatics 5: 3. doi:10.1186/1471-2105-5-3. PMC 317694. PMID 14715090. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=317694.
- "Voltage-gated potassium channel database (VKCDB)" at ualberta.ca
- UMich Orientation of Proteins in Membranes families/superfamily-8 - Spatial positions of voltage gated potassium channels in membranes
Ca2+: Calcium channel Ligand-gatedNa+: Sodium channel Constitutively activeProton gatedK+: Potassium channel Voltage-gatedKvα1-6 (1.1, 1.2, 1.3, 1.4, 1.5, 1.6, 1.7, 1.8) · (2.1, 2.2) · (3.1, 3.2, 3.3, 3.4) · (4.1, 4.2, 4.3) · (5.1) · (6.1, 6.2, 6.3, 6.4)
Kvα7-12 (7.1, 7.2, 7.3, 7.4, 7.5) · (8.1, 8.2) · (9.1, 9.2, 9.3) · (10.1, 10.2) · (11.1/hERG, 11.2, 11.3) · (12.1, 12.2, 12.3)
Kvβ (1, 2, 3) · KCNIP (1, 2, 3, 4) · minK/ISK · minK/ISK-like · MiRP (1, 2, 3) · Shaker geneOther Cl-: Chloride channelHVCN1Generalsee also disorders
B memb: cead, trns (1A, 1C, 1F, 2A, 3A1, 3A2-3, 3D), othrCategories:- Ion channels
- Electrophysiology
Wikimedia Foundation. 2010.
