- Actin
-
Actin Identifiers Symbol Actin Pfam PF00022 Pfam clan CL0108 InterPro IPR004000 PROSITE PDOC00340 SCOP 2btf Available protein structures: Pfam structures PDB RCSB PDB; PDBe PDBsum structure summary Actin is a globular, roughly 42-kDa moonlighting protein found in all eukaryotic cells (the only known exception being nematode sperm) where it may be present at concentrations of over 100 μM. It is also one of the most highly-conserved proteins, differing by no more than 20% in species as diverse as algae and humans. Actin is the monomeric subunit of two types of filaments in cells: microfilaments, one of the three major components of the cytoskeleton, and thin filaments, part of the contractile apparatus in muscle cells. Thus, actin participates in many important cellular processes including muscle contraction, cell motility, cell division and cytokinesis, vesicle and organelle movement, cell signaling, and the establishment and maintenance of cell junctions and cell shape. Many of these processes are mediated by extensive and intimate interactions of actin with cellular membranes.[1] In vertebrates, three main groups of actin isoforms, alpha, beta, and gamma have been identified. The alpha actins are found in muscle tissues and are a major constituent of the contractile apparatus. The beta and gamma actins co-exist in most cell types as components of the cytoskeleton, and as mediators of internal cell motility.
Contents
Formation of thin filament
Genetics
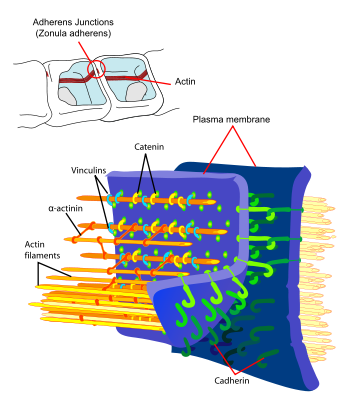
Principal interactions of structural proteins are at cadherin-based adherens junction. Actin filaments are linked to α-actinin and to the membrane through vinculin. The head domain of vinculin associates to E-cadherin via α-, β-, and γ-catenins. The tail domain of vinculin binds to membrane lipids and to actin filaments.
The protein actin is one of the most highly conserved throughout evolution because it interacts with a large number of other proteins, with 80.2% sequence conservation at the gene level between Homo sapiens and Saccharomyces cerevisiae (a species of yeast), and 95% conservation of the primary structure of the protein product.
Although most yeasts have only a single actin gene, higher eukaryotes, in general, express several isoforms of actin encoded by a family of related genes. Mammals have at least six actin isoforms coded by separate genes,[2] which are divided into three classes (alpha, beta and gamma) according to their isoelectric point. In general, alpha actins are found in muscle (α-skeletal, α-aortic smooth, α-cardiac, and γ2-enteric smooth), whereas beta and gamma isoforms are prominent in non-muscle cells (β- and γ1-cytoplasmic). Although the amino acid sequences and in vitro properties of the isoforms are highly similar, these isoforms cannot completely substitute for one another in vivo.[3]
The typical actin gene has an approximately 100-nucleotide 5' UTR, a 1200-nucleotide translated region, and a 200-nucleotide 3' UTR. The majority of actin genes are interrupted by introns, with up to 6 introns in any of 19 well-characterised locations. The high conservation of the family makes actin the favoured model for studies comparing the introns-early and introns-late models of intron evolution.
All non-spherical prokaryotes appear to possess genes such as MreB, which encode homologues of actin; these genes are required for the cell's shape to be maintained. The plasmid-derived gene ParM encodes an actin-like protein whose polymerised form is dynamically unstable, and appears to partition the plasmid DNA into the daughter cells during cell division by a mechanism analogous to that employed by microtubules in eukaryotic mitosis.[4] Actin is found in both smooth and rough endoplasmic reticulums.
Functions
Actin forms microfilaments which are typically one of the most dynamic of the three subclasses of the eukaryotic cytoskeleton.
In turn this gives actin major functions in cells:
- To form microfilaments to give mechanical support to cells, and provide trafficking routes through the cytoplasm to support signal transduction.
- To allow cell motility in cells which undergo amoeboid motion using pseudopods (see actoclampin molecular motors) and phagocytosis, for example of bacteria by macrophages.
- In metazoan muscle cells, to be the scaffold on which myosin proteins generate force to support muscle contraction.
- In non-muscle cells, to be a track for cargo transport myosins (non-conventional myosins) such as myosin V and VI. Non-conventional myosins use ATP hydrolysis to transport cargo, such as vesicles and organelles, in a directed fashion much faster than diffusion. Myosin V walks towards the barbed end of actin filaments, while myosin VI walks toward the pointed end. Most actin filaments are arranged with the barbed end toward the cellular membrane and the pointed end toward the cellular interior. This arrangement allows myosin V to be an effective motor for export of cargos, and myosin VI to be an effective motor for import.
Directionality
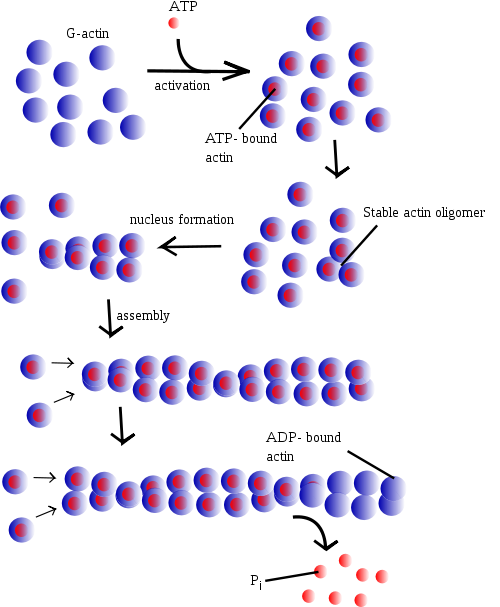
The polarity of an actin filament can be determined by decorating the microfilament with myosin "S1" fragments, creating barbed (+) and pointed (-) ends on the filament. An S1 fragment is composed of the head and neck domains of myosin II. Under physiologic conditions, G-actin (the monomer form) is transformed to F-actin (the polymer form) by ATP, where the role of ATP is essential.
Nucleation and polymerization
Actin polymerization and depolymerization is necessary in chemotaxis and cytokinesis. Nucleating factors are necessary to stimulate actin polymerization. One such nucleating factor is the ARP complex which acts as a barbed end of actin in its shape to stimulate the nucleation of G-actin (or monomeric actin). The Arp2/3 complex can also bind to actin filaments at 70 degrees to form new actin branches off of existing actin filaments. Also, actin filaments themselves bind ATP, and hydrolysis of this ATP stimulates destabilization of the polymer.
The growth of actin filaments can be regulated by thymosin and profilin. Thymosin binds to G-actin to buffer the polymerizing process while profilin binds to G-actin to exchange ADP for ATP promoting the monomeric addition to the barbed, plus end.
Microfilaments
Individual subunits of microfilaments are known as globular actin (G-actin). G-actin subunits assemble into long filamentous polymers called F-actin. Two parallel F-actin strands must rotate 166 degrees to layer correctly on top of each other. This creates the double helix structure of the microfilaments of the cytoskeleton. Microfilaments measure approximately 7 nm in diameter with a loop of the helix repeating every 37 nm.
Actomyosin filaments
In muscle, actin is the major component of thin filaments, which, together with the motor protein myosin (which forms thick filaments), are arranged into actomyosin myofibrils. These fibrils comprise the mechanism of muscle contraction. Using the hydrolysis of ATP for energy, myosin heads undergo a cycle during which they attach to thin filaments, exerting a tension, and then depending on the load, perform a power stroke that causes the thin filaments to slide past, shortening the muscle.
In contractile bundles, the actin-bundling protein alpha-actinin separates each thin filament by ~35 nm. This increase in distance allows thick filaments to fit in between and interact, enabling deformation or contraction. In deformation, one end of myosin is bound to the plasma membrane while the other end "walks" toward the plus end of the actin filament. This pulls the membrane into a different shape relative to the cell cortex. For contraction, the myosin molecule is usually bound to two separate filaments and both ends simultaneously "walk" toward their filament's plus end, sliding the actin filaments closer to each other. This results in the shortening, or contraction, of the actin bundle (but not the filament). This mechanism is responsible for muscle contraction and cytokinesis, the division of one cell into two.
Nuclear actin
Actin is essential for transcription from RNA polymerases I, II and III. In Pol I transcription, actin and myosin (MYO1C, which binds DNA) act as a molecular motor. For Pol II transcription, β-actin is needed for the formation of the pre-initiation complex. Pol III contains β-actin as a subunit. Actin can also be a component of chromatin remodeling complexes as well as pre-mRNP particles (that is, precursor messenger RNA bundled in proteins), and is involved in nuclear export of RNAs and proteins.[5]
History
Actin was first observed experimentally in 1887 by W.D. Halliburton, who extracted a protein from muscle that 'coagulated' preparations of myosin, which he dubbed "myosin-ferment."[6] However, Halliburton was unable to further characterise his findings, and the discovery of actin is credited instead to Brúnó F. Straub, a young biochemist working in Albert Szent-Györgyi's laboratory at the Institute of Medical Chemistry at the University of Szeged, Hungary.
In 1942, Straub developed a novel technique for extracting muscle protein that allowed him to isolate substantial amounts of relatively pure actin. Straub's method is essentially the same as that used in laboratories today. Szent-Gyorgyi had previously described the more viscous form of myosin produced by slow muscle extractions as 'activated' myosin, and, since Straub's protein produced the activating effect, it was dubbed actin. The hostilities of World War II meant that Szent-Gyorgyi and Straub were unable to publish the work in Western scientific journals; it became well known in the West only in 1945, when it was published as a supplement to the Acta Physiologica Scandinavica.[7]
Straub continued to work on actin, and in 1950 reported that actin contains bound ATP [8] and that, during polymerisation of the protein into microfilaments, the nucleotide is hydrolysed to ADP and inorganic phosphate (which remain bound in the microfilament). Straub suggested that the transformation of ATP-bound actin to ADP-bound actin played a role in muscular contraction. In fact, this is true only in smooth muscle, and was not supported through experimentation until 2001.[9]
The crystal structure of G-actin was solved in 1990 by Kabsch and colleagues.[10] In the same year, a model for F-actin was proposed by Holmes and colleagues.[11] The model was derived by fitting a helix of G-actin structures according to low-resolution fiber diffraction data from the filament. Several models of the filament have been proposed since. However, there is still no high-resolution X-ray structure of F-actin.
The Listeria bacteria use the cellular machinery to move around inside the host cell, by inducing directed polymerisation of actin by the ActA transmembrane protein, thus pushing the bacterial cell around.
See also
- MreB — one of the actin homologues in bacteria
- Motor protein — converts chemical energy into mechanical work
- ACTA1 — alpha actin 1
- ACTB — beta actin
- ACTG1 — gamma actin 1
- Actin remodeling — affect on cell structure and shape
- Active matter
References
- ^ Doherty GJ and McMahon HT (2008). "Mediation, Modulation and Consequences of Membrane-Cytoskeleton Interactions". Annual Review of Biophysics 37: 65–95. doi:10.1146/annurev.biophys.37.032807.125912. PMID 18573073. http://arjournals.annualreviews.org/doi/abs/10.1146/annurev.biophys.37.032807.125912.
- ^ Vandekerckhove J, Weber K (December 1978). "At least six different actins are expressed in a higher mammal: an analysis based on the amino acid sequence of the amino-terminal tryptic peptide". J. Mol. Biol. 126 (4): 783–802. doi:10.1016/0022-2836(78)90020-7. PMID 745245. http://linkinghub.elsevier.com/retrieve/pii/0022-2836(78)90020-7.
- ^ Khaitlina SY (2001). "Functional specificity of actin isoforms". Int. Rev. Cytol. 202: 35–98. doi:10.1016/S0074-7696(01)02003-4. PMID 11061563.
- ^ Garner EC, Campbell CS, Weibel DB, Mullins RD (March 2007). "Reconstitution of DNA segregation driven by assembly of a prokaryotic actin homolog". Science 315 (5816): 1270–4. doi:10.1126/science.1138527. PMC 2851738. PMID 17332412. http://www.sciencemag.org/cgi/pmidlookup?view=short&pmid=17332412.
- ^ Zheng B, Han M, Bernier M, Wen JK (May 2009). "Nuclear actin and actin-binding proteins in the regulation of transcription and gene expression". FEBS J. 276 (10): 2669–85. doi:10.1111/j.1742-4658.2009.06986.x. PMC 2978034. PMID 19459931. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2978034.
- ^ Halliburton, W.D. (1887). "On muscle plasma". J. Physiol. 8 (3-4): 133. PMC 1485127. PMID 16991477. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1485127.
- ^ Szent-Gyorgyi, A. (1945). "Studies on muscle". Acta Physiol Scandinav 9 (Suppl): 25.
- ^ Straub FB, Feuer G (1989). "Adenosinetriphosphate. The functional group of actin. 1950". Biochim. Biophys. Acta 1000: 180–95. PMID 2673365.
- ^ Bárány M, Barron JT, Gu L, Bárány K (December 2001). "Exchange of the actin-bound nucleotide in intact arterial smooth muscle". J. Biol. Chem. 276 (51): 48398–403. doi:10.1074/jbc.M106227200. PMID 11602582. http://www.jbc.org/cgi/pmidlookup?view=long&pmid=11602582.
- ^ Kabsch W, Mannherz HG, Suck D, Pai EF, Holmes KC (September 1990). "Atomic structure of the actin:DNase I complex". Nature 347 (6288): 37–44. doi:10.1038/347037a0. PMID 2395459.
- ^ Holmes KC, Popp D, Gebhard W, Kabsch W (September 1990). "Atomic model of the actin filament". Nature 347 (6288): 44–9. doi:10.1038/347044a0. PMID 2395461.
Proteins of the cytoskeleton Human ActinsI (MYO1A, MYO1B, MYO1C, MYO1D, MYO1E, MYO1F, MYO1G, MYO1H) · II (MYH1, MYH2, MYH3, MYH4, MYH6, MYH7, MYH7B, MYH8, MYH9, MYH10, MYH11, MYH13, MYH14, MYH15, MYH16) · III (MYO3A, MYO3B) · V (MYO5A, MYO5B, MYO5C) · VI (MYO6) · VII (MYO7A, MYO7B) · IX (MYO9A, MYO9B) · X (MYO10) · XV (MYO15A) · XVIII (MYO18A, MYO18B) · LC (MYL1, MYL2, MYL3, MYL4, MYL5, MYL6, MYL6B, MYL7, MYL9, MYLIP, MYLK, MYLK2, MYLL1)OtherOtherEpithelial keratins
(soft alpha-keratins)Hair keratins
(hard alpha-keratins)Ungrouped alphaNot alphaType 3Type 4Type 5OtherOtherNonhuman Histology: muscle tissue (TH H2.00.05, H3.3) Smooth
muscleStriated
muscleCostamere/
DAPCMembrane/
extracellularIntracellularDystrophin · Dystrobrevin (A, B) · Syntrophin (A, B1, B2, G1, G2) · Syncoilin · Dysbindin · Synemin/desmuslin
related: NOS1 · Caveolin 3Myofilament (thin filament/actin, thick filament/myosin, elastic filament/titin, nebulin)
Troponin (T, C, I)GeneralNeuromuscular junction · Motor unit · Muscle spindle · Excitation-contraction coupling · Sliding filament mechanismBothFiberCellsOtherOther/
ungroupedAntigens: Autoantigens Dehydrogenase Transglutaminase Nucleoporins Other Acetylcholine receptor · Actin · Apolipoprotein H · Cardiolipin · Centromere · Filaggrin(Citrullinate) · Gangliosides · Sp100 nuclear antigen · Thrombin · TopoisomeraseCategories:- Cytoskeleton
- Structural proteins
- Autoantigens
Wikimedia Foundation. 2010.