- Coenzyme Q
-
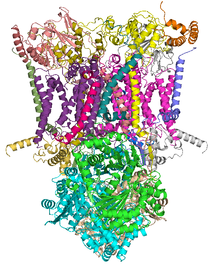
ubiquinol—cytochrome-c reductase Crystal structure of mitochondrial cytochrome bc1 complex bound with ubiquinone.[1] Identifiers EC number 1.10.2.2 CAS number 9027-03-6 Databases IntEnz IntEnz view BRENDA BRENDA entry ExPASy NiceZyme view KEGG KEGG entry MetaCyc metabolic pathway PRIAM profile PDB structures RCSB PDB PDBe PDBsum Gene Ontology AmiGO / EGO Search PMC articles PubMed articles In enzymology, a ubiquinol—cytochrome-c reductase (EC 1.10.2.2) is an enzyme that catalyzes the chemical reaction
- QH2 + 2 ferricytochrome c
 Q + 2 ferrocytochrome c + 2 H+
Q + 2 ferrocytochrome c + 2 H+
Thus, the two substrates of this enzyme are dihydroquinone (QH2) and ferri- (Fe3+) cytochrome c, whereas its 3 products are quinone (Q), ferro- (Fe2+) cytochrome c, and H+.
This enzyme belongs to the family of oxidoreductases, specifically those acting on diphenols and related substances as donor with a cytochrome as acceptor. This enzyme participates in oxidative phosphorylation. It has four cofactors: cytochrome C1, cytochrome b-562, cytochrome b-566 and a 2-Iron ferredoxin.
The coenzyme Q : cytochrome c — oxidoreductase, sometimes called the cytochrome bc1 complex, and at other times complex III, is the third complex in the electron transport chain (EC 1.10.2.2), playing a critical role in biochemical generation of ATP (oxidative phosphorylation). Complex III is a multisubunit transmembrane lipoprotein encoded by both the mitochondrial (cytochrome b) and the nuclear genomes (all other subunits). Complex III is present in the mitochondria of all animals and all aerobic eukaryotes and the inner membranes of most eubacteria. Mutations in Complex III cause exercise intolerance as well as multisystem disorders.
Contents
Nomenclature
The systematic name of this enzyme class is ubiquinol:ferricytochrome-c oxidoreductase. Other names in common use include:
- coenzyme Q-cytochrome c reductase,
- dihydrocoenzyme Q-cytochrome c reductase,
- reduced ubiquinone-cytochrome c reductase, complex III,
- (mitochondrial electron transport),
- ubiquinone-cytochrome c reductase,
- ubiquinol-cytochrome c oxidoreductase,
- reduced coenzyme Q-cytochrome c reductase,
- ubiquinone-cytochrome c oxidoreductase,
- reduced ubiquinone-cytochrome c oxidoreductase,
- mitochondrial electron transport complex III,
- ubiquinol-cytochrome c-2 oxidoreductase,
- ubiquinone-cytochrome b-c1 oxidoreductase,
- ubiquinol-cytochrome c2 reductase,
- ubiquinol-cytochrome c1 oxidoreductase,
- CoQH2-cytochrome c oxidoreductase,
- ubihydroquinol:cytochrome c oxidoreductase,
- coenzyme QH2-cytochrome c reductase, and
- QH2:cytochrome c oxidoreductase.
Structure
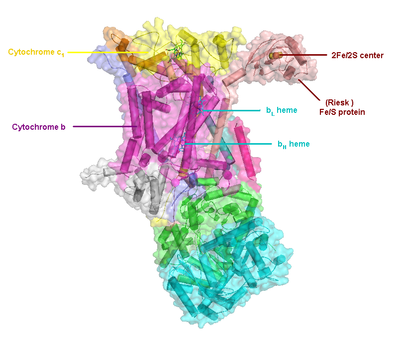
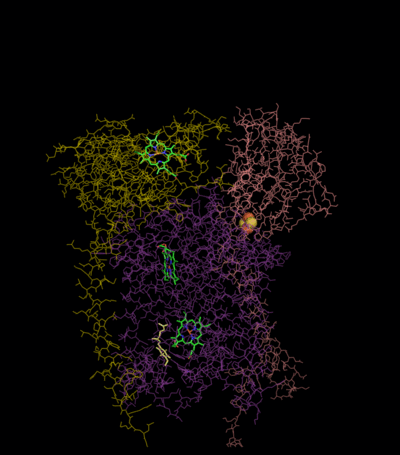
Compared to the other major proton-pumping subunits of the electron transport chain, the number of subunits found can be small, as small as three polypeptide chains. This number does increase, and eleven subunits are found in higher animals.[2] Three subunits have prosthetic groups. The cytochrome b subunit has two b-type hemes (bL and bH), the cytochrome c subunit has one c-type heme (c1), and the Rieske Iron Sulfur Protein subunit (ISP) has a two iron, two sulfur iron-sulfur cluster (2Fe•2S).
Structures of complex III: PDB 1KYO, PDB 1L0L
Reaction
It catalyzes the reduction of cytochrome c by oxidation of coenzyme Q (CoQ) and the concomitant pumping of 4 protons from the mitochondrial matrix to the intermembrane space:
- QH2 + 2 cytochrome c (FeIII) + 2 H+in → Q + 2 cytochrome c (FeII) + 4 H+out
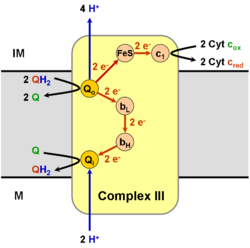
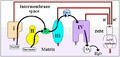
In the process called Q cycle,[3][4] two protons are consumed from the matrix (M), four protons are released into the inter membrane space (IM) and two electrons are passed to cytochrome c.
Reaction Mechanism
The reaction mechanism for complex III (Cytochrome bc1 , Coenzyme Q: Cytochrome C Oxidoreductase) is known as the ubiquinone ("Q") cycle. In this cycle four protons get released into the Positive "P" side (inter membrane space), but only two protons get taken up from the Negative "N" side (matrix). As a result a proton gradient is formed across the membrane. In the overall reaction, two ubiquinols are oxidized to ubiquinones and one ubiquinone is reduced to ubiquinol. In the complete mechanism, two electrons are transferred from ubiquinol to ubiquinone, via two cytochrome c intermediates.
Overall:
- 2 x QH2 oxidised to Q
- 1 x Q reduced to QH2
- 2 x Cyt c1 reduced
- 4 x H+ released into intermembrane space
- 2 x H+ picked up from matrix
The reaction proceeds according to the following steps:
Round 1:
- Cytochrome b binds a ubiquinol and a ubiquinone.
- The 2Fe/2S center and BL heme each pull an electron off the bound ubiquinol, releasing two hydrogens into the intermembrane space.
- One electron is transferred to cytochrome c1 from the 2Fe/2S centre, whilst another is transferred from the BL heme to the BH Heme.
- Cytochrome c1 transfers its electron to cytochrome c (not to be confused with cytochrome c1), and the BH Heme transfers its electron to a nearby ubiquinone, resulting in the formation of a ubisemiquinone.
- Cytochrome c diffuses. The first ubiquinol (now oxidised to ubiquinone) is released, whilst the semiquinone remains bound.
Round 2:
- A second ubiquinol is bound by cytochrome b.
- The 2Fe/2S center and BL heme each pull an electron off the bound ubiquinol, releasing two hydrogens into the intermembrane space.
- One electron is transferred to cytochrome c1 from the 2Fe/2S centre, whilst another is transferred from the BL heme to the BH Heme.
- Cytocrome c1 then transfers its electron to cytochrome c, whilst the nearby semiquinone picks up a second electron from the BH Heme, along with two protons from the matrix.
- The second ubiquinol (now oxidised to ubiquinone), along with the newly formed ubiquinol are released.[5]
Inhibitors of complex III
There are three distinct groups of Complex III inhibitors.
- Antimycin A binds to the Qi site and inhibits the transfer of electrons in Complex III from heme bH to oxidized Q (Qi site inhibitor).
- Myxothiazol and stigmatellin binds to the Qo site and inhibits the transfer of electrons from reduced QH2 to the Rieske Iron sulfur protein. Myxothiazol and stigmatellin bind to distinct pockets within the Qo site.
- Myxothiazol binds very close to cytochrome bL (hence termed a "proximal" inhibitor).
- Stigmatellin binds near the Rieske Iron sulfur protein, with which it strongly interacts.
Some have been commercialized as fungicides (the strobilurin derivatives, best known of which is azoxystrobin; QoI inhibitors) and as anti-malaria agents (atovaquone).
Oxygen free radicals
A small fraction of electrons leave the electron transport chain before reaching complex IV. Premature electron leakage to oxygen results in the formation of superoxide. The relevance of this otherwise minor side reaction is that superoxide and other reactive oxygen species are highly toxic and are thought to play a role in several pathologies, as well as aging (the free radical theory of aging).[6] Electron leakage occurs mainly at the Qo site and is stimulated by antimycin A. Antimycin A locks the b hemes in the reduced state by preventing their re-oxidation at the Qi site, which, in turn, causes the steady-state concentrations of the Qo semiquinone to rise, the latter species reacting with oxygen to form superoxide. The effect of high membrane potential is thought to have a similar effect.[7] Superoxide produced at the Qo site can be released both into the mitochondrial matrix[8][9] and into the intermembrane space (from where it can reach the cytosol.[8][10] This could be explained by the fact that Complex III might produce superoxide as membrane permeable HOO• rather than as membrane impermeable O2-..[9]
Mutations in Complex III genes in human disease
Mutations in Complex III-related genes typically manifest as exercise intolerance.[11][12] Other mutations have been reported to cause septo-optic displaisa[13] and multisystem disorders.[14] However, mutations in BCS1L, a gene responsible for proper maturation of Complex III, can result in Björnstad syndrome and the GRACILE syndrome, which in neonates are lethal conditions that have multisystem and neurologic manifestations typifying severe mitochondrial disorders. The pathogenicity of several mutations has been verified in model systems such as yeast.[15]
The extent to which these various pathologies are due to bioenergetic deficits or overeproduction of superoxide is presently unknown.
See also
Additional images
References
- ^ PDB 1ntz; Gao X, Wen X, Esser L, Quinn B, Yu L, Yu CA, Xia D (August 2003). "Structural basis for the quinone reduction in the bc1 complex: a comparative analysis of crystal structures of mitochondrial cytochrome bc1 with bound substrate and inhibitors at the Qi site". Biochemistry 42 (30): 9067–80. doi:10.1021/bi0341814. PMID 12885240.
- ^ Iwata S, Lee JW, Okada K, Lee JK, Iwata M, Rasmussen B, Link TA, Ramaswamy S, Jap BK (July 1998). "Complete structure of the 11-subunit bovine mitochondrial cytochrome bc1 complex". Science 281 (5373): 64–71. doi:10.1126/science.281.5373.64. PMID 9651245.
- ^ Kramer DM, Roberts AG, Muller F, Cape J, Bowman MK (2004). "Q-cycle bypass reactions at the Qo site of the cytochrome bc1 (and related) complexes". Meth. Enzymol.. Methods in Enzymology 382: 21–45. doi:10.1016/S0076-6879(04)82002-0. ISBN 9780121827861. PMID 15047094.
- ^ Crofts AR (2004). "The cytochrome bc1 complex: function in the context of structure". Annu. Rev. Physiol. 66: 689–733. doi:10.1146/annurev.physiol.66.032102.150251. PMID 14977419.
- ^ Ferguson SJ, Nicholls D, Ferguson S (2002). Bioenergetics (3rd ed.). San Diego: Academic. pp. 114–117. ISBN 0-12-518121-3.
- ^ Muller, F. L., Lustgarten, M. S., Jang, Y., Richardson, A. and Van Remmen, H. (2007). "Trends in oxidative aging theories". Free Radic. Biol. Med. 43 (4): 477–503. doi:10.1016/j.freeradbiomed.2007.03.034. PMID 17640558.
- ^ Skulachev VP (May 1996). "Role of uncoupled and non-coupled oxidations in maintenance of safely low levels of oxygen and its one-electron reductants". Q. Rev. Biophys. 29 (2): 169–202. PMID 8870073.
- ^ a b Muller F (2000). "The nature and mechanism of superoxide production by the electron transport chain: Its relevance to aging". AGE 23 (4): 227–253. doi:10.1007/s11357-000-0022-9.
- ^ a b Muller FL, Liu Y, Van Remmen H (November 2004). "Complex III releases superoxide to both sides of the inner mitochondrial membrane". J. Biol. Chem. 279 (47): 49064–73. doi:10.1074/jbc.M407715200. PMID 15317809.
- ^ Han D, Williams E, Cadenas E (January 2001). "Mitochondrial respiratory chain-dependent generation of superoxide anion and its release into the intermembrane space". Biochem. J. 353 (Pt 2): 411–6. doi:10.1042/0264-6021:3530411. PMC 1221585. PMID 11139407. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1221585.
- ^ DiMauro S (November 2006). "Mitochondrial myopathies". Curr Opin Rheumatol 18 (6): 636–41. doi:10.1097/01.bor.0000245729.17759.f2. PMID 17053512.
- ^ DiMauro S (June 2007). "Mitochondrial DNA medicine". Biosci. Rep. 27 (1–3): 5–9. doi:10.1007/s10540-007-9032-5. PMID 17484047.
- ^ Schuelke M, Krude H, Finckh B, Mayatepek E, Janssen A, Schmelz M, Trefz F, Trijbels F, Smeitink J (March 2002). "Septo-optic dysplasia associated with a new mitochondrial cytochrome b mutation". Ann. Neurol. 51 (3): 388–92. doi:10.1002/ana.10151. PMID 11891837.
- ^ Wibrand F, Ravn K, Schwartz M, Rosenberg T, Horn N, Vissing J (October 2001). "Multisystem disorder associated with a missense mutation in the mitochondrial cytochrome b gene". Ann. Neurol. 50 (4): 540–3. doi:10.1002/ana.1224. PMID 11601507.
- ^ Fisher N, Castleden CK, Bourges I, Brasseur G, Dujardin G, Meunier B (March 2004). "Human disease-related mutations in cytochrome b studied in yeast". J. Biol. Chem. 279 (13): 12951–8. doi:10.1074/jbc.M313866200. PMID 14718526.
Further reading
- Marres CM, Slater EC (1977). "Polypeptide composition of purified QH2:cytochrome c oxidoreductase from beef-heart mitochondria". Biochim. Biophys. Acta. 462 (3): 531–548. doi:10.1016/0005-2728(77)90099-8. PMID 597492.
- Rieske JS (1976). "Composition, structure, and function of complex III of the respiratory chain". Biochim. Biophys. Acta. 456 (2): 195–247. PMID 788795.
- Wikstrom M, Krab K, Saraste M (1981). "Proton-translocating cytochrome complexes". Annu. Rev. Biochem. 50: 623–655. doi:10.1146/annurev.bi.50.070181.003203. PMID 6267990.
External links
- cytochrome bc1 complex site (Edward A. Berry) at lbl.gov
- cytochrome bc1 complex site (Antony R. Crofts) at uiuc.edu
- PROMISE Database: cytochrome bc1 complex at scripps.edu
- Interactive Molecular Model of Complex III (Requires MDL Chime)
- UMich Orientation of Proteins in Membranes families/superfamily-3 - Calculated positions of bc1 and related complexes in membranes
- MeSH Coenzyme+Q-Cytochrome-c+Reductase
Oxidoreductases: diphenol family (EC 1.10) 1.10.1 1.10.2 Coenzyme Q - cytochrome c reductase1.10.3 1.10.99 Other B enzm: 1.1/2/3/4/5/6/7/8/10/11/13/14/15-18, 2.1/2/3/4/5/6/7/8, 2.7.10, 2.7.11-12, 3.1/2/3/4/5/6/7, 3.1.3.48, 3.4.21/22/23/24, 4.1/2/3/4/5/6, 5.1/2/3/4/99, 6.1-3/4/5-6 Metabolism: Citric acid cycle enzymes Cycle Anaplerotic to acetyl-CoAPyruvate dehydrogenase complex (E1, E2, E3)
(regulated by Pyruvate dehydrogenase kinase and Pyruvate dehydrogenase phosphatase)to succinyl-CoAto oxaloacetateMitochondrial
electron transport chain/
oxidative phosphorylationPrimaryComplex I/NADH dehydrogenase · Complex II/Succinate dehydrogenase · Coenzyme Q · Complex III/Coenzyme Q - cytochrome c reductase · Cytochrome c · Complex IV/Cytochrome c oxidase
Coenzyme Q10 synthesis: COQ2 · COQ3 · COQ4 · COQ5 · COQ6 · COQ7 · COQ9 · COQ10A · COQ10B · PDSS1 · PDSS2OtherETC Other B memb: cead, trns (1A, 1C, 1F, 2A, 3A1, 3A2-3, 3D), othr Categories:- EC 1.10.2
- Enzymes of known structure
- Cellular respiration
- Iron-sulfur proteins
- Integral membrane proteins
- QH2 + 2 ferricytochrome c
Wikimedia Foundation. 2010.