- Cytochrome b6f complex
-
Cytochrome b6f complex 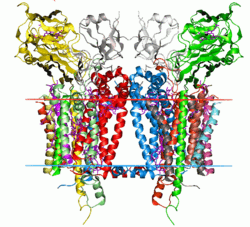
Crystal structure of the cytochrome b6f complex from C. reinhardtii (1q90). Hydrocarbon boundaries of the lipid bilayer are shown by red and blue dots. Identifiers EC number 1.10.99.1 Databases IntEnz IntEnz view BRENDA BRENDA entry ExPASy NiceZyme view KEGG KEGG entry MetaCyc metabolic pathway PRIAM profile PDB structures RCSB PDB PDBe PDBsum Search PMC articles PubMed articles The cytochrome b6f complex (plastoquinol—plastocyanin reductase; EC 1.10.99.1) is an enzyme found in the thylakoid membrane in chloroplasts of plants, cyanobacteria, and green algae, catalyzing the transfer of electrons from plastoquinol to plastocyanin.[1] The reaction is analogous to the reaction catalyzed by cytochrome bc1 (Complex III) of the mitochondrial electron transport chain. For photosynthesis, the cytochrome b6f complex transfers electrons between the two reaction complexes from Photosystem II to Photosystem I.
Contents
Enzyme structure
The cytochrome b6f complex is a dimer, with each monomer composed of eight subunits.[2] These consist of four large subunits: a 32 kDa cytochrome f with a c-type cytochrome, a 25 kDa cytochrome b6 with a low- and high-potential heme group, a 19 kDa Rieske iron-sulfur protein containing a [2Fe-2S] cluster, and a 17 kDa subunit IV; along with four small subunits (3-4 kDa): PetG, PetL, PetM, and PetN.[2][3] The total molecular weight is 217 kDa.
The crystal structure of cytochrome b6f complexes from Chlamydomonas reinhardtii, Mastigocladus laminosus, and Nostoc sp. PCC 7120 have been determined.[4][5][6] The complex is structurally similar to cytochrome bc1. Cytochrome b6 and subunit IV are homologous to cytochrome b[7] and the Rieske iron-sulfur proteins of the two complexes are homologous.[8] However, cytochrome f and cytochrome c1 are not homologous.[9]
Cytochrome b6f contains seven prosthetic groups.[10][11] Four are found in both cytochrome b6f and bc1: the c-type heme of cytochrome c1 and f, the two b-type hemes (bp and bn) in bc1 and b6f, and the [2Fe-2S] cluster of the Rieske protein. Three unique prosthetic groups are found in cytochrome b6f: chlorophyll a, β-carotene, and heme cn (also known as heme x).[4]
Biological function
In photosynthesis, the cytochrome b6f complex functions to mediate the transfer of electrons between the two photosynthetic reaction center complexes, from Photosystem II to Photosystem I, while transferring protons from the chloroplast stroma across the thylakoid membrane into the lumen.[12] Electron transport via cytochrome b6f is responsible for creating the proton gradient that drives the synthesis of ATP in chloroplasts.[3]
In a separate reaction, the cytochrome b6f complex plays a central role in cyclic photophosphorylation, when NADP+ is not available to accept electrons from reduced ferredoxin.[1] This cycle results in the creation of a proton gradient by cytochrome b6f, which can be used to drive ATP synthesis. It has also been shown that this cycle is essential for photosynthesis,[13] in which it is proposed to help maintain the proper ratio of ATP/NADPH production for carbon fixation.[14][15]
Reaction mechanism
The cytochrome b6f complex is responsible for "non-cyclic" (1) and "cyclic" (2) electron transfer between two mobile redox carriers, plastoquinol (QH2) and plastocyanin (Pc):
H2O → photosystem II → QH2 → Cyt b6f → Pc → photosystem I → NADPH (1) QH2 → Cyt b6f → Pc → photosystem I → Q (2) Cytochrome b6f catalyzes the transfer of electrons from plastoquinol to plastocyanin, while pumping two protons from the stroma into the thylakoid lumen:
- QH2 + 2Pc(Cu2+) + 2H+ (stroma) → Q + 2Pc(Cu+) + 4H+ (lumen)[1]
This reaction occurs through the Q cycle as in Complex III.[16] Plastoquinone acts as the electron carrier, transferring its two electrons to high- and low-potential electron transport chains (ETC) via a mechanism called electron bifurcation.[17]
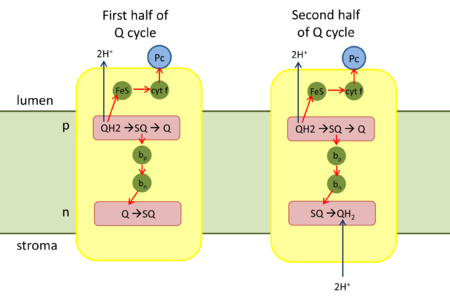
Q cycle
First half of Q cycle
- QH2 binds to the positive 'p' side (lumen side) of the complex. It is oxidized to a semiquinone (SQ) by the iron-sulfur center (high-potential ETC) and releases two protons to the thylakoid lumen.
- The reduced iron-sulfur center transfers its electron through cytochrome f to Pc.
- In the low-potential ETC, SQ transfers its electron to heme bp of cytochrome b6.
- Heme bp then transfers the electron to heme bn.
- Heme bn reduces Q with one electron to form SQ.
Second half of Q cycle
- A second QH2 binds to the complex.
- In the high-potential ETC, one electron reduces another oxidized Pc.
- In the low-potential ETC, the electron from heme bn is transferred to SQ, and the completely reduced Q2- takes up two protons from the stroma to form QH2.
- The oxidized Q and the reduced QH2 that has been regenerated diffuse into the membrane.
Cyclic electron transfer
In contrast to Complex III, cytochrome b6f catalyzes another electron transfer reaction that is central to cyclic photophosphorylation. The electron from ferredoxin (Fd) is transferred to plastoquinone and then the cytochrome b6f complex to reduce plastocyanin, which is reoxidized by P700 in Photosystem I.[18] The exact mechanism for how plastoquinone is reduced by ferredoxin is still under investigation. One proposal is that there exists a ferredoxin:plastoquinone-reductase or an NADP dehydrogenase.[18] Since heme x does not appear to be required for the Q cycle and is not found in Complex III,[12] it has been proposed that it is used for cyclic photophosphorylation by the following mechanism:[17][19]
- Fd (red) + heme x (ox) → Fd (ox) + heme x (red)
- heme x (red) + Fd (red) + Q + 2H+ → heme x (ox) + Fd (ox) + QH2
References
- ^ a b c Berg, Jeremy M. (Jeremy M.); Tymoczko, John L.; Stryer, Lubert.; Stryer, Lubert. Biochemistry. (2007). Biochemistr. New York: W.H. Freeman. ISBN 978-0-7167-8724-2.
- ^ a b Whitelegge, JP.; Zhang, H.; Aguilera, R.; Taylor, RM.; Cramer, WA. (Oct 2002). "Full subunit coverage liquid chromatography electrospray ionization mass spectrometry (LCMS+) of an oligomeric membrane protein: cytochrome b(6)f complex from spinach and the cyanobacterium Mastigocladus laminosus.". Mol Cell Proteomics 1 (10): 816–27. PMID 12438564.
- ^ a b Voet, Donald J. (2011). Biochemistry / Donald J. Voet ; Judith G. Voet. New York, NY: Wiley, J. ISBN 978-0-470-57095-1.
- ^ a b Stroebel, D.; Choquet, Y.; Popot, JL.; Picot, D. (Nov 2003). "An atypical haem in the cytochrome b(6)f complex.". Nature 426 (6965): 413–8. doi:10.1038/nature02155. PMID 14647374.
- ^ Yamashita, E.; Zhang, H.; Cramer, WA. (Jun 2007). "Structure of the cytochrome b6f complex: quinone analogue inhibitors as ligands of heme cn.". J Mol Biol 370 (1): 39–52. doi:10.1016/j.jmb.2007.04.011. PMID 17498743.
- ^ Baniulis, D.; Yamashita, E.; Whitelegge, JP.; Zatsman, AI.; Hendrich, MP.; Hasan, SS.; Ryan, CM.; Cramer, WA. (Apr 2009). "Structure-Function, Stability, and Chemical Modification of the Cyanobacterial Cytochrome b6f Complex from Nostoc sp. PCC 7120.". J Biol Chem 284 (15): 9861–9. doi:10.1074/jbc.M809196200. PMID 19189962.
- ^ Widger, WR.; Cramer, WA.; Herrmann, RG.; Trebst, A. (Feb 1984). "Sequence homology and structural similarity between cytochrome b of mitochondrial complex III and the chloroplast b6-f complex: position of the cytochrome b hemes in the membrane.". Proc Natl Acad Sci U S A 81 (3): 674–8. doi:10.1073/pnas.81.3.674. PMID 6322162.
- ^ Carrell, CJ.; Zhang, H.; Cramer, WA.; Smith, JL. (Dec 1997). "Biological identity and diversity in photosynthesis and respiration: structure of the lumen-side domain of the chloroplast Rieske protein.". Structure 5 (12): 1613–25. PMID 9438861.
- ^ Martinez, SE.; Huang, D.; Szczepaniak, A.; Cramer, WA.; Smith, JL. (Feb 1994). "Crystal structure of chloroplast cytochrome f reveals a novel cytochrome fold and unexpected heme ligation.". Structure 2 (2): 95–105. PMID 8081747.
- ^ Baniulis, D.; Yamashita, E.; Zhang, H.; Hasan, SS.; Cramer, WA.. "Structure-function of the cytochrome b6f complex.". Photochem Photobiol 84 (6): 1349–58. doi:10.1111/j.1751-1097.2008.00444.x. PMID 19067956.
- ^ Cramer, WA.; Zhang, H.; Yan, J.; Kurisu, G.; Smith, JL. (May 2004). "Evolution of photosynthesis: time-independent structure of the cytochrome b6f complex.". Biochemistry 43 (20): 5921–9. doi:10.1021/bi049444o. PMID 15147175.
- ^ a b Kurisu, G.; Zhang, H.; Smith, JL.; Cramer, WA. (Nov 2003). "Structure of the cytochrome b6f complex of oxygenic photosynthesis: tuning the cavity.". Science 302 (5647): 1009–14. doi:10.1126/science.1090165. PMID 14526088.
- ^ Munekage, Y.; Hashimoto, M.; Miyake, C.; Tomizawa, K.; Endo, T.; Tasaka, M.; Shikanai, T. (Jun 2004). "Cyclic electron flow around photosystem I is essential for photosynthesis.". Nature 429 (6991): 579–82. doi:10.1038/nature02598. PMID 15175756.
- ^ Blankenship, Robert E. (2002). Molecular mechanisms of photosynthesi. Oxford ; Malden, MA: Blackwell Science. ISBN 978-0-632-04321-7.
- ^ Bendall, Derek. "Cyclic photophosphorylation and electron transport". Biochimica et Biophysica Acta (BBA) - Bioenergetics. doi:10.1016/0005-2728(94)00195-B.
- ^ Cramer, WA.; Soriano, GM.; Ponomarev, M.; Huang, D.; Zhang, H.; Martinez, SE.; Smith, JL. (Jun 1996). "SOME NEW STRUCTURAL ASPECTS AND OLD CONTROVERSIES CONCERNING THE CYTOCHROME b6f COMPLEX OF OXYGENIC PHOTOSYNTHESIS.". Annu Rev Plant Physiol Plant Mol Biol 47: 477–508. doi:10.1146/annurev.arplant.47.1.477. PMID 15012298.
- ^ a b Cramer, WA.; Zhang, H.; Yan, J.; Kurisu, G.; Smith, JL. (2006). "Transmembrane traffic in the cytochrome b6f complex.". Annu Rev Biochem 75: 769–90. doi:10.1146/annurev.biochem.75.103004.142756. PMID 16756511.
- ^ a b "cyclic electron transfer in plant leaf". PNAS. doi:10.1073/pnas.102306999.
- ^ Cramer, WA.; Yan, J.; Zhang, H.; Kurisu, G.; Smith, JL. (2005). "Structure of the cytochrome b6f complex: new prosthetic groups, Q-space, and the 'hors d'oeuvres hypothesis' for assembly of the complex.". Photosynth Res 85 (1): 133–43. doi:10.1007/s11120-004-2149-5. PMID 15977064.
External links
- 1Q90 - PDB structure of cytochrome b6f complex from Chlamydomonas reinhardtii
- 1VF5 - PDB structure of cytochrome b6f complex from Mastigocladus laminosus
- 2D2C - PDB structure of cytochrome b6f complex from Mastigocladus laminosus
- 2E74 - PDB structure of cytochrome b6f complex from Mastigocladus laminosus
- 2E75 - PDB structure of cytochrome b6f complex from Mastigocladus laminosus
- 2E76 - PDB structure of cytochrome b6f complex from Mastigocladus laminosus
- 2ZT9 - PDB structure of cytochrome b6f complex from Nostoc sp. PCC 7120
- Structure-Function Studies of the Cytochrome b6f Complex - Current research on cytochrome b6f in William Cramer's Lab at Purdue University, USA
- UMich Orientation of Proteins in Membranes families/superfamily-3 - Calculated positions of b6f and related complexes in membranes
- MeSH Cytochrome+b6f+Complex
ETC Other Categories:- Hemoproteins
- Iron-sulfur proteins
- Light reactions
- Integral membrane proteins
Wikimedia Foundation. 2010.