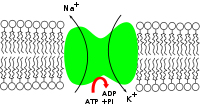
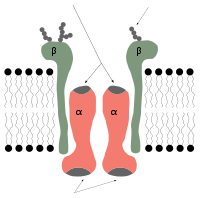
- Na+/K+-ATPase
-
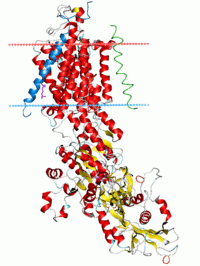 Sodium-potassium pump, E2-Pi state. Calculated hydrocarbon boundaries of the lipid bilayer are shown as blue (intracellular) and red (extracellular) planes
Sodium-potassium pump, E2-Pi state. Calculated hydrocarbon boundaries of the lipid bilayer are shown as blue (intracellular) and red (extracellular) planes
Na+/K+-ATPase (fully sodium-potassium adenosine triphosphatase, also known as the Na+/K+ pump, sodium-potassium pump, or sodium pump, for short) is an enzyme (EC 3.6.3.9) located in the plasma membrane (to be specific, an electrogenic transmembrane ATPase) in all animals.
Contents
Sodium-potassium pumps
Active transport is responsible for cells containing relatively high concentrations of potassium ions but low concentrations of sodium ions. The mechanism responsible for this is the sodium-potassium pump, which moves these two ions in opposite directions across the plasma membrane. This was investigated by following the passage of radioactively labeled ions across the plasma membrane of certain cells. It was found that the concentrations of sodium and potassium ions on the two other sides of the membrane are interdependent, suggesting that the same carrier transports both ions. It is now known that the carrier is an ATP-ase and that it pumps three sodium ions out of the cell for every two potassium ions pumped in.
The sodium-potassium pump was discovered in the 1950s by a Danish scientist, Jens Christian Skou, who was awarded a Nobel Prize in 1997. It marked an important step forward in our understanding of how ions get into and out of cells, and it has a particular significance for excitable cells such as nervous cells, which depend on it for responding to stimuli and transmitting impulses.
(Advanced Biology - Michael Roberts, Michael Reiss, nasif mohammed, Grace Monger. 2000)
Function
The Na+/K+-ATPase helps maintain resting potential, avail transport, and regulate cellular volume.[citation needed] It also functions as signal transducer/integrator to regulate MAPK pathway, ROS, as well as intracellular calcium. For most animal cells, the Na+/K+-ATPase is responsible for 1/3 of the cell's energy expenditure. For neurons, the Na+/K+-ATPase is responsible for 2/3 of the cell's energy expenditure. [citation needed]
Resting potential
See also: Resting potentialIn order to maintain the cell membrane potential, cells keep a low concentration of sodium ions and high levels of potassium ions within the cell (intracellular). The sodium-potassium pump moves 3 sodium ions out and moves 2 potassium ions in, thus in total removing one positive charge carrier from the intracellular space. Please see Mechanism for details.
Not only the mechanism of the sodium-potassium pump alone is responsible for the generation of the resting membrane potential. Also the selective permeability of the cell's plasma membrane for the different Ions plays an important role. All mechanisms involved are explained in the main article on generation of the resting membrane potential. The importance of membrane permeability is explained in a video on Khan Academy.
Transport
Export of sodium from the cell provides the driving force for several secondary active transporters membrane transport proteins, which import glucose, amino acids, and other nutrients into the cell by use of the sodium gradient.
Another important task of the Na+-K+ pump is to provide a Na+ gradient that is used by certain carrier processes. In the gut, for example, sodium is transported out of the reabsorbing cell on the blood (interstitial fluid) side via the Na+-K+ pump, whereas, on the reabsorbing (luminal) side, the Na+-Glucose symporter uses the created Na+ gradient as a source of energy to import both Na+ and glucose, which is far more efficient than simple diffusion. Similar processes are located in the renal tubular system.
Controlling cell volume
One of the important functions of Na+-K+ pump is to maintain the volume of the cell. Inside the cell, there are many proteins and other organic compounds that cannot escape from the cell. Most, being negatively charged, collect around them a large number of positive ions. All these substances tend to cause the osmosis of water into the cell, which, unless checked, can cause the cell to swell up and lyse. The Na+-K+ pump is a mechanism to prevent this. The pump transports 3 Na+ ions out of the cell and in exchange takes 2 K+ ions into the cell. As the membrane is far less permeable to Na+ ions than K+ ions, the sodium ions have a tendency to stay there. In addition, while the cell membrane is impermeable to sodium ions, potassium leak channels embedded in the membrane allow for K+ ions to "leak" back out of the cell down their concentration gradient. This represents a continual net loss of ions out of the cell. The opposing osmotic tendency that results operates to drive the water molecules out of the cells. Furthermore, when the cell begins to swell, this automatically activates the Na+-K+ pump, which moves still more ions to the exterior.
Functioning as signal transducer
Within the last decade, many independent labs have demonstrated that, in addition to the classical ion transporting, this membrane protein can also relay extracellular ouabain-binding signalling into the cell through regulation of protein tyrosine phosphorylation. The downstream signals through ouabain-triggered protein phosphorylation events include to activate the mitogen-activated protein kinase (MAPK) signal cascades, mitochondrial reactive oxygen species (ROS) production, as well as activation of phospholipase C (PLC) and inositol triphosphate (IP3) receptor (IP3R) in different intracellular compartments.[1]
Protein-protein interactions play very important role in Na+-K+ pump-mediated signal transduction. For example, Na+-K+ pump interacts directly with Src, a non-receptor tyrosine kinase, to form a signaling receptor complex.[2] Src kinase is inhibited by Na+-K+ pump, while, upon ouabain binding, Src kinase domain will be released and then activated. Based on this scenario, NaKtide, a peptide Src inhibitor derived from Na+-K+ pump, was developed as a functional ouabain antagonist.[3] Na+-K+ pump also interacts with ankyrin, IP3R, PI3K, PLC-gamma and cofilin.[4]
Mechanism
- The pump, with good binds ATP, binds 3 intracellular Na+ ions.
- ATP is hydrolyzed, leading to phosphorylation of the pump at a highly conserved aspartate residue and subsequent release of ADP.[citation needed]
- A conformational change in the pump exposes the Na+ ions to the outside. The phosphorylated form of the pump has a low affinity for Na+ ions, so they are released.[citation needed]
- The pump binds 2 extracellular K+ ions. This causes the dephosphorylation of the pump, reverting it to its previous conformational state, transporting the K+ ions into the cell.[citation needed]
- The unphosphorylated form of the pump has a higher affinity for Na+ ions than K+ ions, so the two bound K+ ions are released. ATP binds, and the process starts again.[citation needed]
Regulation
Endogenous
The Na+/K+-ATPase is upregulated by cAMP.[5] Thus, substances causing an increase in cAMP upregulate the Na+/K+-ATPase. These include the ligands of the Gs-coupled GPCRs. In contrast, substances causing a decrease in cAMP downregulate the Na+/K+-ATPase. These include the ligands of the Gi-coupled GPCRs.
Note: Early studies indicated the opposite effect, but these were later found to be inaccurate due to additional complicating factors. [citation needed]
Exogenous
The Na+-K+-ATPase can be pharmacologically modified by administrating drugs exogenously.
For instance, Na+-K+-ATPase found in the membrane of heart cells is an important target of cardiac glycosides (for example digoxin and ouabain), inotropic drugs used to improve heart performance by increasing its force of contraction.
Contraction of any muscle is dependent on a 100- to 10,000-times-higher-than-resting intracellular Ca2+ concentration, which, as soon as it is put back again on its normal level by a carrier enzyme in the plasma membrane, and a calcium pump in sarcoplasmic reticulum, muscle relaxes.
Since this carrier enzyme (Na+-Ca2+ translocator) uses the Na gradient generated by the Na+-K+ pump to remove Ca2+ from the intracellular space, slowing down the Na+-K+ pump results in a permanently-higher Ca2+ level in the muscle, which will eventually lead to stronger contractions.
Discovery
Na+/K+-ATPase was discovered by Jens Christian Skou in 1957 while working as assistant professor at the Department of Physiology, University of Aarhus, Denmark. He published his work that year.[6]
In 1997, he received one-half of the Nobel Prize in Chemistry "for the first discovery of an ion-transporting enzyme, Na+, K+ -ATPase."[7]
Genes
- Alpha: ATP1A1[1], ATP1A2[2], ATP1A3[3], ATP1A4[4]. #1 predominates in kidney. #2 is also known as "alpha(+)"
- Beta: ATP1B1[5], ATP1B2, ATP1B3[6], ATP1B4
See also
References
- ^ Yuan Z, Cai T, Tian J, Ivanov AV, Giovannucci DR, Xie Z (September 2005). "Na/K-ATPase tethers phospholipase C and IP3 receptor into a calcium-regulatory complex". Molecular Biology of the Cell 16 (9): 4034–45. doi:10.1091/mbc.E05-04-0295. PMC 1196317. PMID 15975899. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1196317.
- ^ Tian J, Cai T, Yuan Z, et al. (January 2006). "Binding of Src to Na+/K+-ATPase forms a functional signaling complex". Molecular Biology of the Cell 17 (1): 317–26. doi:10.1091/mbc.E05-08-0735. PMC 1345669. PMID 16267270. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1345669.
- ^ Li Z, Cai T, Tian J, et al. (July 2009). "NaKtide, a Na/K-ATPase-derived peptide Src inhibitor, antagonizes ouabain-activated signal transduction in cultured cells". The Journal of Biological Chemistry 284 (31): 21066–76. doi:10.1074/jbc.M109.013821. PMC 2742871. PMID 19506077. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2742871.
- ^ Lee K, Jung J, Kim M, Guidotti G (January 2001). "Interaction of the alpha subunit of Na,K-ATPase with cofilin". The Biochemical Journal 353 (2): 377–85. doi:10.1042/0264-6021:3530377. PMC 1221581. PMID 11139403. http://www.biochemj.org/bj/353/0377/bj3530377.htm.
- ^ Sodium in Health and Disease Michel Burnier. Informa HealthCare. New York, New York.
- ^ Skou JC (February 1957). "The influence of some cations on an adenosine triphosphatase from peripheral nerves". Biochimica et Biophysica Acta 23 (2): 394–401. doi:10.1016/0006-3002(57)90343-8. PMID 13412736.
- ^ Chemistry 1997
External links
Hydrolases: acid anhydride hydrolases (EC 3.6) 3.6.1 3.6.2 3.6.3-4: ATPase 3.6.3Cu++ (3.6.3.4)Ca+ (3.6.3.8)Na+/K+ (3.6.3.9)H+/K+ (3.6.3.10)ATP4AOther P-type ATPase3.6.43.6.5: GTPase 3.6.5.1: Heterotrimeric G protein3.6.5.2: Small GTPase > Ras superfamily3.6.5.3: Protein-synthesizing GTPase3.6.5.5-6: Polymerization motorsF- and V-type ATPase (3.A.2) P-type ATPase (3.A.3) - 3.A.3.1.1: Na+/K+ transporting: ATP1A1, ATP1A2, ATP1A3, ATP1A4, ATP1B1, ATP1B2, ATP1B3, ATP1B4, ATP1G1
- 3.A.3.1.2: H+/K+, H+/K+ exchanging: ATP4A, ATP4B
- 3.A.3.1.4: H+/K+ transporting, nongastric: ATP12A
- 3.A.3.2: Ca+ (SERCA, PMCA, SPCA) / Ca++ transporting: ATP2A1, ATP2A2, ATP2A3, ATP2B1, ATP2B2, ATP2B3, ATP2B4, ATP2C1
- 3.A.3.8.8: flippase: ATP8A2
Other/ungrouped:
Na+/K+ - H+
Categories:- EC 3.6.3
- Transport proteins
Wikimedia Foundation. 2010.