- mir-96 microRNA
-
microRNA mir-96 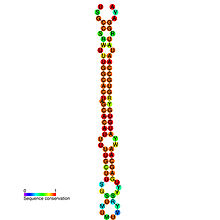
Predicted secondary structure and sequence conservation of mir-96 Identifiers Symbol mir-96 Alt. Symbols MIR96 Rfam RF00669 miRBase MI0000098 miRBase family MIPF0000072 Other data RNA type Gene; miRNA Domain(s) Eukaryota GO 0035068 0035195 SO 0001244
miR-96 microRNA precursor is a small non-coding RNA that regulates gene expression. microRNAs are transcribed as ~80 nucleotide precursors and subsequently processed by the Dicer enzyme to give a ~23 nucleotide products. In this case the mature sequence comes from the 5' arm of the precursor. [1] The mature products are thought to have regulatory roles through complementarity to mRNA. These microRNAs are expressed specifically in the inner ear and the adult eye.[2][3]miR-96 is thought to be conserved within Nephrozoa, i.e. the Deuterostomes and Protostomes. [4]
Variation within the seed region of mature miR-96 has been associated with autosomal dominant, progressive hearing loss in humans and mice. The homozygous mutant mice were profoundly deaf, showing no cochlear responses. Heterozygous mice and humans progressively lose the ability to hear. [5] [6] [7] Five genes, of 132 predicted targets, have been experimentally validated as targets of miR-96: Aqp5, Celsr2, Myrip, Odf2 and Ryk.[6]
Microarray analysis of 4-day old wildtype and mutant mice showed that in the 3' UTR of upregulated genes, there was a significant enrichment in heptamers complementary to miR-96, implying that miR-96 normally affects a wide range of target genes, and that the mutation results in a loss of normal targets. Among the downregulated genes, there is a significant enrichment in heptamers complementary to the mutant miR-96, so the mutant miR-96 has gained novel targets.[6] Among the downregulated genes were five of particular interest; Ocm, Pitpnm1, Prestin, Ptprq and Gfi1, all of which are strongly and specifically expressed in hair cells. Mice mutant for the latter three exhibit deafness and hair cell degeneration. [8] [9] [10]
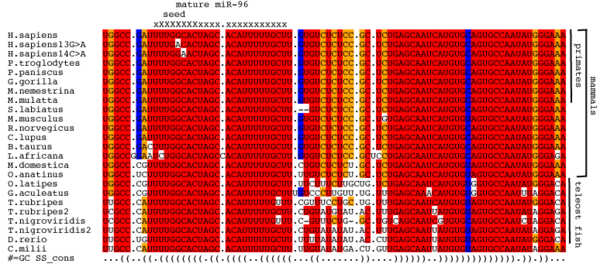 A multiple sequence alignment of precursor miR-96 molecules. Highly conserved nucleotides are coloured in red, less well conserved nucleotides are coloured orange and non-conserved nucleotides are coloured blue or white. The columns corresponding to the mature and seed sequence are indicated above the alignment. The canonical human sequence and the two human variant sequences that are implicated in hearing loss (13G>A and 14C>A) are in the first, second and third rows respectively.
A multiple sequence alignment of precursor miR-96 molecules. Highly conserved nucleotides are coloured in red, less well conserved nucleotides are coloured orange and non-conserved nucleotides are coloured blue or white. The columns corresponding to the mature and seed sequence are indicated above the alignment. The canonical human sequence and the two human variant sequences that are implicated in hearing loss (13G>A and 14C>A) are in the first, second and third rows respectively.
References
- ^ Mourelatos Z, Dostie J, Paushkin S, Sharma A, Charroux B, Abel L, Rappsilber J, Mann M, Dreyfuss G (2002). "miRNPs: a novel class of ribonucleoproteins containing numerous microRNAs.". Genes Dev 16 (6): 720–8. doi:10.1101/gad.974702. PMC 155365. PMID 11914277. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=155365.
- ^ Weston MD, Pierce ML, Rocha-Sanchez S, Beisel KW, Soukup GA (2006). "MicroRNA gene expression in the mouse inner ear.". Brain Res 1111 (1): 95–104. doi:10.1016/j.brainres.2006.07.006. PMID 16904081.
- ^ Xu S, Witmer PD, Lumayag S, Kovacs B, Valle D (2007). "MicroRNA (miRNA) transcriptome of mouse retina and identification of a sensory organ-specific miRNA cluster.". J Biol Chem 282 (34): 25053–66. doi:10.1074/jbc.M700501200. PMID 17597072.
- ^ Wheeler BM, Heimberg AM, Moy VN, Sperling EA, Holstein TW, Heber S, Peterson KJ (2009). "The deep evolution of metazoan microRNAs.". Evol Dev 11 (1): 50–68. doi:10.1111/j.1525-142X.2008.00302.x. PMID 19196333.
- ^ Mencía A, Modamio-Høybjør S, Redshaw N, Morín M, Mayo-Merino F, Olavarrieta L, Aguirre LA, del Castillo I, Steel KP, Dalmay T, Moreno F, Moreno-Pelayo MA (2009). "Mutations in the seed region of human miR-96 are responsible for nonsyndromic progressive hearing loss.". Nat Genet 41 (5): 609–13. doi:10.1038/ng.355. PMID 19363479.
- ^ a b c Lewis MA, Quint E, Glazier AM, Fuchs H, De Angelis MH, Langford C, van Dongen S, Abreu-Goodger C, Piipari M, Redshaw N, Dalmay T, Moreno-Pelayo MA, Enright AJ, Steel KP (2009). "An ENU-induced mutation of miR-96 associated with progressive hearing loss in mice.". Nat Genet 41 (5): 614–8. doi:10.1038/ng.369. PMC 2705913. PMID 19363478. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2705913.
- ^ Soukup GA (2009). "Little but loud: Small RNAs have a resounding affect on ear development.". Brain Res 1277: 104–14. doi:10.1016/j.brainres.2009.02.027. PMC 2700218. PMID 19245798. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2700218.
- ^ Liberman MC, Gao J, He DZ, Wu X, Jia S, Zuo J (2002). "Prestin is required for electromotility of the outer hair cell and for the cochlear amplifier.". Nature 419 (6904): 300–4. doi:10.1038/nature01059. PMID 12239568.
- ^ Goodyear RJ, Legan PK, Wright MB, Marcotti W, Oganesian A, Coats SA, Booth CJ, Kros CJ, Seifert RA, Bowen-Pope DF, Richardson GP (2003). "A receptor-like inositol lipid phosphatase is required for the maturation of developing cochlear hair bundles.". J Neurosci 23 (27): 9208–19. PMID 14534255.
- ^ Wallis D, Hamblen M, Zhou Y, Venken KJ, Schumacher A, Grimes HL, Zoghbi HY, Orkin SH, Bellen HJ (2003). "The zinc finger transcription factor Gfi1, implicated in lymphomagenesis, is required for inner ear hair cell differentiation and survival.". Development 130 (1): 221–32. doi:10.1242/dev.00190. PMID 12441305.
Media reports
Further reading
- Kondkar AA, Bray MS, Leal SM, et al. (2010). "VAMP8/endobrevin is overexpressed in hyperreactive human platelets: suggested role for platelet microRNA.". J. Thromb. Haemost. 8 (2): 369–78. doi:10.1111/j.1538-7836.2009.03700.x. PMID 19943878.
- Modamio-Høybjør S, Moreno-Pelayo MA, Mencía A, et al. (2004). "A novel locus for autosomal dominant nonsyndromic hearing loss, DFNA50, maps to chromosome 7q32 between the DFNB17 and DFNB13 deafness loci.". J. Med. Genet. 41 (2): e14. doi:10.1136/jmg.2003.012500. PMC 1735673. PMID 14757864. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1735673.
- Guttilla IK, White BA (2009). "Coordinate regulation of FOXO1 by miR-27a, miR-96, and miR-182 in breast cancer cells.". J. Biol. Chem. 284 (35): 23204–16. doi:10.1074/jbc.M109.031427. PMC 2749094. PMID 19574223. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2749094.
External links
- Rfam page for the mir-96 microRNA precursor family
- miRBase page for the mir-96 microRNA precursor family
- HGNC page for the mir-96 microRNA precursor family
- OMIM page for the mir-96 microRNA precursor family
- ENTREZ page for the mir-96 microRNA precursor family
miRNA precursor families 1-100 101-200 201+ Other Categories:
Wikimedia Foundation. 2010.
