- Alcohol dehydrogenase
-
alcohol dehydrogenase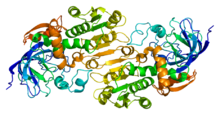 
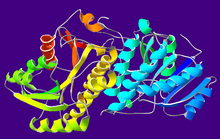
Crystallographic structure of the
homodimer of human ADH5.[1]Identifiers EC number 1.1.1.1 CAS number 9031-72-5 Databases IntEnz IntEnz view BRENDA BRENDA entry ExPASy NiceZyme view KEGG KEGG entry MetaCyc metabolic pathway PRIAM profile PDB structures RCSB PDB PDBe PDBsum Gene Ontology AmiGO / EGO Search PMC articles PubMed articles Alcohol dehydrogenases (ADH) (EC 1.1.1.1) are a group of dehydrogenase enzymes that occur in many organisms and facilitate the interconversion between alcohols and aldehydes or ketones with the reduction of nicotinamide adenine dinucleotide (NAD+ to NADH). In humans and many other animals, they serve to break down alcohols that otherwise are toxic, and they also participate in generation of useful aldehyde, ketone, or alcohol groups during biosynthesis of various metabolites. In yeast, plants, and many bacteria, some alcohol dehydrogenases catalyze the opposite reaction as part of fermentation to ensure a constant supply of NAD+.
Contents
Evolution
Genetic evidence from comparisons of multiple organisms, showed that a glutathione-dependent formaldehyde dehydrogenase, identical to a class III alcohol dehydrogenase (ADH-3/ADH5), probably is the ancestral enzyme for the entire ADH family.[2][3] Early on in evolution, an effective method for eliminating both endogenous and exogenous formaldehyde was important and this capacity has conserved the ancestral ADH-3 through time. From genetic duplications of ADH-3, followed by series of mutations, the other ADHs evolved.[2][3] The ability to produce ethanol from sugar is believed to have initially evolved in yeast. This feature is not adaptive from an energetic point of view, but by making alcohol in such high concentrations so that they were toxic to other organisms, yeast cells could effectively eliminate their competition. Since rotting fruit can contain more than 4% of ethanol, animals eating the fruit needed a system to metabolize exogenous ethanol. This was thought to explain the conservation of ethanol active ADH in other species than yeast, though ADH-3 is now known to also have a major role in nitric oxide signaling.[4][5]
Discovery
The first ever isolated alcohol dehydrogenase (ADH) was purified in 1937 from Saccharomyces cerevisiae (baker’s yeast).[6] Many aspects of the catalytic mechanism for the horse liver ADH enzyme was investigated by Hugo Theorell and coworkers.[7] ADH was also one of the first oligomeric enzymes that had its amino acid sequence and three dimensional structure determined.[8][9][10]
In early 1960 it was discovered in fruit flies of the genus Drosophila.[11]
Properties
The alcohol dehydrogenases comprise a group of several isozymes that catalyse the oxidation of primary and secondary alcohols to aldehydes and ketones, respectively, and also can catalyse the reverse reaction.[11] In mammals this is a redox (reduction/oxidation) reaction involving the coenzyme nicotinamide adenine dinucleotide (NAD+) and cofactor PQQ.
Alcohol dehydrogenase is a dimer with a mass of 80 kDa.[12]
Oxidation of alcohol
Mechanism of action in humans
Steps
- Binding of the coenzyme NAD+;
- Binding of the alcohol substrate by coordination to zinc;
- Deprotonation of His-51;
- Deprotonation of nicotinamide ribose;
- Deprotonation of Ser-48;
- Deprotonation of the alcohol;
- Hydride transfer from the alkoxide ion to NAD+, leading to NADH and a zinc bound aldehyde or ketone;
- Release of the product aldehyde;
The mechanism in yeast and bacteria is the reverse of this reaction. These steps are supported through kinetic studies.[12]
Involved subunits
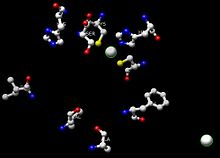
The substrate is coordinated to the zinc and this enzyme has two zinc atoms per subunit. One is the active site, which is involved in catalysis. In the active site, the ligands are Cys-46, Cys-174, His-67, and one water molecule. The other subunit is involved with structure. In this mechanism, the hydride from the alcohol goes to NAD+. Crystal structures indicate that the His-51 deprotonates the nicotinamide ribose, which deprotonates Ser-48. Finally, Ser-48 deprotonates the alcohol, making it an aldehyde.[12] From a mechanistic perspective, if the enzyme adds hydride to the re face of NAD+, the resulting hydrogen is incorporated into the pro-R position. Enzymes that add hydride to the re face are deemed Class A dehydrogenases.
Active site
The active site consists of a zinc atom, His-67, Cys-174, Cys-46, Ser-48, His-51, Ile-269, Val-292, Ala-317, and Phe-319. The zinc coordinates the substrate(alcohol). The zinc is coordinated by Cys-46, Cys-174, and His-67. Phe-319, Ala-317, His-51, Ile-269 and Val-292 stabilize NAD+ by forming hydrogen bonds. His-51 and Ile-269 form hydrogen bonds with the alcohols on nicotinamide ribose. Phe-319, Ala-317 and Val-292 form hydrogen bonds with the amide on NAD+.[12]
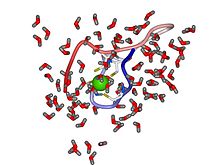
Structural zinc site
Mammalian alcohol dehydrogenases also has a structural zinc site. This Zn ion plays a structural role and is crucial for protein stability. The structures of the catalytic and structural zinc sites in horse liver alcohol dehydrogenase (HLADH) as revealed in crystallographic structures, which has been studied computationally with quantum chemical as well as with classical molecular dynamics methods. The structural zinc site is composed of four closely spaced cysteine ligands (Cys97, Cys100, Cys103, and Cys111 in the amino acid sequence) positioned in an almost symmetric tetrahedron around the Zn ion. A recent study showed that the interaction between zinc and cysteine is governed by primarily an electrostatic contribution with an additional covalent contribution to the binding.[13]
Types
Human
In humans, ADH exists in multiple forms as a dimer and is encoded by at least seven different genes. There are five classes (I-V) of alcohol dehydrogenase, but the hepatic form that is primarily used in humans is class 1. Class 1 consists of α, β, and γ subunits that are encoded by the genes ADH1A, ADH1B, and ADH1C.[14] The enzyme is present at high levels in the liver and the lining of the stomach.[15] It catalyzes the oxidation of ethanol to acetaldehyde:
- CH3CH2OH + NAD+ → CH3CHO + NADH + H+
This allows the consumption of alcoholic beverages, but its evolutionary purpose is probably the breakdown of alcohols naturally contained in foods or produced by bacteria in the digestive tract as alcohols are relatively 'empty' calories, providing little net nutritional benefit.[16]
Another evolutionary purpose may be metabolism of the endogenous alcohol vitamin A (retinol), which generates the hormone retinoic acid, although the function here may be primarily the elimination of toxic levels of retinol.[17]
alcohol dehydrogenase 1A,
α polypeptideIdentifiers Symbol ADH1A Alt. symbols ADH1 Entrez 124 HUGO 249 OMIM 103700 RefSeq NM_000667 UniProt P07327 Other data EC number 1.1.1.1 Locus Chr. 4 q23 alcohol dehydrogenase 1B,
β polypeptideIdentifiers Symbol ADH1B Alt. symbols ADH2 Entrez 125 HUGO 250 OMIM 103720 RefSeq NM_000668 UniProt P00325 Other data EC number 1.1.1.1 Locus Chr. 4 q23 alcohol dehydrogenase 1C,
γ polypeptideIdentifiers Symbol ADH1C Alt. symbols ADH3 Entrez 126 HUGO 251 OMIM 103730 RefSeq NM_000669 UniProt P00326 Other data EC number 1.1.1.1 Locus Chr. 4 q23 Alcohol dehydrogenase is also involved in the toxicity of other types of alcohol: For instance, it oxidizes methanol to produce formaldehyde and ethylene glycol to ultimately yield glycolic and oxalic acids. Humans have at least six slightly different alcohol dehydrogenases. Each is a dimer (i.e., consists of two polypeptides), with each dimer containing two zinc ions Zn2+. One of those ions is crucial for the operation of the enzyme: It is located at the catalytic site and holds the hydroxyl group of the alcohol in place.
Alcohol dehydrogenase activity varies between men and women, between young and old, and among populations from different areas of the world. For example, young women are unable to process alcohol at the same rate as young men because they do not express the alcohol dehydrogenase as highly, although, the inverse is true among the middle-aged.[18] The level of activity may not be dependent only on level of expression but also on allelic diversity among the population.
The human genes that encode class II, III, IV, and V alcohol dehydrogenases are ADH4, ADH5, ADH7, and ADH6, respectively.
alcohol dehydrogenase 4
(class II), π polypeptideIdentifiers Symbol ADH4 Entrez 127 HUGO 252 OMIM 103740 RefSeq NM_000670 UniProt P08319 Other data EC number 1.1.1.1 Locus Chr. 4 q22 alcohol dehydrogenase 5
(class III), χ polypeptideIdentifiers Symbol ADH5 Entrez 128 HUGO 253 OMIM 103710 RefSeq NM_000671 UniProt P11766 Other data EC number 1.1.1.1 Locus Chr. 4 q23 alcohol dehydrogenase 6
(class V)Identifiers Symbol ADH6 Entrez 130 HUGO 255 OMIM 103735 RefSeq NM_000672 UniProt P28332 Other data EC number 1.1.1.1 Locus Chr. 4 q23 alcohol dehydrogenase 7
(class IV), μ or σ polypeptideIdentifiers Symbol ADH7 Entrez 131 HUGO 256 OMIM 600086 RefSeq NM_000673 UniProt P40394 Other data EC number 1.1.1.1 Locus Chr. 4 q23-q24 Yeast and bacteria
Unlike humans, yeast and bacteria (except lactic acid bacteria, and E. coli in certain conditions ) do not ferment glucose to lactate. Instead, they ferment it to ethanol and CO2. The overall reaction can be seen below:
- Glucose + 2 ADP + 2 Pi → 2 ethanol + 2 CO2 + 2 ATP + 2 H2O[19]
In yeast[20] and many bacteria, alcohol dehydrogenase plays an important part in fermentation: pyruvate resulting from glycolysis is converted to acetaldehyde and carbon dioxide, and the acetaldehyde is then reduced to ethanol by an alcohol dehydrogenase called ADH1. The purpose of this latter step is the regeneration of NAD+, so that the energy-generating glycolysis can continue. Humans exploit this process to produce alcoholic beverages, by letting yeast ferment various fruits or grains. It is interesting to note that yeast can produce and consume their own alcohol.
The main alcohol dehydrogenase in yeast is larger than the human one, consisting of four rather than just two subunits. It also contains zinc at its catalytic site. Together with the zinc-containing alcohol dehydrogenases of animals and humans, these enzymes from yeasts and many bacteria form the family of "long-chain"-alcohol dehydrogenases.
Brewer's yeast also has another alcohol dehydrogenase, ADH2, which evolved out of a duplicate version of the chromosome containing the ADH1 gene. ADH2 is used by the yeast to convert ethanol back into acetaldehyde, and it is only expressed when sugar concentration is low. Having these two enzymes allows yeast to produce alcohol when sugar is plentiful (and this alcohol then kills off competing microbes), and then continue with the oxidation of the alcohol once the sugar, and competition, is gone.[21]
Plants
In plants, ADH catalyses the same reaction as in yeast and bacteria to ensure that there is a constant supply of NAD+.Maize has two versions of ADH - ADH1 and ADH2, Arabidopsis thaliana contains only one ADH gene. The structure of Arabidopsis ADH is 47% conserved, relative to ADH from horse liver, structurally and functionally important residues, such as the seven residues that provide ligands for the catalytic and noncatalytic zinc atoms, however are conserved suggesting that the enzymes have a similar structure.[22] ADH is constitutively expressed at low levels in the roots of young plants grown on agar, if the roots lack oxygen, the expression of ADH increases significantly.[23] Its expression is also increased in response to dehydration, low temperatures and to abscisic acid and it plays an important role in fruit ripening, seedling and pollen development.[24] Differences in the sequences of ADH in different species have been used to create phylogenies showing how closely related different species of plants are.[25] It is an ideal gene to use due to its convenient size (2–3 kb in length with a ~1000 nucleotide coding sequence) and low copy number.[24]
Iron-containing
Iron-containing alcohol dehydrogenase 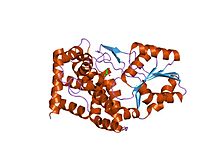
bacillus stearothermophilus glycerol dehydrogenase complex with glycerol Identifiers Symbol Fe-ADH Pfam PF00465 Pfam clan CL0224 InterPro IPR001670 PROSITE PDOC00059 SCOP 1jqa Available protein structures: Pfam structures PDB RCSB PDB; PDBe PDBsum structure summary A third family of alcohol dehydrogenases, unrelated to the above two, are iron-containing ones. They occur in bacteria and fungi. In comparison to enzymes the above families, these enzymes are oxygen-sensitive.[citation needed] Members if the iron-containing alcohol dehydrogenase family include:
- Saccharomyces cerevisiaeAlcohol dehydrogenase 4 (gene ADH4)[26]
- Zymomonas mobilis alcohol dehydrogenase 2 (gene adhB)[27]
- Escherichia coli propanediol oxidoreductase EC 1.1.1.77 (gene fucO),[28] an enzyme involved in the metabolism of fucose and which also seems to contain ferrous ion(s).
- Clostridium acetobutylicum NADPH- and NADH-dependent butanol dehydrogenases EC 1.1.1.- (genes adh1, bdhA and bdhB),[29] enzymes which have activity using butanol and ethanol as substrates.
- E. coli adhE,[30] an iron-dependent enzyme which harbours three different activities: alcohol dehydrogenase, acetaldehyde dehydrogenase (acetylating) EC 1.2.1.10 and pyruvate-formate-lyase deactivase.
- Bacterial glycerol dehydrogenase EC 1.1.1.6 (gene gldA or dhaD).[31]
- Clostridium kluyveri NAD-dependent 4-hydroxybutyrate dehydrogenase (4hbd) EC 1.1.1.61
- Citrobacter freundii and Klebsiella pneumoniae 1,3-propanediol dehydrogenase EC 1.1.1.202 (gene dhaT)
- Bacillus methanolicus NAD-dependent methanol dehydrogenase EC 1.1.1.244[32]
- E. coli and Salmonella typhimurium ethanolamine utilization protein eutG.
- E. coli hypothetical protein yiaY.
Other types
A further class of alcohol dehydrogenases belongs to quinoenzymes and requires quinoid cofactors (e.g., pyrroloquinoline quinone, PQQ) as enzyme-bound electron acceptors. A typical example for this type of enzyme is methanol dehydrogenase of methylotrophic bacteria.
Applications
In fuel cells, alcohol dehydrogenases can be used to catalyze the breakdown of fuel for an ethanol fuel cell. Scientists at Saint Louis University have used carbon-supported alcohol dehydrogenase with poly(methylene green) as an anode, with a nafion membrane, to achieve about 50 μA/cm2.[33]
In biotransformation, alcohol dehydrogenases are often used for the synthesis of enantiomerically pure stereoisomers of chiral alcohols. In contrast to the chemical process, the enzymes yield directly the desired enatiomer of the alcohol by reduction of the corresponding ketone.
Clinical significance
Alcoholism
There have been studies showing that ADH may have an influence on the dependence on ethanol metabolism in alcoholics. Researchers have tentatively detected a few genes to be associated with alcoholism. If the variants of these genes encode slower metabolizing forms of ADH2 and ADH3, there is increased risk of alcoholism. The studies have found that mutations of ADH2 and ADH3 are related to alcoholism in Asian populations. However, research continues in order to identify the genes and their influence on alcoholism.[34]
Drug dependence
Drug dependence is another problem associated with ADH, which researchers think might be linked to alcoholism. One particular study suggests that drug dependence has seven ADH genes associated with it. These results may lead to treatments that target these specific genes. However, more research is necessary.[35]
See also
- Aldehyde dehydrogenase
- Oxidoreductase
- Blood alcohol content for rates of metabolism
References
- ^ PDB 1m6h; Sanghani PC, Robinson H, Bosron WF, Hurley TD (September 2002). "Human glutathione-dependent formaldehyde dehydrogenase. Structures of apo, binary, and inhibitory ternary complexes". Biochemistry 41 (35): 10778–86. doi:10.1021/bi0257639. PMID 12196016.
- ^ a b Danielsson O, Jörnvall H (October 1992). ""Enzymogenesis": classical liver alcohol dehydrogenase origin from the glutathione-dependent formaldehyde dehydrogenase line". Proc. Natl. Acad. Sci. U.S.A. 89 (19): 9247–51. doi:10.1073/pnas.89.19.9247. PMC 50103. PMID 1409630. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=50103.
- ^ a b Persson B, Hedlund J, Jörnvall H (December 2008). "Medium- and short-chain dehydrogenase/reductase gene and protein families : the MDR superfamily". Cell. Mol. Life Sci. 65 (24): 3879–94. doi:10.1007/s00018-008-8587-z. PMC 2792335. PMID 19011751. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2792335.
- ^ Staab CA, Hellgren M, Höög JO (December 2008). "Medium- and short-chain dehydrogenase/reductase gene and protein families : Dual functions of alcohol dehydrogenase 3: implications with focus on formaldehyde dehydrogenase and S-nitrosoglutathione reductase activities". Cell. Mol. Life Sci. 65 (24): 3950–60. doi:10.1007/s00018-008-8592-2. PMID 19011746.
- ^ Godoy L, Gonzàlez-Duarte R, Albalat R (2006). "S-Nitrosogluthathione reductase activity of amphioxus ADH-3: insights into the nitric oxide metabolism". Int. J. Biol. Sci. 2 (3): 117–24. PMC 1458435. PMID 16763671. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1458435.
- ^ Negelein E, Wulff HJ (1937). Biochem. Z. 293: 351.
- ^ Theorell H, McKee JS (October 1961). "Mechanism of action of liver alcohol dehydrogenase". Nature 192 (4797): 47–50. doi:10.1038/192047a0. PMID 13920552.
- ^ Jörnvall H, Harris JI (April 1970). "Horse liver alcohol dehydrogenase. On the primary structure of the ethanol-active isoenzyme". Eur. J. Biochem. 13 (3): 565–76. doi:10.1111/j.1432-1033.1970.tb00962.x. PMID 5462776.
- ^ Brändén CI, Eklund H, Nordström B, Boiwe T, Söderlund G, Zeppezauer E, Ohlsson I, Akeson A (August 1973). "Structure of liver alcohol dehydrogenase at 2.9-angstrom resolution". Proc. Natl. Acad. Sci. U.S.A. 70 (8): 2439–42. doi:10.1073/pnas.70.8.2439. PMC 433752. PMID 4365379. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=433752.
- ^ Hellgren M (2009). Enzymatic studies of alcohol dehydrogenase by a combination of in vitro and in silico methods, Ph.D. thesis. Stockholm, Sweden: Karolinska Institutet. p. 70. ISBN 978-91-7409-567-8. http://diss.kib.ki.se/2009/978-91-7409-567-8/thesis.pdf.
- ^ a b Sofer W, Martin PF (1987). "Analysis of alcohol dehydrogenase gene expression in Drosophila". Annual Review of Genetics 21: 203–25. doi:10.1146/annurev.ge.21.120187.001223. PMID 3327463.
- ^ a b c d Hammes-Schiffer S, Benkovic SJ (2006). "Relating protein motion to catalysis". Annual Review of Biochemistry 75: 519–41. doi:10.1146/annurev.biochem.75.103004.142800. PMID 16756501.
- ^ Erik G. Brandt, Mikko Hellgren, Tore Brinck, Tomas Bergman and Olle Edholm (2009). "Molecular dynamics study of zinc binding to cysteines in a peptide mimic of the alcohol dehydrogenase structural zinc site". Phys. Chem. Chem. Phys. (PCCP) 11 (6): 975–83. doi:10.1039/b815482a. PMID 19177216.
- ^ Sultatos LG, Pastino GM, Rosenfeld CA, Flynn EJ (March 2004). "Incorporation of the genetic control of alcohol dehydrogenase into a physiologically based pharmacokinetic model for ethanol in humans". Toxicological Sciences : an Official Journal of the Society of Toxicology 78 (1): 20–31. doi:10.1093/toxsci/kfh057. PMID 14718645.
- ^ Farrés, james; Moreno, Alberto; Crosas, Bernat; Peralba, Josep M; Allali-Hassani, Abdellah; Hjelmqvist, Lars; Jörnvall, Hans; Parés, Xavier (1994). "Alcohol Dehydrogenase of Class IV (σσ-ADH) from Human Stomach cDNA Sequence and Structure/Function Relationships". European Journal of Biochemistry 224 (2): 549–557. doi:10.1111/j.1432-1033.1994.00549.x. PMID 7925371.
- ^ Kovacs B, Stöppler MC. "Alcohol and Nutrition". MedicineNet, Inc.. http://www.medicinenet.com/alcohol_and_nutrition/article.htm. Retrieved 2011-06-07.
- ^ Duester G (September 2008). "Retinoic acid synthesis and signaling during early organogenesis". Cell 134 (6): 921–31. doi:10.1016/j.cell.2008.09.002. PMC 2632951. PMID 18805086. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2632951.
- ^ Parlesak A, Billinger MH, Bode C, Bode JC (2002). "Gastric alcohol dehydrogenase activity in man: influence of gender, age, alcohol consumption and smoking in a caucasian population". Alcohol and Alcoholism (Oxford, Oxfordshire) 37 (4): 388–93. doi:10.1093/alcalc/37.4.388. PMID 12107043.
- ^ Cox, Michael; Nelson, David R.; Lehninger, Albert L (2005). Lehninger principles of biochemistry. San Francisco: W.H. Freeman. p. 180. ISBN 0-7167-4339-6.
- ^ Leskovac, V.; Trivic, S.; Peričin, D. (December 2002). "The three zinc-containing alcohol dehydrogenases from baker's yeast, Saccharomyces cerevisiae". FEMS Yeast Research 2 (4): 481–494. doi:10.1111/j.1567-1364.2002.tb00116.x. PMID 12702265.
- ^ Coghlan A (2006-12-23). "Festive special: The brewer's tale - life". New Scientist. http://www.newscientist.com/channel/life/mg19225831.100-festive-special-the-brewers-tale.html. Retrieved 2009-04-27.
- ^ C Chang and E M Meyerowitz (1986-03-01). "Molecular cloning and DNA sequence of the Arabidopsis thaliana alcohol dehydrogenase gene". Proceedings of the National Academy of Sciences of the United States of America 83 (5): 1408–12. doi:10.1073/pnas.83.5.1408. PMC 323085. PMID 2937058. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=323085.
- ^ Chung, Hwa-Jee; Robert J. Ferl (October). "Arabidopsis Alcohol Dehydrogenase Expression in Both Shoots and Roots Is Conditioned by Root Growth Environment". Plant Physiology 121 (2): 429–436. doi:10.1104/pp.121.2.429. PMC 59405. PMID 10517834. http://www.plantphysiol.org/cgi/content/full/121/2/429.
- ^ a b Thompson, C.; Fernandes, C.; De Souza, O.; De Freitas, L.; Salzano, F. (2010). "Evaluation of the impact of functional diversification on Poaceae, Brassicaceae, Fabaceae, and Pinaceae alcohol dehydrogenase enzymes". Journal of molecular modeling 16 (5): 919–928. doi:10.1007/s00894-009-0576-0. PMID 19834749.
- ^ Jarvinen, P.; Palme, A.; Orlando Morales, L.; Lannenpaa, M.; Keinanen, M.; Sopanen, T.; Lascoux, M.. "Phylogenetic relationships of Betula species (Betulaceae) based on nuclear ADH and chloroplast matK sequences - Järvinen et al. 91 (11): 1834 - American Journal of Botany". American Journal of Botany (Amjbot.org) 91 (11): 1834. doi:10.3732/ajb.91.11.1834. http://www.amjbot.org/cgi/content/abstract/91/11/1834. Retrieved 2010-07-04.
- ^ Williamson VM, Paquin CE (September 1987). "Homology of Saccharomyces cerevisiae ADH4 to an iron-activated alcohol dehydrogenase from Zymomonas mobilis". Mol. Gen. Genet. 209 (2): 374–81. PMID 2823079.
- ^ Conway T, Sewell GW, Osman YA, Ingram LO (June 1987). "Cloning and sequencing of the alcohol dehydrogenase II gene from Zymomonas mobilis". J. Bacteriol. 169 (6): 2591–7. PMC 212129. PMID 3584063. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=212129.
- ^ Conway T, Ingram LO (July 1989). "Similarity of Escherichia coli propanediol oxidoreductase (fucO product) and an unusual alcohol dehydrogenase from Zymomonas mobilis and Saccharomyces cerevisiae". J. Bacteriol. 171 (7): 3754–9. PMC 210121. PMID 2661535. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=210121.
- ^ Walter KA, Bennett GN, Papoutsakis ET (November 1992). "Molecular characterization of two Clostridium acetobutylicum ATCC 824 butanol dehydrogenase isozyme genes". J. Bacteriol. 174 (22): 7149–58. PMC 207405. PMID 1385386. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=207405.
- ^ Kessler D, Leibrecht I, Knappe J (April 1991). "Pyruvate-formate-lyase-deactivase and acetyl-CoA reductase activities of Escherichia coli reside on a polymeric protein particle encoded by adhE". FEBS Lett. 281 (1-2): 59–63. doi:10.1016/0014-5793(91)80358-A. PMID 2015910.
- ^ Truniger V, Boos W (March 1994). "Mapping and cloning of gldA, the structural gene of the Escherichia coli glycerol dehydrogenase". J. Bacteriol. 176 (6): 1796–800. PMC 205274. PMID 8132480. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=205274.
- ^ de Vries GE, Arfman N, Terpstra P, Dijkhuizen L (August 1992). "Cloning, expression, and sequence analysis of the Bacillus methanolicus C1 methanol dehydrogenase gene". J. Bacteriol. 174 (16): 5346–53. PMC 206372. PMID 1644761. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=206372.
- ^ Moore CM, Minteer SD, Martin RS (February 2005). "Microchip-based ethanol/oxygen biofuel cell". Lab on a Chip 5 (2): 218–25. doi:10.1039/b412719f. PMID 15672138.
- ^ Sher KJ, Grekin ER, Williams NA (2005). "The development of alcohol use disorders". Annual Review of Clinical Psychology 1: 493–523. doi:10.1146/annurev.clinpsy.1.102803.144107. PMID 17716097.
- ^ Luo X, Kranzler HR, Zuo L, Wang S, Schork NJ, Gelernter J (February 2007). "Multiple ADH genes modulate risk for drug dependence in both African- and European-Americans". Human Molecular Genetics 16 (4): 380–90. doi:10.1093/hmg/ddl460. PMC 1853246. PMID 17185388. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1853246.
External links
- PDBsum has links to three-dimensional structures of various alcohol dehydrogenases contained in the Protein Data Bank
- ExPASy contains links to the alcohol dehydrogenase sequences in Swiss-Prot, to a Medline literature search about the enzyme, and to entries in other databases.
- Radio Free Genome created a musical score from a sequence of alcohol dehydrogenase. MP3 audio version and an open source version of the software used to create it is available.
Oxidoreductases: alcohol oxidoreductases (EC 1.1) 1.1.1: NAD/NADP acceptor Alcohol dehydrogenase · Aldo-keto reductase (1A1, 1B1, 1B10, 1C1, 1C3, 1C4, 7A2) · Aldose reductase · Carbohydrate dehydrogenases · Carnitine dehydrogenase · DXP reductoisomerase · Glucose-6-phosphate dehydrogenase · Glycerol-3-phosphate dehydrogenase · HMG-CoA reductase · 3-hydroxyacyl-CoA dehydrogenase · Beta-hydroxybutyryl-CoA dehydrogenase · Isocitrate dehydrogenase · IMP dehydrogenase · Β-Ketoacyl ACP reductase · Lactate dehydrogenase · Malate dehydrogenase · Phosphogluconate dehydrogenase · L-threonine dehydrogenase · L-xylulose reductase · Sorbitol dehydrogenase
Hydroxysteroid dehydrogenase: 3 Beta (3-beta-HSD, NSDHL) · 11 Beta (HSD11B1, HSD11B2) · 17 Beta1.1.2: cytochrome acceptor 1.1.3: oxygen acceptor 1.1.4: disulfide as acceptor 1.1.5: quinone/similar acceptor 1.1.99: other acceptors This article includes text from the public domain Pfam and InterPro IPR001670
Categories:- Genes on chromosome 4
- Protein families
- EC 1.1.1
- NADH-dependent enzymes
- Iron enzymes
- Zinc enzymes
Wikimedia Foundation. 2010.