- Herpes simplex virus
-
This article is about the virus. For information about the disease caused by the virus, see Herpes simplex.
Herpes simplex virus 
TEM micrograph of a herpes simplex virus. Virus classification Group: Group I (dsDNA) Family: Herpesviridae Subfamily: Alphaherpesvirinae Genus: Simplexvirus Species Herpes simplex virus 1 (HSV-1)
Herpes simplex virus 2 (HSV-2)Herpes simplex virus 1 and 2 (HSV-1 and HSV-2), also known as Human herpes virus 1 and 2 (HHV-1 and -2), are two members of the herpes virus family, Herpesviridae, that infect humans.[1] Both HSV-1 (which produces most cold sores) and HSV-2 (which produces most genital herpes) are ubiquitous and contagious. They can be spread when an infected person is producing and shedding the virus.
Symptoms of herpes simplex virus infection include watery blisters in the skin or mucous membranes of the mouth, lips or genitals.[1] Lesions heal with a scab characteristic of herpetic disease. Sometimes, the viruses cause very mild or atypical symptoms during outbreaks. However, as neurotropic and neuroinvasive viruses, HSV-1 and -2 persist in the body by becoming latent and hiding from the immune system in the cell bodies of nerves. After the initial or primary infection, some infected people experience sporadic episodes of viral reactivation or outbreaks. In an outbreak, the virus in a nerve cell becomes active and is transported via the nerve's axon to the skin, where virus replication and shedding occur and cause new sores.[2]
Contents
Transmission
Main article: Herpes simplexHSV-1 and -2 are transmitted from contact with an infectious area of the skin during reactivations of the virus. Although less likely, the herpes viruses can be transmitted during latency. Transmission is likely to occur during symptomatic reactivation of the virus that causes visible and typical skin sores. Asymptomatic reactivation means that the virus causes atypical, subtle or hard to notice symptoms that are not identified as an active herpes infection. Atypical symptoms are often attributed to other causes such as a yeast infection.[3][4] HSV-1 is usually acquired orally during childhood, but may also be sexually transmitted. HSV-2 is primarily a sexually transmitted infection but rates of HSV-1 genital infections are increasing.[3]
Both viruses may also be transmitted vertically during childbirth, although the real risk is very low.[5] The risk of infection is minimal if the mother has no symptoms or exposed blisters during delivery. The risk is considerable when the mother gets the virus for the first time during late pregnancy.[6]
Symptoms resulting from primary infection with HSV are usually much more severe than subsequent outbreaks, as the body has not had a chance to produce antibodies. This first outbreak of oral herpes (cold sores) carries a low (≈1%) risk of developing aseptic meningitis.[1]
Microbiology
Viral structure
Animal herpes viruses all share some common properties. The structure of herpes viruses consists of a relatively large double-stranded, linear DNA genome encased within an icosahedral protein cage called the capsid, which is wrapped in a lipid bilayer called the envelope. The envelope is joined to the capsid by means of a tegument. This complete particle is known as the virion.[7] HSV-1 and HSV-2 each contain at least 74 genes (or open-reading frames, ORFs) within their genomes,[8] although speculation over gene crowding allows as many as 84 unique protein coding genes by 94 putative ORFs.[9] These genes encode a variety of proteins involved in forming the capsid, tegument and envelope of the virus, as well as controlling the replication and infectivity of the virus. These genes and their functions are summarized in the table below.
The genomes of HSV1 and HSV2 are complex and contain two unique regions called the long unique region (UL) and the short unique region (US). Of the 74 known ORFs, UL contains 56 viral genes, whereas US contains only 12.[8] Transcription of HSV genes is catalyzed by RNA polymerase II of the infected host.[8] Immediate early genes, which encode proteins that regulate the expression of early and late viral genes, are the first to be expressed following infection. Early gene expression follows, to allow the synthesis of enzymes involved in DNA replication and the production of certain envelope glycoproteins. Expression of late genes occurs last; this group of genes predominantly encode proteins that form the virion particle.[8]
Five proteins from (UL) form the viral capsid; UL6, UL18, UL35, UL38 and the major capsid protein UL19.[7]
Cellular entry
Entry of HSV into the host cell involves interactions of several glycoproteins on the surface of the enveloped virus, with receptors on the surface of the host cell. The envelope covering the virus particle, when bound to specific receptors on the cell surface, will fuse with the host cell membrane and create an opening, or pore, through which the virus enters the host cell.
The sequential stages of HSV entry are analogous to those of other viruses. At first, complementary receptors on the virus and the cell surface bring the viral and cell membranes into proximity. In an intermediate state, the two membranes begin to merge, forming a hemifusion state. Finally, a stable entry pore is formed through which the viral envelope contents are introduced to the host cell.[10] In the case of a herpes virus, initial interactions occur when a viral envelope glycoprotein called glycoprotein C (gC) binds to a cell surface particle called heparan sulfate. A second glycoprotein, glycoprotein D (gD), binds specifically to at least one of three known entry receptors. These include herpesvirus entry mediator(HVEM), nectin-1 and 3-O sulfated heparan sulfate. The receptor provides a strong, fixed attachment to the host cell. These interactions bring the membrane surfaces into mutual proximity and allow for other glycoproteins embedded in the viral envelope to interact with other cell surface molecules. Once bound to the HVEM, gD changes its conformation and interacts with viral glycoproteins H (gH) and L (gL), which form a complex. The interaction of these membrane proteins results in the hemifusion state. Afterward, gB interaction with the gH/gL complex creates an entry pore for the viral capsid.[10] Glycoprotein B interacts with glycosaminoglycans on the surface of the host cell.
Genetic inoculation
After the viral capsid enters the cellular cytoplasm, it is transported to the cell nucleus. Once attached to the nucleus at a nuclear entry pore, the capsid ejects its DNA contents via the capsid portal. The capsid portal is formed by twelve copies of portal protein, UL6, arranged as a ring; the proteins contain a leucine zipper sequence of amino acids which allow them to adhere to each other.[11] Each icosahedral capsid contains a single portal, located in one vertex.[12][13] The DNA exits the capsid in a single linear segment.[14]
Immune evasion
HSV evades the immune system through interference with MHC class I presentation of antigen on the cell surface. It achieves this through blockade of the TAP transporter induced by the secretion of ICP-47[15] by HSV. TAP maintains the integrity of the MHC class I molecule before it is transported via the golgi apparatus for recognition by CD8+ CTLs on the cell surface. ICP-47 disrupts this integrity, preventing the capture of cytosolic proteins for CTL recognition and thus evades CTL destruction.
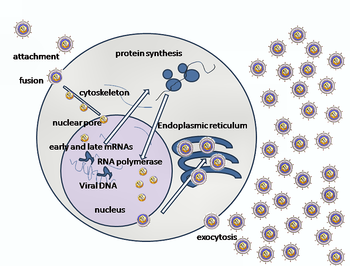
Replication
Following infection of a cell, herpes virus proteins, called immediate-early, early, and late, are produced. Research using flow cytometry on another member of the herpes virus family, Kaposi's sarcoma-associated herpesvirus, indicates the possibility of an additional lytic stage, delayed-late.[16] These stages of lytic infection, particularly late lytic, are distinct from the latency stage. In the case of HSV-1, no protein products are detected during latency, whereas they are detected during the lytic cycle.
The early proteins transcribed are used in the regulation of genetic replication of the virus. On entering the cell, an α-TIF protein joins the viral particle and aids in immediate-early transcription. The virion host shutoff protein (VHS or UL41) is very important to viral replication.[17] This enzyme shuts off protein synthesis in the host, degrades host mRNA, helps in viral replication, and regulates gene expression of viral proteins. The viral genome immediately travels to the nucleus but the VHS protein remains in the cytoplasm.[18][19]
The late proteins are used in to form the capsid and the receptors on the surface of the virus. Packaging of the viral particles — including the genome, core and the capsid - occurs in the nucleus of the cell. Here, concatemers of the viral genome are separated by cleavage and are placed into pre-formed capsids. HSV-1 undergoes a process of primary and secondary envelopment. The primary envelope is acquired by budding into the inner nuclear membrane of the cell. This then fuses with the outer nuclear membrane releasing a naked capsid into the cytoplasm. The virus acquires its final envelope by budding into cytoplasmic vesicles.[20]
Latent infection
HSVs may persist in a quiescent but persistent form known as latent infection, notably in neural ganglia.[1] HSV-1 tends to reside in the trigeminal ganglia, while HSV-2 tends to reside in the sacral ganglia, but note that these are tendencies only, not fixed behavior. During such latent infection of a cell, HSVs express Latency Associated Transcript (LAT) RNA. LAT is known to regulate the host cell genome and interferes with natural cell death mechanisms. By maintaining the host cells, LAT expression preserves a reservoir of the virus, which allows subsequent, usually symptomatic, periodic recurrences or "outbreaks" characteristic of non-latency. Whether or not recurrences are noticeable (symptomatic) or not, viral shedding occurs to produce further infections (usually in a new host, if any). A protein found in neurons may bind to herpes virus DNA and regulate latency. Herpes virus DNA contains a gene for a protein called ICP4, which is an important transactivator of genes associated with lytic infection in HSV-1.[21] Elements surrounding the gene for ICP4 bind a protein known as the human neuronal protein Neuronal Restrictive Silencing Factor (NRSF) or human Repressor Element Silencing Transcription Factor (REST). When bound to the viral DNA elements, histone deacetylation occurs atop the ICP4 gene sequence to prevent initiation of transcription from this gene, thereby preventing transcription of other viral genes involved in the lytic cycle.[21][22] Another HSV protein reverses the inhibition of ICP4 protein synthesis. ICP0 dissociates NRSF from the ICP4 gene and thus prevents silencing of the viral DNA.[23]
The virus can be reactivated by illnesses such as colds and influenza, eczema, emotional and physical stress, gastric upset, fatigue or injury, by menstruation and possibly exposure to bright sunlight.
Viral genome
The open reading frames (ORFs) of HSV-1[8][24] Gene Protein Function/description Gene Protein Function/description UL1 Glycoprotein L [1] Surface and membrane UL38 UL38; VP19C [2] Capsid assembly and DNA maturation UL2 UL2 [3] Uracil-DNA glycosylase UL39 UL39 [4] Ribonucleotide reductase (Large subunit) UL3 UL3 [5] unknown UL40 UL40 [6] Ribonucleotide reductase (Small subunit) UL4 UL4 [7] unknown UL41 UL41; VHS [8] Tegument protein; Virion host shutoff[17] UL5 UL5 [9] DNA replication UL42 UL42 [10] DNA polymerase processivity factor UL6 Portal protein UL-6 Twelve of these proteins constitute the capsid portal ring through which DNA enters and exits the capsid.[11][12][13] UL43 UL43 [11] Membrane protein UL7 UL7 [12] Virion maturation UL44 Glycoprotein C [13] Surface and membrane UL8 UL8 [14] DNA helicase/primase complex-associated protein UL45 UL45 [15] Membrane protein; C-type lectin[25] UL9 UL9 [16] Replication origin-binding protein UL46 VP11/12 [17] Tegument proteins UL10 Glycoprotein M [18] Surface and membrane UL47 UL47; VP13/14 [19] Tegument protein UL11 UL11 [20] virion exit and secondary envelopment UL48 VP16 (Alpha-TIF) [21] Virion maturation; activate IEGs by interacting with the cellular transcription factors Oct-1 and HCF. Binds to the sequence 5'TAATGARAT3'. UL12 UL12 [22] Alkaline exonuclease UL49 UL49A [23] Envelope protein UL13 UL13 [24] Serine-threonine protein kinase UL50 UL50 [25] dUTP diphosphatase UL14 UL14 [26] Tegument protein UL51 UL51 [27] Tegument protein UL15 Terminase [28] Processing and packaging of DNA UL52 UL52 [29] DNA helicase/primase complex protein UL16 UL16 [30] Tegument protein UL53 Glycoprotein K [31] Surface and membrane UL17 UL17 [32] Processing and packaging DNA UL54 IE63; ICP27 [33] Transcriptional regulation UL18 VP23 [34] Capsid protein UL55 UL55 [35] Unknown UL19 VP5 [36] Major capsid protein UL56 UL56 [37] Unknown UL20 UL20 [38] Membrane protein US1 ICP22; IE68 [39] Viral replication UL21 UL21 [40] Tegument protein[26] US2 US2 [41] Unknown UL22 Glycoprotein H [42] Surface and membrane US3 US3 [43] Serine/threonine-protein kinase UL23 Thymidine kinase [44] Peripheral to DNA replication US4 Glycoprotein G [45] Surface and membrane UL24 UL24 [46] unknown US5 Glycoprotein J [47] Surface and membrane UL25 UL25 [48] Processing and packaging DNA US6 Glycoprotein D [49] Surface and membrane UL26 P40; VP24; VP22A [50] Capsid protein US7 Glycoprotein I [51] Surface and membrane UL27 Glycoprotein B [52] Surface and membrane US8 Glycoprotein E [53] Surface and membrane UL28 ICP18.5 [54] Processing and packaging DNA US9 US9 [55] Tegument protein UL29 UL29; ICP8 [56] Major DNA-binding protein US10 US10 [57] Capsid/Tegument protein UL30 DNA polymerase [58] DNA replication US11 US11; Vmw21 [59] Binds DNA and RNA UL31 UL31 [60] Nuclear matrix protein US12 ICP47; IE12 [61] Inhibits MHC class I pathway by preventing binding of antigen to TAP UL32 UL32 [62] Envelope glycoprotein RS1 ICP4; IE175 [63] Major transcriptional activator. Essential for progression beyond the immediate-early phase of infection. IEG transcription repressor. UL33 UL33 [64] Processing and packaging DNA ICP0 ICP0; IE110; α0 [65] E3 ubiquitin ligase that activates viral gene transcription and counteracts the interferon response UL34 UL34 [66] Inner nuclear membrane protein LRP1 LRP1 [67] Latency-related protein UL35 VP26 [68] Capsid protein LRP2 LRP2 [69] Latency-related protein UL36 UL36 [70] Large tegument protein RL1 RL1; ICP34.5 [71] Neurovirulence factor. Antagonizes PKR by de-phosphorylating eIF4a. UL37 UL37 [72] Capsid assembly LAT none [73] Latency-associated transcript Treatment and vaccine development
- For more details on treatment of herpes simplex virus, see Herpes simplex.
- For more information on vaccines, see Herpes simplex vaccine
Herpes viruses establish lifelong infections and the virus cannot currently be eradicated from the body. Treatment usually involves general-purpose antiviral drugs that interfere with viral replication, reducing the physical severity of outbreak-associated lesions and lowering the chance of transmission to others. Studies of vulnerable patient populations have indicated that daily use of antivirals such as acyclovir and valacyclovir can reduce reactivation rates.[4]
In vitro research has indicated that Aloe Vera may be effective against genital herpes.[27]
Connection between facial sores and Alzheimer's disease
In the presence of a certain gene variation (APOE-epsilon4 allele carriers), a possible link between HSV-1 (i.e., the virus that causes cold sores or oral herpes) and Alzheimer’s disease was reported in 1979.[28] HSV-1 appears to be particularly damaging to the nervous system and increases one’s risk of developing Alzheimer’s disease. The virus interacts with the components and receptors of lipoproteins, which may lead to the development of Alzheimer's disease.[29] This research identifies HSVs as the pathogen most clearly linked to the establishment of Alzheimer’s.[30] Without the presence of the gene allele, HSV-1 does not appear to cause any neurological damage or increase the risk of Alzheimer’s.[31] Many more Alzheimer's disease susceptibility genes, including the major players APOE, clusterin, complement receptor 1 and PICALM are involved in the herpes simplex life cycle as curated in this database
References
- ^ a b c d Ryan KJ, Ray CG (editors) (2004). Sherris Medical Microbiology (4th ed.). McGraw Hill. pp. 555–62. ISBN 0838585299.
- ^ "Herpes simplex". DermNet NZ — New Zealand Dermatological Society. 2006-09-16. http://www.dermnetnz.org/viral/herpes-simplex.html. Retrieved 2006-10-15.
- ^ a b Gupta R, Warren T, Wald A (2007). "Genital herpes". Lancet 370 (9605): 2127–37. doi:10.1016/S0140-6736(07)61908-4. PMID 18156035.
- ^ a b Koelle DM, Corey L (2008). "Herpes simplex: insights on pathogenesis and possible vaccines". Annual Review of Medicine 59: 381–95. doi:10.1146/annurev.med.59.061606.095540. PMID 18186706.
- ^ Corey L, Wald A (2009). "Maternal and Neonatal HSV Infections". New England Journal of Medicine 361 (14): 1376–85. doi:10.1056/NEJMra0807633. PMC 2780322. PMID 19797284. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2780322.
- ^ Kimberlin DW (2007). "Herpes simplex virus infections of the newborn". Semin. Perinatol. 31 (1): 19–25. doi:10.1053/j.semperi.2007.01.003. PMID 17317423.
- ^ a b Mettenleiter TC, Klupp BG, Granzow H (2006). "Herpesvirus assembly: a tale of two membranes". Curr. Opin. Microbiol. 9 (4): 423–9. doi:10.1016/j.mib.2006.06.013. PMID 16814597.
- ^ a b c d e McGeoch DJ, Rixon FJ, Davison AJ (2006). "Topics in herpesvirus genomics and evolution". Virus Res. 117 (1): 90–104. doi:10.1016/j.virusres.2006.01.002. PMID 16490275.
- ^ Rajcáni J, Andrea V, Ingeborg R (2004). "Peculiarities of herpes simplex virus (HSV) transcription: an overview". Virus Genes 28 (3): 293–310. doi:10.1023/B:VIRU.0000025777.62826.92. PMID 15266111.
- ^ a b Subramanian RP, Geraghty RJ (2007). "Herpes simplex virus type 1 mediates fusion through a hemifusion intermediate by sequential activity of glycoproteins D, H, L, and B". Proc. Natl. Acad. Sci. U.S.A. 104 (8): 2903–8. doi:10.1073/pnas.0608374104. PMC 1815279. PMID 17299053. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1815279.
- ^ a b Cardone G, Winkler DC, Trus BL, Cheng N, Heuser JE, Newcomb WW, Brown JC, Steven AC (May 2007). "Visualization of the Herpes Simplex Virus Portal in situ by Cryo-electron Tomography". Virology 361 (2): 426–34. doi:10.1016/j.virol.2006.10.047. PMC 1930166. PMID 17188319. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1930166.
- ^ a b Trus BL, Cheng N, Newcomb WW, Homa FL, Brown JC, Steven AC (November 2004). "Structure and Polymorphism of the UL6 Portal Protein of Herpes Simplex Virus Type 1". Journal of Virology 78 (22): 12668–71. doi:10.1128/JVI.78.22.12668-12671.2004. PMC 525097. PMID 15507654. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=525097.
- ^ a b Nellissery JK, Szczepaniak R, Lamberti C, Weller SK (2007-06-20). "A Putative Leucine Zipper within the Herpes Simplex Virus Type 1 UL6 Protein Is Required for Portal Ring Formation". Journal Virology 81 (17): 8868–77. doi:10.1128/JVI.00739-07. PMC 1951442. PMID 17581990. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1951442.
- ^ Newcomb WW, Booy FP, Brown JC (2007). "Uncoating the Herpes Simplex Virus Genome". J. Mol. Biol. 370 (4): 633–42. doi:10.1016/j.jmb.2007.05.023. PMC 1975772. PMID 17540405. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1975772.
- ^ Abbas et al (2009) Cellular and Molecular Immunology, Elsevier Inc.
- ^ Adang LA, Parsons CH, Kedes DH (2006). "Asynchronous Progression through the Lytic Cascade and Variations in Intracellular Viral Loads Revealed by High-Throughput Single-Cell Analysis of Kaposi's Sarcoma-Associated Herpesvirus Infection". J. Virol. 80 (20): 10073–82. doi:10.1128/JVI.01156-06. PMC 1617294. PMID 17005685. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1617294.
- ^ a b Matis J, Kúdelová M (2001). "Early shutoff of host protein synthesis in cells infected with herpes simplex viruses". Acta Virol. 45 (5–6): 269–77. PMID 12083325.
- ^ Taddeo B, Roizman B (2006). "The Virion Host Shutoff Protein (UL41) of Herpes Simplex Virus 1 Is an Endoribonuclease with a Substrate Specificity Similar to That of RNase A". J. Virol. 80 (18): 9341–5. doi:10.1128/JVI.01008-06. PMC 1563938. PMID 16940547. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1563938.
- ^ Skepper JN, Whiteley A, Browne H, Minson A (June 2001). "Herpes Simplex Virus Nucleocapsids Mature to Progeny Virions by an Envelopment → Deenvelopment → Reenvelopment Pathway". J. Virol. 75 (12): 5697–702. doi:10.1128/JVI.75.12.5697-5702.2001. PMC 114284. PMID 11356979. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=114284.
- ^ Granzow H, Klupp BG, Fuchs W, Veits J, Osterrieder N, Mettenleiter TC (April 2001). "Egress of Alphaherpesviruses: Comparative Ultrastructural Study". J. Virol. 75 (8): 3675–84. doi:10.1128/JVI.75.8.3675-3684.2001. PMC 114859. PMID 11264357. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=114859.
- ^ a b Pinnoji RC, Bedadala GR, George B, Holland TC, Hill JM, Hsia SC (2007). "Repressor element-1 silencing transcription factor/neuronal restrictive silencer factor (REST/NRSF) can regulate HSV-1 immediate-early transcription via histone modification". Virol. J. 4: 56. doi:10.1186/1743-422X-4-56. PMC 1906746. PMID 17555596. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1906746.
- ^ Bedadala GR, Pinnoji RC, Hsia SC (2007). "Early growth response gene 1 (Egr-1) regulates HSV-1 ICP4 and ICP22 gene expression". Cell Res. 17 (6): 546–55. doi:10.1038/cr.2007.44. PMID 17502875.
- ^ Roizman B, Gu H, Mandel G (2005). "The first 30 minutes in the life of a virus: unREST in the nucleus". Cell Cycle 4 (8): 1019–21. doi:10.4161/cc.4.8.1902. PMID 16082207.
- ^ Search in UniProt Knowledgebase (Swiss-Prot and TrEMBL) for: HHV1
- ^ Wyrwicz LS, Ginalski K, Rychlewski L (2007). "HSV-1 UL45 encodes a carbohydrate binding C-type lectin protein". Cell Cycle 7 (2): 269–71. doi:10.4161/cc.7.2.5324. PMID 18256535.
- ^ Vittone V, Diefenbach E, Triffett D, Douglas MW, Cunningham AL, Diefenbach RJ (2005). "Determination of Interactions between Tegument Proteins of Herpes Simplex Virus Type 1". J. Virol. 79 (15): 9566–71. doi:10.1128/JVI.79.15.9566-9571.2005. PMC 1181608. PMID 16014918. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1181608.
- ^ Vogler BK, Ernst E (October 1999). "Aloe vera: a systematic review of its clinical effectiveness". The British Journal of General Practice 49 (447): 823–8. PMC 1313538. PMID 10885091. http://openurl.ingenta.com/content/nlm?genre=article&issn=0960-1643&volume=49&issue=447&spage=823&aulast=Vogler.
- ^ Middleton PJ, Petric M, Kozak M, Rewcastle NB, McLachlan DR (May 1980). "Herpes-simplex viral genome and senile and presenile dementias of Alzheimer and Pick". Lancet 315 (8176): 1038. doi:10.1016/S0140-6736(80)91490-7. PMID 6103379.
- ^ Dobson CB, Itzhaki RF (1999). "Herpes simplex virus type 1 and Alzheimer's disease". Neurobiol. Aging 20 (4): 457–65. doi:10.1016/S0197-4580(99)00055-X. PMID 10604441.
- ^ Pyles RB (November 2001). "The association of herpes simplex virus and Alzheimer's disease: a potential synthesis of genetic and environmental factors" (PDF). Herpes 8 (3): 64–8. PMID 11867022. http://www.ihmf.com/journal/download/83pyles(64)vol864.pdf.
- ^ Itzhaki RF, Lin WR, Shang D, Wilcock GK, Faragher B, Jamieson GA (January 1997). "Herpes simplex virus type 1 in brain and risk of Alzheimer's disease". Lancet 349 (9047): 241–4. doi:10.1016/S0140-6736(96)10149-5. PMID 9014911.
External links
- Genital Herpes - Public Health Agency of Canada
- Herpes simplex: Host viral protein interactions: A database of HSV-1 interacting host proteins
Sexually transmitted diseases and infections (STD/STI) (primarily A50–A64, 090–099) Bacterial Protozoal Parasitic Viral AIDS (HIV-1/HIV-2) · Cervical cancer, vulvar cancer & Genital warts (condyloma), Penile cancer, Anal cancer (Human papillomavirus (HPV)) · Hepatitis B (Hepatitis B virus) · Herpes simplex (HSV1/HSV2) · Molluscum contagiosum (MCV)General
inflammationInfectious skin disease: Viral cutaneous conditions, including viral exanthema (B00–B09, 050–059) Ungrouped unknown/multiple: Asymmetric periflexural exanthem of childhood · Post-vaccination follicular eruption · Lipschütz ulcer · Eruptive pseudoangiomatosis · Viral-associated trichodysplasia · Gianotti–Crosti syndromeBaltimore (virus classification) DNA I: dsDNA viruses HerpesviralesUnassignedNLCDV: Asfarviridae · Iridoviridae · Marseilleviridae · Megaviridae · Mimiviridae · Phycodnaviridae · Poxviridae
nonenveloped: Adenoviridae · Papillomaviridae · Papovaviridae (obsolete) · Polyomaviridae
Ascoviridae · Baculoviridae · Corticoviridae · Fuselloviridae · Guttaviridae · Lipothrixviridae · Nimaviridae · Plasmaviridae · Rudiviridae · TectiviridaeII: ssDNA viruses non-enveloped: Parvoviridae (Parvovirus B19)
ungrouped: Circoviridae · Geminiviridae · Inoviridae · Microviridae · NanoviridaeRNA III: dsRNA viruses Birnaviridae · Chrysoviridae · Cystoviridae · Hypoviridae · Partitiviridae · Reoviridae (Rotavirus) · TotiviridaeIV: (+)ssRNA viruses (primarily icosahedral) PicornaviralesDicistroviridae · Iflaviridae · Marnaviridae · Picornaviridae (Enterovirus, Rhinovirus) · SecoviridaeTymoviralesUnassignedAstroviridae · Barnaviridae · Bromoviridae · Caliciviridae · Closteroviridae · Comoviridae · Flaviviridae · Flexiviridae · Leviviridae · Luteoviridae · Narnaviridae · Nodaviridae · Potyviridae · Sequiviridae · Tetraviridae · Togaviridae (Rubella virus) · TombusviridaeV: (-)ssRNA viruses (primarily helical) RT VI: ssRNA-RT viruses VII: dsDNA-RT viruses Categories:- Herpesviruses
- Sexually transmitted diseases and infections
Wikimedia Foundation. 2010.