- Coronaviridae
-
Coronaviridae Virus classification Group: Group IV ((+)ssRNA) Order: Nidovirales Family: Coronaviridae Subfamilies Coronavirinae
TorovirinaeCoronaviruses are enveloped, single-stranded, positive-sense RNA viruses (27-31kb) with club-shaped surface about 120-160 nm in diameter that resemble a “corona”.
Contents
Virology
The 5' and 3' ends of the genome have a cap and poly (A) tract, respectively. The envelope consists of two viral-specified structural glycoproteins S and M. Glycoprotein S is located on the outer membrane of the envelope and is responsible for the club-shaped projections. On the other hand, glycoprotein M is a transmembrane molecule and is located on the inner part of the envelope. Another important structural protein is the phosphoprotein N, which is responsible for the helical symmetry of the nucleocapsid that encloses the genomic RNA.
Taxonomy
The Coronaviridae are a family of viruses, including the following subfamilies: [1]
- Subfamily: Coronavirinae
- Genera: Alphacoronavirus, Betacoronavirus, Gammacoronavirus
- Genera: Alphacoronavirus, Betacoronavirus, Gammacoronavirus
- Subfamily: Torovirinae
- Genera: Bafinivirus, Torovirus
In April of 2008, the following proposals were ratified by the ICTV: [2]
- 2005.260V.04 To create the following species in the genus Coronavirus in the family Coronaviridae, named Goose coronavirus, Pigeon coronavirus, Duck coronavirus.
- 2006.009V.04 To create a species in the genus Coronavirus in the family Coronaviridae, named Human coronavirus NL63.
- 2006.010V.04 To create a species in the genus Coronavirus in the family Coronaviridae, named Human coronavirus HKU1.
- 2006.011V.04 To create a species in the genus Coronavirus in the family Coronaviridae, namedEquine coronavirus.
In July of 2009, the following proposals were ratified by the ICTV: [3]
- 2008.085-122V.A.v3.Coronaviridae
- 2008.085V Create a new subfamily in the family Coronaviridae, order Nidovirales
- 2008.086V Name the new subfamily Coronavirinae
- 2008.087V Create a new genus in the proposed subfamily Coronavirinae
- 2008.088V Name the new genus Alphacoronavirus
- 2008.089V Assign three existing species (Human coronavirus 229E, Human coronavirus NL63, Porcine epidemic diarrhea virus) and five new species proposed in 2008.091-095V.01 to the proposed new genus Alphacoronavirus
- 2008.090V Designate proposed species Alphacoronavirus 1 as type species of the genus Alphacoronavirus
- 2008.091V Create new species named Alphacoronavirus 1 in the new genus
- 2008.092V Create new species named Rhinolophus bat coronavirus HKU2 in the new genus
- 2008.093V Create new species named Scotophilus bat coronavirus 512 in the new genus
- 2008.094V Create new species named Miniopterus bat coronavirus 1 in the new genus
- 2008.095V Create new species named Miniopterus bat coronavirus HKU8 in the new genus
- 2008.096V Create a new genus in the proposed subfamily Coronavirinae
- 2008.097V Name the new genus Betacoronavirus
- 2008.098V Assign the existing species Human coronavirus HKU1 and six new species proposed in
- 2008.100-105V.01 to the proposed genus Betacoronavirus
- 2008.099V Designate proposed species Murine coronavirus as type species of the genus Betacoronavirus
- 2008.108V Assign the two species proposed in 2008.110,111V.01 to the new genus
- 2008.109V Designate proposed species Avian coronavirus as type species of the new genus
- 2008.110V Create species named Avian coronavirus in the new genus
- 2008.111V Create species named Beluga whale coronavirus SW1 in the new genus
- 2008.112V Create a new subfamily in the family Coronaviridae, order Nidovirales
- 2008.113V Name the new subfamily Torovirinae
- 2008.114V Create a new genus in the subfamily Torovirinae
- 2008.115V Name the new genus Bafinivirus
- 2008.116V Assign the species White breamVirus (proposed in 2008.118V.01) to the new genus
- 2008.117V Designate species White bream virus as type species in the new genus
- 2008.118V Create species named White bream virus in the new genus
- 2008.119V Remove the genus Torovirus from the family Coronaviridae
- 2008.120V Reassign the genus Torovirus to the subfamily Torovirinae
- 2008.121V.U Remove (abolish) 18 species (Human enteric coronavirus, Human coronavirus OC43, Bovine coronavirus, Porcine hemagglutinating encephalomyelitis virus, Equine coronavirus, Murine hepatitis virus, Puffinosis coronavirus, Rat coronavirus, Transmissible gastroenteritis virus, Canine coronavirus, Feline coronavirus, Infectious bronchitis virus, Duck coronavirus, Goose coronavirus, Pheasant coronavirus, Pigeon coronavirus, Turkey coronavirus, Severe acute respiratory syndrome coronavirus) from the genus Coronavirus
- 2008.122V.U Reassign species Human coronavirus 229E, Human coronavirus NL63 and Porcine epidemic diarrhea virus to the new genus Alphacoronavirus and Human coronavirus HKU1 to the new genus Betacoronavirus
Transmission
Coronaviruses are transmitted by faecal-oral route or by aerosols of respiratory secretions.
Pathogenesis
Coronaviruses infect a wide range of mammals and birds and occur worldwide. Although most diseases are mild, sometimes they can cause more severe situations in humans, such as, for example, the infection of the respiratory tract known as Severe Acute Respiratory Syndrome (SARS). They can also cause enteric infections in very young infants and, in rare situations, neurological syndromes.
SARS
Human infection by SARS coronavirus appears to be limited to the respiratory tract where infection of susceptible cells leads to damage to the pneumocytes resulting in a histological picture of diffuse alveolar damage and a clinical picture of adult respiratory distress syndrome. Diarrhea is also present but there is limited evidence of damage to the intestinal epithelium. The damage to the respiratory tree appears limited to the lower respiratory tract and there is evidence that the immune response plays a part in the outcome of patients with SARS.[4]
Binding and Entry
Coronaviruses bind to host cells primarily through interactions between viral spike glycoproteins and specific host cell surface glycoproteins. Some coronaviruses also bind to sialic acids on glycoproteins and glycolipids via their spike and/or hemaglutinin esterase glycoproteins. The interactions between coronaviruses and host cell receptors are critical determinants of species-specificity, tissue tropism, and virulence.[4]
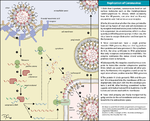
Replication
Coronaviruses have single-stranded, positive-sense RNA genomes of about 30 kilobases, by far the largest non-segmented RNA virus genomes currently known. The key functions required for coronavirus RNA synthesis are encoded by the viral replicase gene. The gene comprises more than 20,000 nucleotides and encodes two replicase polyproteins, pp1a and pp1ab, that are proteolytically processed by viral proteases. Over the past years, it has become clear that the unique size of the coronavirus genome and the special mechanism that coronaviruses (and several other nidoviruses) have evolved to produce an extensive set of subgenome-length RNAs is linked to the production of a number of nonstructural proteins (nsps) that is unprecedented among RNA viruses. Many of these replicase cleavage products are in fact multidomain proteins themselves, thus further increasing the complexity of protein functions and interactions. Structural studies suggest that several nsps, following their release from larger precursor molecules, form dimers or even multimers. The various pp1a/pp1ab precursors and processing products are thought to assemble into large, membrane-associated complexes that, in a temporally coordinated manner, catalyze the reactions involved in RNA replication and transcription and, it is presumed, serve yet other functions in the viral life cycle.[4][5]
Coronaviruses also exhibit ribosomal frameshifting and polymerase stuttering as part of their complex replicative cycle.[6][7][8][9][10]
Genomic Cis-Acting Elements
In common with the genomes of all other RNA viruses, coronavirus genomes contain cis-acting RNA elements that ensure the specific replication of viral RNA by a virally encoded RNA-dependent RNA polymerase. The embedded cis-acting elements devoted to coronavirus replication constitute a small fraction of the total genome, but this is, it is presumed, a reflection of the fact that coronaviruses have the largest genomes of all RNA viruses. The boundaries of cis-acting elements essential to replication are fairly well-defined, and an increasingly well-resolved picture of the RNA secondary structures of these regions is emerging. However, we are in only the early stages of understanding how these cis-acting structures and sequences interact with the viral replicase and host cell components, and much remains to be done before we understand the precise mechanistic roles of such elements in RNA synthesis.[4]
Genome Packaging
The assembly of infectious coronavirus particles requires the selection of viral genomic RNA from a cellular pool that contains an abundant excess of non-viral and viral RNAs. Among the seven to ten specific viral mRNAs synthesized in virus-infected cells, only the full-length genomic RNA is packaged efficiently into coronavirus particles. Studies have revealed cis-acting elements and trans-acting viral factors involved in coronavirus genome encapsidation and packaging. Understanding the molecular mechanisms of genome selection and packaging is critical for developing antiviral strategies and viral expression vectors based on the coronavirus genome.[4]
References
- ^ "ICTV 2009 MASTER SPECIES LIST VERSION 4". 20 March 2010. http://talk.ictvonline.org/cfs-filesystemfile.ashx/__key/CommunityServer.Components.PostAttachments/00.00.00.12.31/ICTV_2D00_Master_2D00_Species_2D00_List_2D00_2009_5F00_v4.xls
- ^ Carstens, E. B.; Ball, L. A. (July, 2009). "Ratification vote on taxonomic proposals to the International Committee on Taxonomy of Viruses (2008)". Archives of Virology (Springer Wien) 154 (7): 1181–1188. doi:10.1007/s00705-009-0400-2. ISSN 1432-8798. PMID 19495937. http://www.springerlink.com/content/n5633x0287h34370/fulltext.pdf.
- ^ Carstens, E. B. (January, 2010). "Ratification vote on taxonomic proposals to the International Committee on Taxonomy of Viruses (2009)". Archives of Virology (Springer Wien) 155 (1): 133–146. doi:10.1007/s00705-009-0547-x. ISSN 1432-8798. PMID 19960211. http://www.springerlink.com/content/55520571803m75n3/fulltext.pdf.
- ^ a b c d e Thiel V (editor). (2007). Coronaviruses: Molecular and Cellular Biology (1st ed.). Caister Academic Press. ISBN 978-1-904455-16-5. [1]. http://www.horizonpress.com/cor.
- ^ Enjuanes L, et al. (2008). "Coronavirus Replication and Interaction with Host". Animal Viruses: Molecular Biology. Caister Academic Press. ISBN [[Special:BookSources/978-1-904455-22-6]|978-1-904455-22-6]]]. [http://www.horizonpress.com/avir. http://www.horizonpress.com/hsp/abs/absavir.html.
- ^ Namy O, Moran SJ, Stuart DI, Gilbert RJ, Brierley I. "A mechanical explanation of RNA pseudoknot function in programmed ribosomal frameshifting." Nature. 2006 May 11;441(7090):244-7.
- ^ Plant EP, Dinman JD. "Comparative study of the effects of heptameric slippery site composition on -1 frameshifting among different eukaryotic systems." RNA. 2006 Apr;12(4):666-73. Epub 2006 Feb 22.
- ^ Su MC, Chang CT, Chu CH, Tsai CH, Chang KY. "An atypical RNA pseudoknot stimulator and an upstream attenuation signal for -1 ribosomal frameshifting of SARS coronavirus." Nucleic Acids Res. 2005 Jul 29;33(13):4265-75.
- ^ Herrewegh AA, Vennema H, Horzinek MC, Rottier PJ, de Groot RJ. "The molecular genetics of feline coronaviruses: comparative sequence analysis of the ORF7a/7b transcription unit of different biotypes." Virology. 1995 Oct 1;212(2):622-31.
- ^ Jeong YS, Makino S. "Evidence for coronavirus discontinuous transcription." J Virol. 1994 Apr;68(4):2615-23.
External links
Categories: - Subfamily: Coronavirinae
Wikimedia Foundation. 2010.