- NAD+ kinase
-
NAD+ kinase 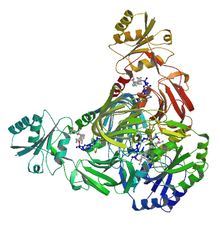
Ribbon diagram of NAD+ kinase in complex with substrates.[1] Identifiers EC number 2.7.1.23 CAS number 9032-66-0 Databases IntEnz IntEnz view BRENDA BRENDA entry ExPASy NiceZyme view KEGG KEGG entry MetaCyc metabolic pathway PRIAM profile PDB structures RCSB PDB PDBe PDBsum Gene Ontology AmiGO / EGO Search PMC articles PubMed articles NAD kinase Identifiers Symbols NADK; DKFZp686L22239; FLJ13052; FLJ37724; FLJ54695; FLJ77769; FLJ78247; FLJ78307; MGC1900; dJ283E3.1 External IDs OMIM: 611616 MGI: 2183149 HomoloGene: 49724 GeneCards: NADK Gene EC number 2.7.1.23 Gene Ontology Molecular function • nucleotide binding
• NAD+ kinase activity
• NAD+ kinase activity
• protein binding
• ATP binding
• transferase activity
• metal ion bindingCellular component • cytosol
• cytosolBiological process • vitamin metabolic process
• water-soluble vitamin metabolic process
• metabolic process
• phosphorylation
• NAD metabolic process
• ATP metabolic processSources: Amigo / QuickGO RNA expression pattern 
More reference expression data Orthologs Species Human Mouse Entrez 65220 192185 Ensembl ENSG00000008130 ENSMUSG00000029063 UniProt O95544 Q3TCP7 RefSeq (mRNA) NM_001198993.1 NM_138671 RefSeq (protein) NP_001185922.1 NP_619612 Location (UCSC) Chr 1:
1.68 – 1.71 MbChr 4:
154.94 – 154.97 MbPubMed search [1] [2] NAD+ kinase (EC 2.7.1.23, NADK) is an enzyme that converts nicotinamide adenine dinucleotide (NAD+) into NADP+, through phosphorylating the NAD+ coenzyme.[2] NADP+ is an essential coenzyme in metabolism and provides reducing power to biosynthetic processes such as fatty acid biosynthesis.[3] The structure of the NADK from the archaean Archaeoglobus fulgidus has been determined.[1]
In humans, NAD+ kinase is encoded by the NADK gene.[4]
Contents
Reaction
See also
References
- ^ a b PDB 1SUW; Liu J, Lou Y, Yokota H, Adams PD, Kim R, Kim SH (November 2005). "Crystal structures of an NAD kinase from Archaeoglobus fulgidus in complex with ATP, NAD, or NADP". J. Mol. Biol. 354 (2): 289–303. doi:10.1016/j.jmb.2005.09.026. PMID 16242716.
- ^ Magni G, Orsomando G, Raffaelli N (2006). "Structural and functional properties of NAD kinase, a key enzyme in NADP biosynthesis". Mini reviews in medicinal chemistry 6 (7): 739–46. doi:10.2174/138955706777698688. PMID 16842123.
- ^ Pollak N, Dölle C, Ziegler M (2007). "The power to reduce: pyridine nucleotides – small molecules with a multitude of functions". Biochem. J. 402 (2): 205–18. doi:10.1042/BJ20061638. PMC 1798440. PMID 17295611. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1798440.
- ^ Lerner F, Niere M, Ludwig A, Ziegler M (October 2001). "Structural and functional characterization of human NAD kinase". Biochem. Biophys. Res. Commun. 288 (1): 69–74. doi:10.1006/bbrc.2001.5735. PMID 11594753.
Further reading
- Andersson B, Wentland MA, Ricafrente JY, et al. (1996). "A "double adaptor" method for improved shotgun library construction". Anal. Biochem. 236 (1): 107–13. doi:10.1006/abio.1996.0138. PMID 8619474.
- Yu W, Andersson B, Worley KC, et al. (1997). "Large-Scale Concatenation cDNA Sequencing". Genome Res. 7 (4): 353–8. doi:10.1101/gr.7.4.353. PMC 139146. PMID 9110174. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=139146.
- Strausberg RL, Feingold EA, Grouse LH, et al. (2003). "Generation and initial analysis of more than 15,000 full-length human and mouse cDNA sequences". Proc. Natl. Acad. Sci. U.S.A. 99 (26): 16899–903. doi:10.1073/pnas.242603899. PMC 139241. PMID 12477932. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=139241.
- Ota T, Suzuki Y, Nishikawa T, et al. (2004). "Complete sequencing and characterization of 21,243 full-length human cDNAs". Nat. Genet. 36 (1): 40–5. doi:10.1038/ng1285. PMID 14702039.
- Gerhard DS, Wagner L, Feingold EA, et al. (2004). "The Status, Quality, and Expansion of the NIH Full-Length cDNA Project: The Mammalian Gene Collection (MGC)". Genome Res. 14 (10B): 2121–7. doi:10.1101/gr.2596504. PMC 528928. PMID 15489334. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=528928.
- Stelzl U, Worm U, Lalowski M, et al. (2005). "A human protein-protein interaction network: a resource for annotating the proteome". Cell 122 (6): 957–68. doi:10.1016/j.cell.2005.08.029. PMID 16169070.
- Rual JF, Venkatesan K, Hao T, et al. (2005). "Towards a proteome-scale map of the human protein-protein interaction network". Nature 437 (7062): 1173–8. doi:10.1038/nature04209. PMID 16189514.
- Gregory SG, Barlow KF, McLay KE, et al. (2006). "The DNA sequence and biological annotation of human chromosome 1". Nature 441 (7091): 315–21. doi:10.1038/nature04727. PMID 16710414.
- Ewing RM, Chu P, Elisma F, et al. (2007). "Large-scale mapping of human protein–protein interactions by mass spectrometry". Mol. Syst. Biol. 3 (1): 89. doi:10.1038/msb4100134. PMC 1847948. PMID 17353931. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1847948.
- Pollak N, Niere M, Ziegler M (2008). "NAD kinase levels control the NADPH concentration in human cells". J. Biol. Chem. 282 (46): 33562–71. doi:10.1074/jbc.M704442200. PMID 17855339.
External links
Transferases: phosphorus-containing groups (EC 2.7) 2.7.1-2.7.4:
phosphotransferase/kinase
(PO4)Hexo- · Gluco- · Fructo- (Hepatic) · Galacto- · Phosphofructo- (1, Liver, Muscle, Platelet, 2) · Riboflavin · Shikimate · Thymidine (ADP-thymidine) · NAD+ · Glycerol · Pantothenate · Mevalonate · Pyruvate · Deoxycytidine · PFP · Diacylglycerol · Phosphoinositide 3 (Class I PI 3, Class II PI 3) · Sphingosine · Glucose-1,6-bisphosphate synthase2.7.2: COOH acceptor2.7.6: diphosphotransferase
(P2O7)2.7.7: nucleotidyltransferase
(PO4-nucleoside)DNA-directed DNA polymerase: DNA polymerase I · DNA polymerase II · DNA polymerase III holoenzyme
DNA nucleotidylexotransferase/Terminal deoxynucleotidyl transferase
RNA-directed DNA polymerase: Reverse transcriptase (Telomerase)RNA nucleotidyltransferaseRNA polymerase/DNA-directed RNA polymerase: RNA polymerase I · RNA polymerase II · RNA polymerase III · RNA polymerase IV · Primase · RNA-dependent RNA polymerase
PNPaseUridylyltransferaseGlucose-1-phosphate uridylyltransferase · Galactose-1-phosphate uridylyltransferaseGuanylyltransferasemRNA capping enzymeOther2.7.8: miscellaneous PhosphatidyltransferasesCDP-diacylglycerol—glycerol-3-phosphate 3-phosphatidyltransferase · CDP-diacylglycerol—serine O-phosphatidyltransferase · CDP-diacylglycerol—inositol 3-phosphatidyltransferase · CDP-diacylglycerol—choline O-phosphatidyltransferaseGlycosyl-1-phosphotransferase2.7.10-2.7.13: protein kinase
(PO4; protein acceptor)see tyrosine kinasessee serine/threonine-specific protein kinases2.7.12: protein-dual-specificitysee serine/threonine-specific protein kinases2.7.13: protein-histidineThis transferase article is a stub. You can help Wikipedia by expanding it. This article about metabolism is a stub. You can help Wikipedia by expanding it.

