- Folding@home
-
Folding@home 
Original author(s) Vijay Pande Developer(s) Stanford University / Pande lab Initial release 2000-10-01 Stable release Windows:
Uniprocessor: 6.23[1]
GPU: 6.41[2]
Mac OS X:
x86-64 SMP: 6.29.3[1]
Linux:
SMP: 6.34[1][2]
PlayStation 3: 1.4[3]Preview release Windows, Mac, and Linux:
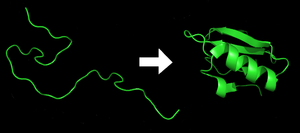
7.1.38[4] / October 10, 2011[5]Operating system Microsoft Windows, Mac OS X, Linux Platform Cross-platform Available in English Type Distributed computing License Proprietary[6] Website folding.stanford.edu Folding@home is a distributed computing project designed to use spare processing power on personal computers to perform simulations of disease-relevant protein folding and other molecular dynamics, and to improve on the methods of doing so. Also referred to as FAH or F@h, much of its work attempts to determine how proteins reach their final structure, which is of significant academic interest and has implications to both disease research and nanotechnology. To a lesser degree Folding@home also tries to predict that final structure from only the initial amino acid sequence, which has applications in drug design.[7] Folding@home is developed and operated by the Pande Laboratory at Stanford University, under the direction of Vijay Pande.[8] The goal of the project is to "understand protein folding, misfolding, and related diseases".[9][10]
Folding@home's accurate simulations of protein folding and misfolding enable the scientific community to better understand the development of many diseases, including Alzheimer's disease, Parkinson's disease, cancer, Creutzfeldt–Jakob disease, Huntington's disease, cystic fibrosis, sickle-cell anaemia, HIV, Chagas disease, influenza, osteogenesis imperfecta, autism,[11] and alpha 1-antitrypsin deficiency, among others.[12] More fundamentally, understanding the process of protein folding — how proteins assemble themselves into a functional state — is one of the outstanding problems of molecular biology.[13] The Pande lab has produced ninety-five scientific research papers as a direct result of the project, and Folding@home has caused paradigm shifts in protein folding theory.[14][15]
In January 2010 the Folding@home project successfully simulated protein folding in the 1.5 millisecond range — a simulation thousands of times longer than ever previously achieved.[16] Folding@home has pioneered the uses of GPUs, multi-core processors, and PlayStation 3s for distributed computing. It remains one of the world's most powerful computing systems, is more powerful than all distributed computing projects under BOINC combined, and is one of the world's largest projects. Folding@home's distributed simulations remain accurate compared to results from laboratory research, a "grand challenge" in computational biology.[17] The Pande lab's goal is to refine Folding@home's methods to the level where it will become an essential tool for molecular medical research, and they collaborate with various scientific institutions and laboratories across the world.[18][19]
Contents
Biomedical significance
Further information: Protein foldingProteins are an essential component to many biological functions, and participate in virtually every process within cells. They often act as enzymes, performing biochemical reactions including cell signaling, transportation, cellular regulation, and others. As structural elements, some proteins act as a type of skeleton for cells, and as antibodies other proteins help the immune system.[12] Before a protein can take on these roles, it must fold into a functional three-dimensional shape based on a particular series of steps, which often occur spontaneously.[20][21] Understanding protein folding is thus critical to understanding what a protein does and how it works.[12] Moreover, when proteins do not fold correctly—that is, when they fold down the wrong pathway and end up misshapen—they can aggregate and cause serious and in some cases life-threatening diseases.[22][23] While the understanding of protein folding requires a combination of theories and experiments, creating experimentally comparable simulations of protein folding dynamics remains difficult and is considered a "holy grail" of computational biology.[24][25]
Folding@home's novel distributed computing methods can address these challenges and allow for better insights into the complexity of protein folding while strongly agreeing with experimental data.[24][26] A highly accurate and thorough exploration of protein configurations in a reasonable amount of time is obtainable through distributed computing, especially over very long simulation timescales.[27] The Pande lab and other researchers can use Folding@home to study aspects of folding, misfolding, and related diseases that would never be seen experimentally; however, many simulations run on Folding@home are used in conjunction with laboratory experiments.[17] Moreover, as of 2011 the Pande lab is performing studies into how protein folding in native cells may be different than in environments such as test tubes used during experiments.[28][29]
While the understanding of protein folding has become increasingly important for disease research, for the last decade simulations of protein molecular dynamics have been severely limited by computational power. Straightforward approaches have exceptional difficulty for all but the most elementary of systems, which has prompted the use of simplified native-centric in silico molecular models.[24][30] Not only are these models insufficient for a comprehensive view of protein folding, but the traditional methods of using a small number of very long simulations cannot accurately capture detailed information into how a protein misfolds.[24][31] Moreover, these simplified models often ignore how solvents such as water might affect the folding process. Folding@home's computational power and distributed computing methods can reveal its influence, especially with explicit solvation simulations, as demonstrated in 2004 in strong agreement with experimental observations.[32] This observation was made possible by Folding@home's assembly of many shorter simulations, which provides a much more complete description of the protein's energy landscape, conformation space, and equilibrium thermodynamics.[18][33] While some of these simulations may work themselves into impossible atomic configurations or into the correct native state, others will illustrate how that protein misfolds.[33] Armed with a knowledge of thermodynamics, Folding@home uses methods that can find these rare events without having to complete many simulations.[34][35] Once the misfolding process is determined, figuring out its prevention can be the next step, and preventative treatments can be applied during the folding process.[23] The Pande lab has used Folding@home to study this for number of years using a variety of different methods.[36]
In addition to the diseases listed below, Folding@home is used to study the dynamics of key components of HIV,[37] details of how the influenza virus recognizes and infects cells,[38] as well as malaria, Chagas disease,[12] and the prions which cause Creutzfeldt–Jakob disease.[39] Folding@home's results have led to major shifts in the understanding of protein folding and its applications for disease, as well as improved protein folding models.[40][41] Folding@home is dedicated to producing significant amounts of results towards protein folding, the diseases that result from protein misfolding, and novel computational methods for doing so.[12] The goal of the first five years of the project was to make significant advances in understanding folding, while the current goal is to understand misfolding and related disease, especially Alzheimer's disease.[17][42]
As a part of Stanford University, a non-profit organization, the Pande lab does not sell the results generated by Folding@home. The large data sets from the project are freely available for other researchers to use upon request, and some can be accessed from the Folding@home website.[43][44] The Pande lab also releases Folding@home's key software to other researchers, so that the algorithms which benefit Folding@home will also help in other scientific areas.[43] Additionally, summaries of all of the scientific findings from Folding@home are posted on the Folding@home website after publication.[14] The full publications are available online or from a local municipal or academic library.[44]
Alzheimer's disease
Alzheimer's disease, a form of dementia which most often affects the elderly, is believed to be caused by specific misfolding and subsequent aggregation of the small 42-residue amyloid beta (Aß) peptide.[42] The severity of the disease depends not only on the amount of Aß, but also on how it misfolds. Current theory holds that toxic non-plaque Aß oligomers (aggregates of many monomers) bind to a surface receptor on neurons and change the structure of the synapse, thereby disrupting neuronal communication and causing neuronal cell death which leads to the associated neurodegenerative consequences.[45][46] Understanding how and why this peptide misfolds could result in key insights into how to cure Alzheimer's Disease, and will also help the Pande lab prepare for similar aggregation studies.[42][47]
Despite this connection, toxic Aß aggregations remain so complex that it was not previously possible to simulate them in atomic resolution. In 2011, the Pande lab explored how their Abeta studies using Folding@home could be used as a starting point for a new Alzheimer's therapy. Folding@home is currently concentrating on Alzheimer's and continues full-scale simulations of amyloid beta and its oligomerization,[42][48] which had previously been a technological challenge to simulate.[47] These studies build off of the Pande lab's 2008 published work into new ways to simulate Abeta oligomerization over long timescales. In the same publication, previous all-atom simulations were performed that led to specific experimentally-tested predictions, such as ways in which to stabilize the protein and prevent the toxic oligomer formation. The Pande lab is focusing their research in this area for rational drug design approaches.[12] Pande described that paper as the "tip of the iceberg" for the Folding@home studies of Alzheimer's, as further results will follow and possibly new therapeutics.[42][47]
Folding@home is also being used to study Aß fragments of different sizes to determine how various natural enzymes affect the structure and folding of Aß. These fragments are tied to senile plaques, a pathological marker of Alzheimer's disease in patient's brain. When certain enzymes cleave the amyloid precursor protein, Abeta peptides are produced, while the action of other enzymes can instead produce p3 peptides, much smaller fragments of Aß. Folding@home is simulating one of these smaller peptides in water in an effort to determine how the length of Aß affects its overall structure.[49]
In 2010, several possible drug leads predicted by Folding@home went from the test tube to testing on living tissue, and in close cooperation with the Nanomedicine Center for Protein Folding, the drug leads continued to be refined. Additionally, as predicted by FAH's simulations, a stable form of amyloid beta was experimentally verified which the Pande lab believes could be used as a starting point for new Alzheimer's therapy.[42][47] In 2008, Folding@home produced several small drug candidates to fight Alzheimer's Disease, as they appear to inhibit the toxicity of Abeta.[50]
The Pande lab is also using Folding@home to investigate protein–protein interactions, which occur extensively throughout both benign and disease-related biological activities. Interactions involving the common SH3 protein are also being studied, as it has implications in Alzheimers research. The refinement of these simulations has greatly improved the Pande lab's ability to understand a wide variety of biological interactions.[51]
Huntington's disease
Huntington's disease, an incurable neurodegenerative genetic disorder affecting muscle coordination and leading to dementia, is also associated with protein misfolding. Specifically, it is caused by a mutation in the Huntingtin gene, which causes excessively long repetitive chains of the glutamine amino acid in the Huntingtin protein, a protein that plays important roles in nerve cells.[52][53] The likelihood of neuronal cell death is primarily affected by the length of the glutamine chain and the neuron's intracellular exposure to the misfolded Huntingtin protein.[54] The defective protein causes Huntington's by aggregating most often in the striatum and frontal cortex of patient's brains. The Pande lab is using Folding@home to study these aggregates, as well as predict how they form.[12] How this aggregation occurs has been largely unknown, but in 2009 a paper based off of Folding@home's results and published in the Journal of Molecular Biology investigated possible mechanisms for the aggregation formation, and the implications into how to prevent it.[53] These studies will be useful for drug design approaches against the disease, and will serve as a foundation for methods to stop the aggregation formation. Additionally, some of the methods used to study Huntington's are also being used for Alzheimer's research.[12]
In 2010, Folding@home researcher Veena Thomas proposed a novel therapeutic strategy for HD, which may be funded by the NIH. This strategy could be used to bring the results from Folding@home directly to a therapeutic.[12]
Cancer
More than half of all known cancers involve mutations in p53, a tumor suppressor protein present in every cell which signals for cell death in the event of damage to a cell's DNA. If p53 becomes mutated, breaks down, or fails to fold properly, it will no longer be able to cause the "stop signal" for cell division. Cells will consequently divide and grow uncontrollably to form tumors.[55] Folding@home is used to study specific properties of p53 in order to understand and predict these mutations, and in 2007 work began to develop inhibitor proteins that deactivate damaged p53.[12] The Pande lab has also developed a novel experimentally-verified method for predicting how mutations affect p53, which has had reasonable success in the identification of deleterious mutations such as those linked to cancer.[55] This supplemented work performed in 2005 that studied how mutations affect the folding of p53, and which mutations are relevant to cancer. While it was the first peer-reviewed publication from a distributed computing project related to cancer,[56] the study also agreed well with experiments and offered insights that were previously unobtainable.[57] Following these results, the Pande lab have expanded their work to other p53-related diseases.[12]
The Pande lab is also performing research into protein chaperones. These are proteins that assist in the folding of other molecules, assembly of oligomeric structures, the prevention of potential damage caused by protein misfolding, and other functions. They are needed for these purposes by rapidly growing cancerous cells.[58] Using Folding@home and working closely with the Protein Folding Center, they plan to find ways to inhibit chaperones involved in cancer. Using Folding@home for a more comprehensive visualization of their functions, the Pande lab and the Protein Folding Center collectively plan to engineer modified chaperonins to inhibit the folding of particular proteins associated with human diseases such as cancer and Alzheimer's. While this approach has been used before, they believe that this project, if successful, could lead to an interesting new drug against cancer or at least make major advances in that area.[58]
Folding@home is also used to study the folding of several other proteins which have mutations tied to cancer, such as the enzyme src Kinase and certain forms of the Engrailed Homeodomain. These proteins also have a great deal of experimental data for comparison, and serve as a great system for the understanding of folding and misfolding.[59][60] Additionally, the Pande lab is using Folding@home to understand the dynamics of a small knottin protein and how it can be used to bind to contrast agents for imaging scan or drugs.[61] Finally, some forms of interleukin-2, an important signaling protein for the immune system, have been used as immunotherapy for cancer. The Pande lab believes that Folding@home's simulations of its dynamics will lead to insights into how to design other therapeutics.[62]
Parkinson's disease
Parkinson's disease is a degenerative disorder of the central nervous system, characterized by shaking, rigidity, slowness of movement, and dementia. The Pande lab has performed preliminary studies on the properties of alpha-synuclein, a key natively unfolded protein.[12] Particular mutations of alpha-synuclein can aggregate to form toxic fibrils, and while the mechanism of this aggregation remains largely unknown, it can lead to the Parkinson's disease and other conditions.[63] The Pande lab is also testing how Folding@home's methods apply to this problem, and in 2005, Pande presented results from FAH at a National Parkinson Foundation conference.[12]
Osteogenesis imperfecta
Osteogenesis imperfecta is a non-curable genetic bone disorder. Those with the disease are unable to successfully make functional connective bone tissue. This is lethal for many but can also induce a higher rate of miscarriages.[12] The disease is caused by mutations in the Type-1 collagen protein, the most common form of collagen and found abundantly throughout the body. Although some of these mutations of collagen can lead to serious morphological disorders, more benign forms can cause brittle bones and other subtleties.[12] Folding@home has performed simulations of collagen, and has produced a paper on Osteogenesis imperfecta outlining new molecular simulation techniques and revealing new insights into how collagen misfolds. The Pande lab believes these results will be useful for later computational studies of collagen.[64]
Diabetes
Amylin is a misfolded peptide involved in Type II diabetes. While amylin is natively unfolded, it forms an alpha helix structure upon contact with cellular membranes. Moreover, it can aggregate into large deposits on these membranes, inducing cell death of insulin-producing cells, which may be relevant to the development of the disease. Around 95% of patients with Type II diabetes exhibit these aggregate deposits. As of 2011, Folding@home is simulating amylin with the goal to understand how this aggregation forms and to design drugs to prevent it.[65]
Antibiotics
The ribosome is a large biological machine that synthesizes proteins from mRNA. It is the target for approximately half of all known antibiotics, which usually kill bacteria by preventing their ribosomes from making new and essential proteins. Folding@home is simulating the ribosome in detail using new state-of-the-art calculation methods.[12] Results from these simulations have significantly helped the Pande lab prepare to study more complex biomedical problems. The Pande lab is also using Folding@home to perform antibiotic drug design calculations.[12]
Drug design
The Pande lab is using Folding@home to explore how to model and accurately estimate the binding energy of small molecules to a protein. Accurate predictions of binding affinities have the potential to significantly lower the development cost of new drugs.[66] Additionally, Folding@home is utilized to find prime binding sites on protein surfaces by simulating interactions between ligand binding sites with different molecules. This has a direct application to computational drug design.[67] Folding@home is also performing calculations on beta-lactamase, a protein that plays important roles in drug resistance. The Pande lab hopes that by understanding its dynamics, they may be able to design drugs to deactivate it.[68]
Participation
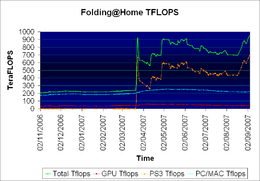
Folding@home computing power shown, by device type, in teraFLOPS as recorded semi-daily from November 2006 until September 2007. Note the large spike in total compute power after March 22, when the PlayStation 3 client was released.
Interest and participation in the project has grown steadily since its launch.[69][70] As of November 17, 2011, Folding@home has about 439,000 active CPUs, about 37,000 active GPUs, and about 21,000 active PS3s, for a total of about 6.7 native petaFLOPS, (9.1 x86 petaFLOPS) more computing power than the combined efforts of all distributed computing projects under BOINC.[71][72] A large majority of this performance comes from the GPU and PS3 clients.[71] Folding@home divides its simulations in such a way that it gains a near-linear speedup for every additional processor.[26][73]
In 2007, Guinness recognized Folding@home as the most powerful distributed computing cluster in the world.[74] Folding@home is also one of the world's largest distributed computing projects.[71] This large and powerful network allows FAH to do work not possible any other way, including through the use of supercomputers,[75] which are typically expensive to operate and often shared.[17][26]
Google collaboration
There used to be cooperation between Folding@home and Google Labs in the form of Google Toolbar.[76][77] Google Compute started in March 2002,[78][79] back when Folding@home only had about 10,000 active CPUs, and pushed Folding@home up to about 30,000 active CPUs.[18] The Google Compute clients had certain limits: it could only run the TINKER core and had limited username and team options.[80] Folding@home is no longer supported on Google Toolbar,[81][82] and even the old Google Toolbar client will not work. Google Compute officially ended in October 2005, and the Google Compute homepage recommends that users wishing to continue contributions to the project download the official Folding@home client instead.[83][84]
Genome@home
Genome@home was another distributed computing project from the Pande lab, a sister project to Folding@home. The goal of Genome@home was protein design and its applications. The project was officially concluded on March 8, 2004, although data was still collected until April 15.[85] It accumulated a large database of protein sequences, which will be used for important scientific purposes for many years by the Pande lab and many others throughout the world.[85][86]
Folding@home clients absorbed the remainder of the Genome@home project on March 12, 2004.[86] The work that was started by the Genome@home project has since been completed using the Folding@home network, (as Work Units without deadlines) and no new work is being distributed by this project.[15] All donators were encouraged to download the Folding@home client (the FAH 4.xx client had a Genome@home option), and once the Genome@home work was complete these clients were asked to donate their processing power to the Folding@home project instead.[86]
PetaFLOPS milestones
Native petaFLOPS threshold Date crossed Fastest Supercomputer at Date Crossed Note 1 1.0 September 16, 2007 0.2806 petaFLOP BlueGene/L[87] 2.0 May 7, 2008 0.4782 petaFLOP BlueGene/L[88] 3.0 August 20, 2008 1.042 petaFLOP Roadrunner[89] 4.0 September 28, 2008 1.042 petaFLOP Roadrunner[89] 5.0 February 18, 2009 1.105 petaFLOP Roadrunner[90] 6.0 November 10, 2011 8.162 petaFLOP K computer[91] On September 16, 2007, the Folding@home project officially attained a sustained performance level higher than one native petaFLOPS, becoming the first computing system of any kind in the world to do so,[92][93] although it had erroneously almost reached that level in March of that year.[94][95] On May 7, 2008, the project attained a sustained performance level higher than two native petaFLOPS,[96] followed by the three and four native petaFLOPS milestones on August 20 and September 28, 2008 respectively.[97][98] Then on February 18, 2009, Folding@home achieved a performance level of just above five native petaFLOPS, thereby becoming the first computing system to surpass that performance, just as it was for the other four milestones.[99][100] Finally, on November 10, 2011, Folding@home crossed the six native petaFLOP barrier with the equivalent of nearly eight x86 petaFLOPS.[101]
Starting in March 2009, Folding@home began reporting performance in both Native FLOPS and x86 FLOPS.[102] As different hardware can perform the same computations in different ways, "x86 FLOPS" are an estimate of how many FLOPS a given calculation would take on the standard x86 CPU architecture.[103] Despite being conservative in the conversions,[104][105] "x86" FLOPS are consistently much greater than the "Native" FLOPS.[103] A detailed explanation of the difference between the two figures is given in the Folding@home FLOP FAQ.[103]
Points
Distributed computing projects are often driven by a sense of collegiate competition to compute the most for the project. Folding@home quantitatively assesses this through points.[106][107] Contributors to Folding@home may select user names to track their contributions.[1] Rankings and other statistics for both individuals and teams are posted to the Folding@home website, with third party statistics sites also available.[108]
Points are determined by the performance of each contributor's folding hardware relative to a reference machine. One or more Work Units from a project are benchmarked on that machine before any donors can do any work on that project. This generates a fair system of equal pay for equal work.[106] However, certain projects require substantially more computer resources (disk space, network usage, RAM, etc.) than others. Contributors can opt in for these projects, and are rewarded with additional bonus points.[106] On some high performance platforms such as the multi-core SMP client, these bonus points can also be rewarded to contributors who reliably and promptly complete Work Units.[106][107][109] The goal is to align the points system with the value of the scientific results.[110] The points can foster friendly competition between donors.[107]
Passkeys
Passkeys are a unique identifier that securely tie contributions directly to each donor.[111] Passkeys are generated from a case-sensitive hash function based on a donor's email address and username. The passkey is then automatically emailed directly to that donor,[112][113] creating a secure system in which only the intended donor receives the passkey.[111] A Folding@home contributor can then enter in the passkey into the client. This not only separates them from any policy issues arising from another donor with that same username, but it also allows for bonus points to be awarded for running the SMP high-performance client.[111]
Teams
Users can register their contributions under a team, and teams register the combined score of all their members.[114] A user can start their own team, or they can join an existing team to add the points they receive to a larger collective.[1] By default, each client is configured to donate under Team 0, but this can be changed in the client's configuration screen.[115]
Teams can be used for troubleshooting or recruitment purposes, but can also keep donors motivated.[116] In some cases, team membership also has a sense of camaraderie, and some teams have their own community-driven sources of help such as a forum.[117][118] In addition, rivalries between teams create friendly competition that benefits the folding community,[119] and members can also have intra-team competitions for top spots.[120][121][122] However, regardless of username or team affiliation, all contributions go to the same place and have the same scientific value.[117]
Software
Folding@home software on the user's end consists of three components: a client, work units, and cores.
Client
Folding@home is powered by volunteers who have installed a client program on their personal computer.[17] The project differs from other distributed computing projects, such as those under BOINC, by offering a variety of clients such as those for multi-core processors, graphics processing units, and PlayStation 3s, complementing the standard client designed for uniprocessor systems. These high performance clients use significantly more computing, memory, and network resources, but return results very quickly, which is of major scientific value.[123] Folding@home is the first project to use GPUs, PS3s, or multi-core processors for distributed computing,[124][125][126] and they have the capability of completing simulations in a few weeks or months rather than years with traditional approaches using a single processor.[123] By supporting machines with varied functionality, Folding@home gains a very rich set of hardware on which to run many different types of customized calculations, thereby achieving a significant increase in performance.[104] The Pande lab's goal is to apply this to dramatically advance the capabilities of Folding@home, as well as apply simulations to further study of protein folding and its related diseases.[123] With all of these clients coupled with new simulation methodologies, they plan to address questions previously considered impossible to tackle computationally, and make even greater impacts on knowledge of folding and misfolding-related diseases.[123]
Each client is the software with which the user interacts, and manages the other software components behind the scenes.[115] Through the client, the user may pause the folding process, open an event log, check the work progress, or view personal statistics.[115] These clients run continuously in the background, using otherwise unused processing power.[1] These clients are designed to run FAH's calculations at an extremely low priority, and will back off to allow other computer programs to have more processing power.[127][128] Although modern computer chips are designed to be able to operate continuously without degrading,[127][129] if users wish to reduce power consumption or heat production, the maximum percentage of CPU power used can be adjusted if desired.[115] If interrupted by a computer shutdown or other means, the client will resume work at almost the same point at startup.[130] For users with machines with multiple processor units, multiple clients may be installed on one machine, and users may be credited by clients on multiple machines.[15]
For security and scientific integrity reasons, the Pande lab does not publicly release the source code of the clients.[6][131] Significant work goes into minimizing security issues in all of Folding@home's software.[15][132] For example, clients can be downloaded only from the official Folding@home website or its commercial partners.[6] It will upload and download data only from Stanford's Folding@home data servers, (over port 8080, with 80 as an alternative)[133] and will only interact with FAH computer files.[15][132] Moreover, it does not normally need computer administrative privileges,[132] so from a security standpoint it behaves similar to but is even more secure than a web browser.[18][133]
Folding@home's first client was a screensaver, which would run Folding@home while the computer was not otherwise in use.[115][134] Later, the Pande lab tested clients on the open source BOINC framework; however, this approach became unworkable and was abandoned in June 2006.[135] Both BOINC and Folding@home clients fell short, for neither client type had enough ability to be compatible with the other. BOINC lacked many features that FAH needed, and FAH lacked features that BOINC needed.[136]
Graphics processing units
GPUs are computer chips used to accelerate 3D graphics, which are most commonly found in video games. GPUs have the capability to significantly out-perform CPUs in terms of Floating Point OPerations, (FLOPs) at the cost of lower generality.[135][137] For this reason, high-performance computing is increasingly utilizing the GPU specialized hardware. In the first large-scale test of GPU scientific reliability, the Pande lab found that although GPUs lack built-in memory error detection and correction, reliable scientific computation can be performed on consumer-grade hardware, as long as sufficient measures are taken (such as Folding@home's built-in error detection) to ensure data integrity.[138]
Despite their potential benefits, scientific computing on GPUs has previously remained inefficient and difficult. The Pande lab has been able to write OpenMM, an open source molecular dynamics library optimized to take full advantage of the GPU architecture and gain large speed increases over conventional single-CPU implementations.[139] As an abstraction layer, OpenMM allows molecular dynamics simulations to be efficiently run across a variety of computer architectures and platforms, something previously problematic in scientific software development.[140] GPUs remain the most powerful platform available in terms of FLOPS; as of September 23, 2011, GPU clients account for 71% of the entire project's FLOP throughput.[71]
The first generation of Folding@home's Windows GPU client (GPU1) was released to the public on October 2, 2006,[135] and delivered a 20-30X speedup for certain calculations over its CPU-based Gromacs counterparts.[141] It was the first time GPUs had been used for either distributed computing or major molecular dynamics calculations.[124] The Pande lab learned much about the development of GPGPU software, but citing a need to improve scientific accuracies over DirectX,[141][142] it was succeeded by GPU2, the second generation successor of the client on April 10, 2008.[143] Several months later, GPU1 was officially retired on June 6.[144] Compared to GPU1, GPU2 was more scientifically reliable and productive, ran on ATI and CUDA-enabled Nvidia GPUs, and supported more advanced algorithms, larger proteins, and real-time visualization of the protein simulation.[145][146] Following this, the third generation of Folding@home's GPU client (GPU3) was released on May 25, 2010. While backwards compatible to GPU2, GPU3 is more stable and efficient, has additional scientific capabilities,[147][148] and uses the OpenCL framework and the Pande lab's OpenMM library.[148][149] Although it does not natively support the Linux operating system, it can be run under WINE for donors with Nvidia graphics cards.[150][151]
PlayStation 3
Main article: Life with PlayStationFolding@home can also take advantage of the computing power of PlayStation 3s, to achieve performance previously only possible on supercomputers. Unlike Microsoft's Xbox,[104] the PS3 is well suited for Folding@home simulations. Its main Cell processor delivers a 20x speed increase over PCs for certain calculations,[17][18] allowing the Pande lab to address problems previously considered impossible to tackle computationally.[104][152] This speed and high efficiency introduced other opportunities for worthwhile optimizations in other area. Moreover, the PS3's streaming processor radically changes the view on the tradeoff between computational efficiency and overall accuracy, allowing for employment of complex mathematical molecular models with even greater accuracy at little extra computational cost.[153] These additional capabilities also allow for greater insights into disease research.[104]
The PS3 processor takes the middle path between flexibility and speed, performing a limited set of calculations rapidly while still retaining the ability to process other types. This makes it more flexible than FAH's GPUs, but less so than CPUs.[135] However, unlike CPUs and GPUs, donors cannot play video games while running Folding@home calculations.[152] Instead, the Pande lab has specifically designed PS3 Work Units to take approximately eight hours so that they can be completed overnight. These WUs have a deadline of two days.[104]
The PS3's uniform console environment makes support easier, as well as making Folding@home user friendly.[17] The PS3 also has the ability to stream data quickly to its GPU, allowing for real-time atomic detail visualizations of the protein dynamics.[135] The PS3 client was originally a standalone application, but since September 18, 2008 is now a channel of the application Life with PlayStation, developed in a collaborative effort between Sony and the Pande lab.[104] Both Sony and the Pande lab have acknowledged the PS3 client is safe to use.[104]
Multi-core processing client
The Symmetric MultiProcessing (SMP) client fulfills two purposes: it takes advantage of the high-performance capabilities of recent multiprocessor systems, and it helps develop a simulation architecture that will become one of the dominant FAH computing paradigms as multi-core chips become an industry standard over the next several years.[18][123][135] The SMP client is capable of delivering over a 4x calculation speedup over the standard uniprocessor clients.[123]
Folding@home's SMP core handles multi-core CPUs very different from other distributed computing projects, including those under BOINC.[154] Instead of simply doing multiple Work Units simultaneously, single WUs are completed much faster across the multiple CPU cores.[155] This cuts down on the traditional difficulties of scaling a large simulation to many processors. As such, this approach is very scientifically valuable. Some of the Pande lab publications would not have been possible without the SMP client.[155]
On November 13, 2006, first generation SMP Folding@home clients for x86 Microsoft Windows, x86-64 Linux, and x86 Mac OS X were released.[123] These clients used Message Passing Interface (MPI) protocols on the localhost,[123] as at the time the Gromacs cores were not designed to be used with multiple threads. This made Folding@home the first to use MPI for distributed computing software, as it had previously been reserved only for supercomputers.[156] The MPI-based clients worked well in Unix-based operating systems such as Linux and Mac's OS-X, but was particularly troublesome in Windows.[155][156] Despite these difficulties, SMP1 generated significant results that would have been impossible otherwise and which represented a landmark in the simulation of protein folding.[155]
The second generation of the SMP client was released as an open beta on January 24, 2010, and subsequently replaced SMP1.[157] The SMP2 client exchanges the complex MPI for threads, which removes much of the overhead of keeping the cores synchronized.[157][158] The SMP2 client also supports a bonus points system, which non-linearly rewards additional points to donors for quick WU returns and for contributing to next-generation capabilities.[123] Donors who run the SMP2 client receive these extra points if they use a passkey and maintained an 80% successful return of Work Units.[157]
SMP2 also supports extra-large Work Units for users with powerful eight-core CPUs or better. While these WUs consume even more RAM and have more network usage than regular SMP WUs, users who run these are rewarded with a 20% increase over SMP2's bonus point system.[159][160] These powerful computers allow for simulations to be performed on Folding@home that had previously required the use of supercomputing clusters. There is a great scientific need to run these simulations out to long timescales as quickly as possible, so the additional bonus points also serves as an incentive for rapid completions of Work Units. This allows the Pande lab to perform studies of larger molecular systems that would not have been possible anywhere else on Folding@home.[160]
V7
 A sample image of the v7 client in Novice mode running under Windows 7. In addition to a variety of controls and user details, v7 also presents Work Unit information, such as its state, calculation progress, ETA, credit points, and identification
A sample image of the v7 client in Novice mode running under Windows 7. In addition to a variety of controls and user details, v7 also presents Work Unit information, such as its state, calculation progress, ETA, credit points, and identification
The v7 client is the seventh and latest generation of the Folding@home software, currently under heavy development, but available for open beta testing.[4] V7 is a complete rewrite and unification of the previous clients for Microsoft Windows, Mac OS X and Linux operating systems.[161] Following its predecessors, v7 runs Folding@home in the background at very low priority, which allows other applications to use CPU resources as they need.[161] The v7 client is designed to make the installation, start-up, and operation user-friendly for novices, as well as offer greater scientific flexibility than previous clients.[161][162]
V7 consists of several elements. The user interacts with v7's GUI, known as FAHControl. It has Novice, Advanced, and Expert user interface modes, and has the ability to monitor, configure, and control many remote folding clients from a single computer.[161] FAHControl can monitor and direct FAHClient, which runs behind the scenes and in turn manages each FAHSlot (or "slot"). These slots act as replacements for the previously distinct FAH clients, as they may be of Uniprocessor, SMP, or GPU type. Each slot also contains a core and data associated with it, and can download, process, and upload Work Units independently.[161][163] The FAHViewer function, modeled after the PS3 viewer, displays a real-time 3D rendering, if available, of the protein currently being processed.[161][163]
The v7 installer supports Windows 2000, 2003, 2008, XP, Vista, and 7,[161][164] as well as various 32-bit and 64-bit Linux operating systems, such as Debian, Ubuntu, RedHat, Fedora, and CentOS.[161][164] V7 also supports 64-bit OS-X, with a 32-bit version under development.[161] It is the Pande lab's goal to make v7 the recommended client by January 2012 at the latest,[165] and versions of v7 will be frequently released until then.[165]
Work Units
The Work Unit (WU) is the actual data that the client is being asked to process. The client connects to Folding@home servers to retrieve a Work Unit, and returns it upon completion.[166] During transfer, all Work Units are validated through the use of 2048-bit digital signatures.[15] Each WU is identified for its respective protein Project, Run (conformation), Clone (atomic trajectory), and Generation (time steps in the trajectory/simulation).[167] Work Units also have associated deadlines and credit (point) value. If this deadline is is exceeded, the user may not get credit and the unit will be reissued to someone else.[15] As protein folding is serial in nature and each WU is generated from its predecessor, this allows the overall simulation process to proceed normally if a WU is not returned after a certain period of time.[15] Additionally, Work Units for high performance clients have a much shorter deadline than those for the uniprocessor client, as a major part of the scientific benefit is dependent on rapidly completing simulations.[123]
Work units go through several Quality Assurance steps. First, WUs are first tested internally for issues, then they are released to special donors who opt-in for beta WUs, before finally being publicly released across FAH. The goal of this gradual rollout is to keep problematic WUs from becoming fully available.[168]
Unlike BOINC projects such as SETI@home, Folding@home's Work Units are normally processed only once, except in the rare event that errors occur during processing of a WU. If this occurs, the Work Unit will still be reissued to two other users,[169] and if it generates errors for those users as well, at 8am PDT it is automatically marked as "bad" and is pulled from distribution.[170][171] Topics in the Folding@home forum can be used to differentiate between problematic hardware and an actual bad Work Unit.[172]
Work Units are very much tied to the Pande lab's simulation Markov state models, which allow for extensive parallelization of very long simulation processes which otherwise seem intrinsically serial.[166][173] It has been demonstrated that during the folding process, proteins spend much of their time "waiting" in various states, before quickly transitioning to the next configuration. This allows for the unique possibility to simulate only a small fraction of the overall folding timescale, leading to significant speedups.[26] The Pande lab achieves this by first dividing the protein's possible dynamics into a series of related conformation states, and creates WUs to calculate the rates of transition between these states. When the completed WUs are gathered, the Pande lab then runs sophisticated Bayesian Machine Learning algorithms which calculate which states are reasonable and the rates between them.[166] This also averages the molecular simulation ensembles, which is important so that direct, meaningful comparison between in silico simulations and in vitro experiments can be made.[174][175] This system is successful even at the millisecond timescale, compares well to traditional methods and experimental results, and allows for previously intractable problems to be within reach.[16][73]
Any computer can contribute to Folding@home. However, older and slower machines might not be fast enough to complete a Work Unit before the deadline. The Pande lab states that a Pentium 3 450 MHz CPU with SSE or newer is able to complete WUs before they expire.[15]
Cores
Main article: Folding@home coresSpecialized scientific computer programs, referred to as "cores," perform the calculations on the Work Unit behind the scenes.[176] Folding@home's cores are based on modified and optimized versions of molecular dynamics programs for calculation, including GROMACS, AMBER, TINKER, CPMD, SHARPEN, ProtoMol, BrookGPU and Desmond.[176][177] These variants are each given an arbitrary identifier (Core xx). While the same core can be used by various versions of the client, separating the core from the client enables the scientific methods to be updated automatically as needed without a client update.[176]
Some of these cores do explicit molecular dynamics calculations, which simulate atoms based on the forces that the atoms exert on each other.[178] These types of calculations reveal how folding happened, not just what the final outcome is, and often tend to be more efficient and optimized than the Monte Carlo methods used to predict particular final properties of a protein.[178] Other cores, such as those designed for the PS3, perform implicit solvation methods, which treat atoms as a mathematical continuum.[179]
Following the Pande lab's pattern of openness, these scientific cores are open-source software or are under similar licenses.[180][181] During download, the cores are verified by 2048-bit digital signatures.[15][182] This ensures very tight security using the best possible software security measures.[15]
Comparison to other molecular systems
Rosetta@home is a distributed computing project aimed at protein structure prediction and is one of the most successful approaches to this problem.[183] Folding@home and Rosetta@home address very different molecular questions.[184] Although Rosetta@home does not provide information into how proteins fold, it does predict the protein's most likely final structure, which in some cases is used as a basis for Folding@home's projects.[185][186] Rosetta's predictions can help FAH simulate the folding of larger proteins more efficiently. Folding@home can also verify Rosetta@home's results and find additional atomistic details of the protein's kinetics and folding pathway,[185] which is intrinsically much more difficult.[30] Folding@home's accurate simulations have also suggesting important novel implications into the fields of protein folding, structure prediction, and certain folding experiments,[187] and have shown that Rosetta's structure prediction may benefit from thermodynamic sampling aspects of protein folding mechanisms.[188][189]
Folding@home also compares well to Anton, a powerful supercomputer which uses specialized hardware to produce a small number of ultra-long molecular trajectories. It is unique in this ability, and like Folding@home, has also improved particular long-held theories of protein folding. Its longer simulations, while computationally expensive, contain more phase space than any one of Folding@home's many shorter trajectories, which allows Anton to perform a thorough exploration of the required space.[190] As of October 2011, Anton and FAH are the two most powerful molecular dynamics systems,[191] and Anton has also run simulations out to the millisecond range.[192][193] In 2011, the Pande lab built a Markov state model from one of Anton's simulations.[190] It demonstrated that there was little difference between MSMs built from Anton's fewer long trajectories and one assembled from Folding@home's many shorter trajectories. Their analysis also showed that Folding@home's Markov state models significantly improve the analysis of these longer simulations, such as revealing additional relevant folding pathways and information into how the protein carries out its biological function.[190] Folding@home is running further analysis on one of Anton's simulations to better determine how its approaches compare to Anton's methods.[194] It is probable that a combination of Anton's and F@h's simulation methods would be very beneficial,[190] and Pande looks forward to see how Anton and FAH can be used together.[191]
See also
- Storage@home
- List of distributed computing projects
- Software for molecular modeling
- Molecular modeling on GPUs
- Rosetta@home
- Blue Gene
- Molecular dynamics
- Computational biology
External links
- Folding@home homepage
- Folding@home official blog
- Folding@home forum
- Folding@home statistics
- List of publications from F@h results
- "Futures In Biotech" episode (for newcomers)
- 2009 Interview with Vijay Pande about Folding@home Project
- Video of record-breaking 1.5ms protein fold
- Pande lab's OpenMM molecular dynamics library
- Simple multimedia presentation about F@h
- Folding@home Wiki
- Wikipedia Team
Notes
Note 1: It should be noted that accurate measurement of the speed of a supercomputer does not necessarily equate to scientific productivity. Supercomputers are typically tested for brief periods using the legacy LINPACK benchmark. This brief CPU testing is not an accurate indication of their prolonged performance over real-world tasks.[110][195]
References
- ^ a b c d e f Pande lab (2011-09-19). "Download the Folding@home Software Application". Stanford University. http://folding.stanford.edu/English/Download. Retrieved 2011-08-31.
- ^ a b Pande lab. "High Performance Clients". Stanford University. http://folding.stanford.edu/English/DownloadWinOther. Retrieved 2011-08-31.
- ^ "Folding@home for PlayStation3". Sony. 2008. http://www.scei.co.jp/folding/en/update.html. Retrieved 2011-08-31.
- ^ a b "Folding@Home v7 Client Beta Release Page". Stanford University. https://fah-web.stanford.edu/projects/FAHClient/wiki/BetaRelease. Retrieved 2011-09-19.
- ^ Joseph Coffland (2011-10-10). "FAHClient V7.1.38 released (4th Open-Beta)". http://foldingforum.org/viewtopic.php?f=67&t=19795. Retrieved 2011-10-10.
- ^ a b c Pande lab. "Folding@home Distributed Computing Client". Stanford University. http://folding.stanford.edu/English/License. Retrieved 2010-08-26.
- ^ Imran "ihaque" Haque (Pande lab member) (2010-08-11). "Re: FAH really doing anything?". http://foldingforum.org/viewtopic.php?f=17&t=14179#p139017. Retrieved 2011-08-23.
- ^ Pande lab. "Folding@Home Executive summary". Stanford University. http://www.stanford.edu/group/pandegroup/folding/FoldingFAQ.pdf. Retrieved 2011-10-04.
- ^ Pande lab (2011). "Folding@home - Main". Stanford University. http://folding.stanford.edu. Retrieved 2011-10-04.
- ^ Pande lab (2009). "The Science Behind Folding@home" (FAQ). Stanford University. http://folding.stanford.edu/English/Science. Retrieved 2011-08-15.
- ^ Antonella De Jaco, Michael Z. Lin, Noga Dubi, Davide Comoletti, Meghan T. Miller, Shelley Camp, Mark Ellisman, Margaret T. Butko, Roger Y. Tsien, and Palmer Taylor (2010). "Neuroligin Trafficking Deficiencies Arising from Mutations in the a/ß-Hydrolase Fold Protein Family". Journal of Biological Chemistry 285 (37): 28674–28682. doi:10.1074/jbc.M110.139519. PMC 2937894. PMID 20615874. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2937894.
- ^ a b c d e f g h i j k l m n o p q Pande lab. "Folding@home Diseases Studied FAQ" (FAQ). Stanford University. http://folding.stanford.edu/FAQ-diseases.html. Retrieved 2011-09-23.
- ^ Xiche Hu, Dong Xu, Kenneth Hamer, Klaus Schulten, Juergen Koepke, and Hartmut Michel (1991). "Knowledge Based Structure Prediction of the Light-Harvesting Complex II of Rhodospirillum molischianum". http://www.ks.uiuc.edu/Publications/Papers/PDF/HU96/HU96.pdf. Retrieved 2011-09-10.
- ^ a b Pande lab (2011-08-05). "Folding@home - Papers". Stanford University. http://folding.stanford.edu/English/Papers. Retrieved 2011-10-09.
- ^ a b c d e f g h i j k Pande lab (2011-08-18). "Main F@h FAQ" (FAQ). Stanford University. http://folding.stanford.edu/English/FAQ-main. Retrieved 2011-10-23.
- ^ a b Vijay Pande (2010-01-17). "Folding@home: Paper #72: Major new result for Folding@home: Simulation of the millisecond timescale". http://folding.typepad.com/news/2010/01/major-new-result-from-foldinghome-simulation-of-the-millisecond-timescale.html. Retrieved 2011-09-08.
- ^ a b c d e f g Pande lab. "Folding@Home Press FAQ" (FAQ). Stanford University. http://folding.stanford.edu/English/FAQ-Press. Retrieved 2011-08-31.
- ^ a b c d e f "Futures in Biotech 27: Folding@home at 1.3 Petaflops" (Interview, webcast). 2007-12-28. http://twit.tv/fib27.
- ^ Pande lab (2011). "Folding@home - About". Stanford University. http://folding.stanford.edu/English/About. Retrieved 2011-08-31.
- ^ Anfinsen, C.B., Haber, E., Sela, M., White Jr., F.H. (1961). "The kinetics of formation of native ribonuclease during oxidation of the reduced polypeptide chain". Proceedings of the National Academy of Sciences of the United States of America 45 (1): 1309–1314. ISSN 00278424.
- ^ Alberts, Bruce; Alexander Johnson, Julian Lewis, Martin Raff, Keith Roberts, and Peter Walters (2002). "The Shape and Structure of Proteins". Molecular Biology of the Cell; Fourth Edition. New York and London: Garland Science. ISBN 0-8153-3218-1.
- ^ Uversky, V.N. (2009). "Intrinsic disorder in proteins associated with neurodegenerative diseases". Frontiers in Bioscience 14 (14): 5188–5238. doi:10.2741/3594. ISSN 10939946.
- ^ a b "Center for Protein Folding Machinery". Protein Folding Center. 2011. http://www.proteinfoldingcenter.org/. Retrieved 2011-09-30.
- ^ a b c d G. Bowman, V. Volez, and V. S. Pande (2011). "Taming the complexity of protein folding". Current Opinion in Structural Biology 21 (1): 4–11. doi:10.1016/j.sbi.2010.10.006. PMC 3042729. PMID 21081274. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3042729.
- ^ "Bio-X Stanford University: Vijay Pande". Bio-X Stanford University. 2011. http://biox.stanford.edu/clark/pande.html. Retrieved 2011-10-16.
- ^ a b c d Vijay S. Pande, Ian Baker, Jarrod Chapman, Sidney P. Elmer, Siraj Khaliq, Stefan M. Larson, Young Min Rhee, Michael R. Shirts, Christopher D. Snow, Eric J. Sorin, Bojan Zagrovic (2002). "Atomistic protein folding simulations on the submillisecond timescale using worldwide distributed computing". Biopolymers 68 (1): 91–109. doi:10.1002/bip.10219. PMID 12579582.
- ^ Stefan M. Larson, Christopher D. Snow, Michael R. Shirts, and Vijay S. Pande (2002). "Folding@home and Genome@home: Using distributed computing to tackle previously intractable problems in computational biology". Computational Genomics. http://fah-web.stanford.edu/papers/Horizon_Review.pdf.
- ^ Christian "schwancr" Schwantes (Pande lab member) (2011-08-15). "Projects 7808 and 7809 to full fah". http://foldingforum.org/viewtopic.php?f=24&t=19376&start=0#p193378. Retrieved 2011-10-16.
- ^ Vijay Pande (2007-11-16). "Paper highlight: paper #50 & #36". http://folding.typepad.com/news/2007/11/paper-highlig-1.html. Retrieved 2011-09-27.
- ^ a b C. D. Snow, E. J. Sorin, Y. M. Rhee, and V. S. Pande. (2005). "How well can simulation predict protein folding kinetics and thermodynamics?". Annual Reviews of Biophysics 34: 43–69. doi:10.1146/annurev.biophys.34.040204.144447. PMID 15869383.
- ^ TJ Lane (Pande lab member) (2011-06-09). "Re: Course grained Protein folding in under 10 minutes". http://foldingforum.org/viewtopic.php?p=188496#p188392. Retrieved 2011-10-29.
- ^ Young Min Rhee, Eric J. Sorin, Guha Jayachandran, Erik Lindahl, & Vijay S Pande (2004). "Simulations of the role of water in the protein-folding mechanism". Proceedings of the National Academy of Sciences, USA 101 (17): 6456–6461. Bibcode 2004PNAS..101.6456R. doi:10.1073/pnas.0307898101. PMC 404066. PMID 15090647. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=404066.
- ^ a b Vijay Pande (2008-01-23). "Random thoughts on the superbowl and statistics". http://folding.typepad.com/news/2008/01/random-thoughts.html. Retrieved 2011-09-11.
- ^ Vijay Pande (2007-09-29). "What can Thermodynamics tell us?". http://folding.typepad.com/news/2007/09/what-can-thermo.html. Retrieved 2011-10-29.
- ^ Vijay Pande (2007-09-30). "How FAH works: Thermodynamic alchemy". http://folding.typepad.com/news/2007/09/how-fah-works-t.html. Retrieved 2011-10-29.
- ^ Tim "7im" Braun (2011-09-24). "Re: New Invention Unravels Mystery of Protein Folding". http://foldingforum.org/viewtopic.php?f=17&t=19671#p195980. Retrieved 2011-09-24.
- ^ Pande lab. "Project 10125 Description". Stanford University. http://fah-web.stanford.edu/cgi-bin/fahproject.overusingIPswillbebanned?p=10125. Retrieved 2011-09-27.
- ^ Pande lab. "Project 2660 Description". Stanford University. http://fah-web.stanford.edu/cgi-bin/fahproject.overusingIPswillbebanned?p=2660. Retrieved 2011-09-27.
- ^ Pande lab. "Project 6811 Description". Stanford University. http://fah-web.stanford.edu/cgi-bin/fahproject.overusingIPswillbebanned?p=6811. Retrieved 2011-09-27.
- ^ G. R. Bowman and V. S. Pande (2010). "Protein folded states are kinetic hubs". Proceedings of the National Academy of Sciences, USA 107 (24): 10890–10895. Bibcode 2010PNAS..10710890B. doi:10.1073/pnas.1003962107. PMC 2890711. PMID 20534497. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2890711.
- ^ V. S. Pande (2010). "A simple theory of protein folding kinetics". Physical Review Letters 105 (19). Bibcode 2010PhRvL.105s8101P. doi:10.1103/PhysRevLett.105.198101.
- ^ a b c d e f Vijay Pande (2011-08-05). "Results page updated – new key result published in our work in Alzheimer's Disease". http://folding.typepad.com/news/2011/08/results-page-updated-new-key-result-published-in-our-work-in-alzheimers-disease.html. Retrieved 2011-09-10.
- ^ a b Vijay Pande (2008-04-23). "Folding@home and Simbios". http://folding.typepad.com/news/2008/04/foldinghome-and.html. Retrieved 2011-11-09.
- ^ a b Vijay Pande (2011-10-25). "Re: Suggested Changes to F@h Website". http://foldingforum.org/viewtopic.php?f=16&t=19643&p=197898#p197898. Retrieved 2011-10-25.
- ^ Lacor PN,et al.; Buniel, MC; Furlow, PW; Clemente, AS; Velasco, PT; Wood, M; Viola, KL; Klein, WL (January 2007). "Aß Oligomer-Induced Aberrations in Synapse Composition, Shape, and Density Provide a Molecular Basis for Loss of Connectivity in Alzheimer's Disease". Journal of Neuroscience 27 (4): 796–807. doi:10.1523/JNEUROSCI.3501-06.2007. PMID 17251419.
- ^ P. Novick, J. Rajadas, C.W. Liu, N. W. Kelley, M. Inayathullah, and V. S. Pande (2011). Buehler, Markus J.. ed. "Rationally Designed Turn Promoting Mutation in the Amyloid-ß Peptide Sequence Stabilizes Oligomers in Solution". PLoS ONE 6 (7): e21776. doi:10.1371/journal.pone.0021776. PMC 3142112. PMID 21799748. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=3142112.
- ^ a b c d Nicholas W. Kelley, V. Vishal, Grant A. Krafft, and Vijay S. Pande. (2008). "Simulating oligomerization at experimental concentrations and long timescales: A Markov state model approach". Journal of Chemical Physics 129 (21): 214707. Bibcode 2008JChPh.129u4707K. doi:10.1063/1.3010881. PMC 2674793. PMID 19063575. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2674793.
- ^ Pande lab. "Project 6802 Description". Stanford University. http://fah-web.stanford.edu/cgi-bin/fahproject.overusingIPswillbebanned?p=6802. Retrieved 2011-09-27.
- ^ Pande lab. "Project 6871 Description". Stanford University. http://fah-web.stanford.edu/cgi-bin/fahproject.overusingIPswillbebanned?p=6871. Retrieved 2011-09-27.
- ^ Vijay Pande (2008-12-18). "New FAH results on possible new Alzheimer's drug presented". http://folding.typepad.com/news/2008/12/new-fah-results-on-possible-new-alzheimers-drug-presented.html. Retrieved 2011-09-23.
- ^ Pande lab. "Project 700 Description". Stanford University. http://fah-web.stanford.edu/cgi-bin/fahproject.overusingIPswillbebanned?p=700. Retrieved 2011-09-27.
- ^ Walker FO (2007). "Huntington's disease". Lancet 369 (9557): 220. doi:10.1016/S0140-6736(07)60111-1. PMID 17240289.
- ^ a b Nicholas W. Kelley, Xuhui Huang, Stephen Tam, Christoph Spiess, Judith Frydman and Vijay S. Pande (2009). "The predicted structure of the headpiece of the Huntingtin protein and its implications on Huntingtin aggregation". Journal of Molecular Biology 388 (5): 919–27. doi:10.1016/j.jmb.2009.01.032. PMC 2677131. PMID 19361448. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2677131.
- ^ Orr HT (October 2004). "Neurodegenerative disease: neuron protection agency". Nature 431 (7010): 747–8. doi:10.1038/431747a. PMID 15483586.
- ^ a b L.T. Chong, W. C. Swope, J. W. Pitera, and V. S. Pande (2006). "A novel approach for computational alanine scanning: application to the p53 oligomerization domain". Journal of Molecular Biology 357 (3): 1039–1049. doi:10.1016/j.jmb.2005.12.083. PMID 16457841.
- ^ Vijay Pande. "First results from Folding@Home cancer project published". http://web.archive.org/web/20070706192334/http://forum.folding-community.org/viewtopic.php?t=10950. Retrieved 2011-11-09.
- ^ L. T. Chong, C. D. Snow, Y. M. Rhee, and V. S. Pande. (2004). "Dimerization of the p53 Oligomerization Domain: Identification of a Folding Nucleus by Molecular Dynamics Simulations". Journal of Molecular Biology 345 (4): 869–878. doi:10.1016/j.jmb.2004.10.083. PMID 15588832.
- ^ a b Vijay Pande (2007-09-28). "Nanomedicine center". http://folding.typepad.com/news/2007/09/nanomedicine-ce.html. Retrieved 2011-09-23.
- ^ Vijay Pande (2009-12-22). "Release of new Protomol (Core B4) WUs". http://folding.typepad.com/news/2009/12/release-of-new-protomol-core-b4-wus-.html. Retrieved 2011-09-23.
- ^ Pande lab. "Project 180 Description". Stanford University. http://fah-web.stanford.edu/cgi-bin/fahproject.overusingIPswillbebanned?p=180. Retrieved 2011-09-27.
- ^ Pande lab. "Project 7600 Description". Stanford University. http://fah-web.stanford.edu/cgi-bin/fahproject.overusingIPswillbebanned?p=7600. Retrieved 2011-09-27.
- ^ Pande lab. "Project 10113 Description". Stanford University. http://fah-web.stanford.edu/cgi-bin/fahproject.overusingIPswillbebanned?p=10113. Retrieved 2011-09-27.
- ^ Arima K, Uéda K, Sunohara N, Hirai S, Izumiyama Y, Tonozuka-Uehara H, Kawai M (October 1998). "Immunoelectron-microscopic demonstration of NACP/alpha-synuclein-epitopes on the filamentous component of Lewy bodies in Parkinson's disease and in dementia with Lewy bodies". Brain Res. 808 (1): 93–100. doi:10.1016/S0006-8993(98)00734-3. PMID 9795161.
- ^ Sanghyun Park, Randall J. Radmer, Teri E. Klein, and Vijay S. Pande (2005). "A New Set of Molecular Mechanics Parameters for Hydroxyproline and Its Use in Molecular Dynamics Simulations of Collagen-Like Peptides". Journal of Computational Chemistry 26 (15): 1612–1616. doi:10.1002/jcc.20301. PMID 16170799.
- ^ Pande lab. "Project 2974 Description". Stanford University. http://fah-web.stanford.edu/cgi-bin/fahproject.overusingIPswillbebanned?p=2974. Retrieved 2011-09-27.
- ^ Pande lab. "Project 3855 Description". Stanford University. http://fah-web.stanford.edu/cgi-bin/fahproject.overusingIPswillbebanned?p=3855. Retrieved 2011-09-27.
- ^ Pande lab. "Project 2450 Description". Stanford University. http://fah-web.stanford.edu/cgi-bin/fahproject.overusingIPswillbebanned?p=2450. Retrieved 2011-09-27.
- ^ Pande lab. "Project 10115 Description". Stanford University. http://fah-web.stanford.edu/cgi-bin/fahproject.overusingIPswillbebanned?p=10115. Retrieved 2011-09-27.
- ^ Vijay Pande (2007-10-21). "Fun fact: FAH growth over time". http://folding.typepad.com/news/2007/10/fun-fact-fah-gr.html. Retrieved 2011-10-21.
- ^ Pande lab. "Active CPUs" (Image). Stanford University. http://www.stanford.edu/group/pandegroup/images/ActiveCPUs2010.png. Retrieved 2011-08-30.
- ^ a b c d Pande lab (updated automatically). "Client Statistics by OS". Stanford University. http://fah-web.stanford.edu/cgi-bin/main.py?qtype=osstats. Retrieved 2011-09-28.
- ^ "BOINC Combined Credit Overview". http://boincstats.com/stats/project_graph.php?pr=bo. Retrieved 2011-11-17.
- ^ a b Michael R. Shirts and Vijay S. Pande (2001). "Mathematical Foundations of Coupled Parallel Simulations". Physical Review Letters 86 (22): 4983–4987. Bibcode 2001PhRvL..86.4983S. doi:10.1103/PhysRevLett.86.4983. PMID 11384401.
- ^ Joshua Topolsky (2007-10-31). "Folding@Home recognized by Guinness World Records". http://www.engadget.com/2007/10/31/folding-home-recognized-by-guinness-world-records/. Retrieved 2007-11-05.
- ^ Caroline Hadley (2004). "Biologists think bigger". EMBO reports 12 (5): 236–238. doi:10.1038/sj.embor.7400108. http://www.nature.com/embor/journal/v5/n3/full/7400108.html.
- ^ Google (2007). "Your computer's idle time is too precious to waste". http://toolbar.google.com/dc/offerdc.html/. Retrieved 2011-09-06.
- ^ "Google Toolbar Compute - Folding@Home". http://www.powder2glass.com/Google_Toolbar_Compute/. Retrieved 2011-09-06.
- ^ Olsen, Stefanie (2003-03-27). "Google tests distributed computing". CNet News. http://news.cnet.com/Google-tests-distributed-computing/2110-1032_3-994371.html.
- ^ Shankland, Stephen (March 22, 2002). "Google takes on supercomputing". CNet News. http://news.cnet.com/2100-1001-867091.html.
- ^ Tim "7im" Braun (2011-03-28). "Re: Weird issue - Not sure where to post". http://foldingforum.org/viewtopic.php?f=61&t=18008&p=180435&hilit=google+compute#p180435. Retrieved 2011-09-06.
- ^ Tim "7im" Braun (2008-06-19). "Re: How close do you think we are to end of the project?". http://foldingforum.org/viewtopic.php?f=16&t=3205&p=31026&hilit=google+compute#p31026. Retrieved 2011-09-06.
- ^ Tim "7im" Braun (2011-08-19). "Re: ChromiumOS Folding integration?". http://foldingforum.org/viewtopic.php?f=14&t=19388&p=193623&hilit=google+compute#p193637. Retrieved 2011-09-06.
- ^ ChelseaOilman (2005-12-30). "Google is after your CPU cycles". http://hardforum.com/showpost.php?p=1028770683&postcount=4. Retrieved 2011-09-06.
- ^ "Google is after your CPU cycles". 2005-12-30. http://hardforum.com/showthread.php?t=997889. Retrieved 2011-09-06.
- ^ a b "Genome@home Updates". 2002-03-04. http://www.stanford.edu/group/pandegroup/genome/new.html. Retrieved 2011-09-05.
- ^ a b c Pande lab. "Genome@home FAQ" (FAQ). Stanford University. http://genomeathome.stanford.edu/faq.html. Retrieved 2011-09-05.
- ^ "TOP500 List - June 2007". Top500. 06-2007. http://www.top500.org/list/2007/06/100. Retrieved 2011-11-11.
- ^ "TOP500 List - November 2007". Top500. 11-2007. http://www.top500.org/list/2007/11/100. Retrieved 2011-11-11.
- ^ a b "TOP500 List - June 2008". Top500. 06-2008. http://www.top500.org/list/2008/06/100. Retrieved 2011-11-11.
- ^ "TOP500 List - November 2008". Top500. 11-2008. http://www.top500.org/list/2008/11/100. Retrieved 2011-11-11.
- ^ "TOP500 List - June 2011". Top500. 06-2011. http://www.top500.org/list/2011/06/100. Retrieved 2011-11-11.
- ^ Vijay Pande (2007-09-16). "Crossing the petaFLOPS barrier". http://folding.typepad.com/news/2007/09/crossing-the-pe.html. Retrieved 2011-08-28.
- ^ David Nagel (2007-09-19). "Folding@home Achieves Petaflop Milestone - PS3 owners help scientists speed up their research". http://games.ign.com/articles/821/821350p1.html. Retrieved 2011-09-06.
- ^ Mark Wilson (2007-03-25). "PS3 Folding@Home TFLOP Rating Demoted by 50%, PFLOPS Still Possible". http://gizmodo.com/246900/breaking-ps3-foldinghome-tflop-rating-demoted-by-50-pflops-still-possible. Retrieved 2011-09-14.
- ^ Tim Hanlon (2007-03-09). "Playstation 3 continues to top Folding@Home statistics". http://www.gizmag.com/go/7086/. Retrieved 2011-09-14.
- ^ "Folding@Home reach 2 Petaflops". 2008-05-08. http://n4g.com/news/143113/ps3-andamp-foldingahome-reach-2-petaflops/com. Retrieved 2011-09-23.
- ^ "NVIDIA Achieves Monumental Folding@Home Milestone With Cuda". 2008-08-26. http://www.nvidia.com/object/io_1219747545128.html. Retrieved 2011-09-06.
- ^ "3 PetaFLOP barrier". 2008-08-19. http://www.longecity.org/forum/topic/23841-3-petaflop-barrier/. Retrieved 2011-09-23.
- ^ Vijay Pande (2009-02-18). "Folding@home Passes the 5 petaFLOP Mark". http://folding.typepad.com/news/2009/02/foldinghome-passes-the-5-petaflop-mark.html. Retrieved 2011-08-31.
- ^ "Crossing the 5 petaFLOPS barrier". 2009-02-18. http://www.longecity.org/forum/topic/26449-crossing-the-5-petaflops-barrier/. Retrieved 2011-09-23.
- ^ Jesse_V (2011-11-10). "Six Native PetaFLOPS". http://foldingforum.org/viewtopic.php?f=16&t=20011#p198840. Retrieved 2011-11-11.
- ^ Vijay Pande (2009-03-18). "FLOPS". http://folding.typepad.com/news/2009/03/flops.html. Retrieved 2011-10-11.
- ^ a b c Pande lab (2009-04-04). "Folding@home FLOP FAQ" (FAQ). Stanford University. http://folding.stanford.edu/English/FAQ-flops. Retrieved 2011-08-28.
- ^ a b c d e f g h Pande lab (2009-02-05). "PS3 FAQ" (FAQ). Stanford University. http://folding.stanford.edu/FAQ-PS3.html. Retrieved 2011-09-05.
- ^ Bruce Borden (bruce) (2011-07-12). "Re: Are my conversion for GPU flops relativly correct? [sic"]. http://foldingforum.org/viewtopic.php?f=16&t=19118#p191086. Retrieved 2011-09-08.
- ^ a b c d Pande lab (2011-02-16). "Folding@home Points FAQ" (FAQ). Stanford University. http://folding.stanford.edu/English/FAQ-Points. Retrieved 2011-08-31.
- ^ a b c Pande lab (2011-02-16). "Folding@home Points FAQ (New Benchmark Machine -- January 2010)" (FAQ). Stanford University. http://folding.stanford.edu/English/FAQ-PointsNew. Retrieved 2011-08-31.
- ^ "Third Party Contributions - Stats Pages". 2011. http://fahwiki.net/index.php/Third_Party_Contributions#Stats_Pages. Retrieved 2011-08-07.
- ^ Peter Kasson (Pande lab member) (2010-01-24). "Upcoming Release of SMP2 Cores". http://foldingforum.org/viewtopic.php?f=24&t=13038. Retrieved 2011-08-31.
- ^ a b Bruce Borden (bruce) (2010-04-07). "Re: Answers to: Reasons for not using F@H". http://foldingforum.org/viewtopic.php?f=16&t=1164&start=165#p138646. Retrieved 2011-09-05.
- ^ a b c Pande lab (2011-05-24). "Folding@home Passkey FAQ" (FAQ). Stanford University. http://folding.stanford.edu/English/FAQ-passkey. Retrieved 2011-09-06.
- ^ Tim "7im" Braun (2011-09-29). "Re: Passkey when changing username". http://foldingforum.org/viewtopic.php?f=61&t=19712#p196346. Retrieved 2011-10-02.
- ^ Pande lab. "Passkey Form". Stanford University. http://fah-web.stanford.edu/cgi-bin/getpasskey.py. Retrieved 2011-09-06.
- ^ "Default Team". http://fah-web.stanford.edu/cgi-bin/main.py?qtype=teampage&teamnum=0. Retrieved 2011-09-06.
- ^ a b c d e Pande lab (2011-02-10). "Windows Uniprocessor Client Installation Guide". Stanford University. http://folding.stanford.edu/English/WinUNIGuide. Retrieved 2011-09-05.
- ^ MtM (2009-12-17). "Re: New to F@H need startup info". http://foldingforum.org/viewtopic.php?f=47&t=12499#p122465. Retrieved 2011-09-30.
- ^ a b "Re: Why join a team?". 2009-08-28. http://foldingforum.org/viewtopic.php?f=16&t=11242&start=0#p109792. Retrieved 2011-09-08.
- ^ "Official Extreme Overclocking Folding@Home Team Forum". Extreme Overclocking. http://forums.extremeoverclocking.com/forumdisplay.php?f=45. Retrieved 2011-09-08.
- ^ Norman Chan (2009-04-06). "Help Maximum PC's Folding Team Win the Next Chimp Challenge!". http://www.maximumpc.com/article/news/help_maximum_pcs_folding_team_win_next_chimp_challenge. Retrieved 2011-09-06.
- ^ "Team 24 Folding at Home - March Challenge". 2011-02-20. http://forums.overclockers.com.au/showthread.php?t=940743. Retrieved 2011-09-06.
- ^ "Announcing the Immortality Institute Folding@Home Prize". 2008-03-20. http://www.mprize.org/index.php?ctype=news&pagename=blogdetaildisplay&BID=2008032-20053630&detaildisplay=Y. Retrieved 2011-09-06.
- ^ "Announcing the F@H Prize". Immortality Institute. http://imminst.org/announcing-foldinghome-prize. Retrieved 2011-09-06.
- ^ a b c d e f g h i j Pande lab (2010-12-11). "Folding@home SMP FAQ" (FAQ). Stanford University. http://folding.stanford.edu/English/FAQ-SMP. Retrieved 2011-08-31.
- ^ a b Vijay Pande (2008-05-23). "GPU news (about GPU1, GPU2, & NVIDIA support)". http://folding.typepad.com/news/2008/05/gpu-news-gpu1-g.html. Retrieved 2011-09-08.
- ^ Travis Desell1, Anthony Waters, Malik Magdon-Ismail, Boleslaw K. Szymanski, Carlos A. Varela, Matthew Newby, Heidi Newberg, Andreas Przystawik, and David Anderson (2009). "Accelerating the MilkyWay@Home volunteer computing project with GPUs". 8th International Conference on Parallel Processing and Applied Mathematics (PPAM 2009). doi:10.1.1.158.7614. http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.158.7614&rep=rep1&type=pdf.
- ^ M. J. Harvey, G. Giupponi, Vill`a-Freixa, and G. De Fabritiis (2007). "PS3GRID.NET: Building a distributed supercomputer using the PlayStation 3". Distributed & Grid Computing - Science Made Transparent for EVeryone. Principles, Applications and Supporting Communities. http://www.gpugrid.net/pub/ps3grid_chapter.pdf.
- ^ a b John Naylor (2008-02-09). "Answers to: Reasons for not using F@H". http://foldingforum.org/viewtopic.php?f=16&t=1164#p9750. Retrieved 2011-09-05.
- ^ Bruce Borden (bruce) (2011-09-11). "Re: F@h Advertisement Techniques". http://foldingforum.org/viewtopic.php?f=16&t=19555#p195220. Retrieved 2011-10-18.
- ^ Tim "7im" Braun (2008-09-28). "Answers to: Reasons for not using F@H". http://foldingforum.org/viewtopic.php?f=16&t=1164&start=75#p58696. Retrieved 2011-09-05.
- ^ Bruce Borden (bruce) (2008-07-28). "Re: Answers to: Reasons for not using F@H". http://foldingforum.org/viewtopic.php?f=16&t=1164&start=75#p43364. Retrieved 2011-09-05.
- ^ Vijay Pande (2009-08-20). "Importance of software and data integrity". http://folding.typepad.com/news/2009/08/importance-of-software-and-data-integrity.html. Retrieved 2011-10-19.
- ^ a b c Bruce Borden (bruce) (2011-07-18). "Re: Advice for a new user". http://foldingforum.org/viewtopic.php?f=61&t=19163#p191513. Retrieved 2011-09-11.
- ^ a b Pande lab (2009-11-19). "Uninstalling Folding@home FAQ". Stanford University. http://folding.stanford.edu/English/FAQ-Uninstall. Retrieved 2011-09-21.
- ^ M. R. Shirts and V. S. Pande. (2000). "Screen Savers of the World, Unite!". Science 290 (5498): 1903–1904. doi:10.1126/science.290.5498.1903. PMID 17742054.
- ^ a b c d e f Pande lab (2010-05-13). "High Performance FAQ" (FAQ). Stanford University. http://folding.stanford.edu/English/FAQ-highperformance. Retrieved 2011-09-05.
- ^ Tim "7im" Braun (2010-04-02). "Re: Answers to: Reasons for not using F@H". http://foldingforum.org/viewtopic.php?f=16&t=1164&start=135#p137893. Retrieved 2011-09-05.
- ^ Vijay Pande (2007-09-24). "How FAH works: Folding on streaming processors (GPU's and PS3)". http://folding.typepad.com/news/2007/09/how-fah-works-f.html. Retrieved 2011-10-29.
- ^ I. Haque and V. S. Pande (2010). "Hard Data on Soft Errors: A Large-Scale Assessment of Real-World Error Rates in GPGPU". 2010 10th IEEE/ACM International Conference on Cluster, Cloud and Grid Computing (CCGrid): 691–696. doi:10.1109/CCGRID.2010.84.
- ^ M. S. Friedrichs, P. Eastman, V. Vaidyanathan, M. Houston, S. LeGrand, A. L. Beberg, D. L. Ensign, C. M. Bruns, V. S. Pande (2009). "Accelerating Molecular Dynamic Simulation on Graphics Processing Units". Journal of Computational Chemistry 30 (6): 864–72. doi:10.1002/jcc.21209. PMC 2724265. PMID 19191337. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2724265.
- ^ P. Eastman and V. S. Pande (2010). "OpenMM: A Hardware Abstraction Layer for Molecular Simulations". Computing in Science & Engineering 12 (4): 34–39. doi:10.1109/MCSE.2010.27.
- ^ a b Pande lab (2011-03-18). "ATI FAQ" (FAQ). Stanford University. http://folding.stanford.edu/English/FAQ-ATI. Retrieved 2011-08-31.
- ^ Vijay Pande (2008-05-27). "More info about the GPU1 to GPU2 transition". http://folding.typepad.com/news/2008/05/gpu1-to-gpu2-transition-why-does-gpu1-need-to-end.html. Retrieved 2011-09-07.
- ^ Vijay Pande (2008-04-10). "GPU2 open beta". http://folding.typepad.com/news/2008/04/gpu2-open-beta.html. Retrieved 2011-09-07.
- ^ Vijay Pande (2008-06-06). "GPU1 has been retired, GPU2 for NVIDIA release nearing". http://folding.typepad.com/news/2008/06/gpu1-has-been-retired-gpu2-for-nvidia-release-nearing.html. Retrieved 2011-09-07.
- ^ Vijay Pande (2008-04-15). "Updates to the Download page/GPU2 goes live". http://folding.typepad.com/news/2008/04/updates-to-the.html. Retrieved 2011-09-07.
- ^ Vijay Pande (2008-04-11). "GPU2 open beta going well". http://folding.typepad.com/news/2008/04/gpu2-open-bet-1.html. Retrieved 2011-09-07.
- ^ Vijay Pande (2008-01-25). "Code Development Updates". http://folding.typepad.com/news/2008/01/code-developmen.html. Retrieved 2011-09-07.
- ^ a b Vijay Pande (2010-04-24). "Prepping for the GPU3 rolling: new client and NVIDIA FAH GPU clients will (in the future) need CUDA 2.2 or later". http://folding.typepad.com/news/2010/04/prepping-for-the-gpu3-rolling-new-client-and-nvidia-fah-gpu-clients-will-need-cuda-22-or-later.html. Retrieved 2011-09-08.
- ^ Vijay Pande (2010-05-25). "Folding@home: Open beta release of the GPU3 client/core". http://folding.typepad.com/news/2010/05/open-beta-release-of-the-gpu3-clientcore.html. Retrieved 2011-09-07.
- ^ Joseph Coffland (2011-10-13). "Re: FAHClient V7.1.38 released (4th Open-Beta)". http://foldingforum.org/viewtopic.php?f=67&t=19795&start=45#p197198. Retrieved 2011-10-15.
- ^ "NVIDIA GPU3 Linux/Wine Headless Install Guide". 2008-11-08. http://foldingforum.org/viewtopic.php?f=54&t=6793. Retrieved 2011-09-05.
- ^ a b David E. Williams (2006-10-20). "PlayStation's serious side: Fighting disease". CNN. http://edition.cnn.com/2006/TECH/fun.games/09/18/playstation.folding/. Retrieved 2011-10-16.
- ^ Edgar Luttmann, Daniel L. Ensign, Vishal Vaidyanathan, Mike Houston, Noam Rimon, Jeppe Øland, Guha Jayachandran, Mark Friedrichs, Vijay S. Pande (2008). "Accelerating Molecular Dynamic Simulation on the Cell processor and PlayStation 3". Journal of Computational Chemistry 30 (2): 268–274. doi:10.1002/jcc.21054. PMID 18615421.
- ^ Ozzfan (2007-10-01). "Can I have multiple CPUs work on the same task (workunit)?". BOINC. http://boincfaq.mundayweb.com/index.php?language=1&view=115&sessionID=f68532da1ffd360def2213188757c695. Retrieved 2011-10-09.
- ^ a b c d Vijay Pande (2008-06-15). "What does the SMP core do?". http://folding.typepad.com/news/2008/06/what-does-the-smp-core-do.html. Retrieved 2011-09-07.
- ^ a b Vijay Pande (2008-03-08). "New Windows client/core development (SMP and classic clients)". http://folding.typepad.com/news/2008/03/new-windows-cli.html. Retrieved 2011-09-30.
- ^ a b c Peter Kasson (Pande lab member) (2010-01-24). "upcoming release of SMP2 cores". http://foldingforum.org/viewtopic.php?f=24&t=13038#p127406. Retrieved 2011-09-30.
- ^ Vijay Pande (2009-06-17). "How does FAH code development and sysadmin get done?". http://folding.typepad.com/news/2009/06/how-does-fah-code-development-and-sysadmin-get-done.html. Retrieved 2011-10-14.
- ^ Vijay Pande (2011-07-02). "Change in the points system for bigadv work units". http://folding.typepad.com/news/2011/07/change-in-the-points-system-for-bigadv-work-units.html. Retrieved 2011-10-09.
- ^ a b Peter Kasson (Pande lab member) (2009-07-15). "new release: extra-large work units". http://foldingforum.org/viewtopic.php?t=10697. Retrieved 2011-10-09.
- ^ a b c d e f g h i Vijay Pande (2011-03-29). "Client version 7 now in open beta". http://folding.typepad.com/news/2011/03/client-version-7-now-in-open-beta.html. Retrieved 2011-08-14.
- ^ Vijay Pande (2011-03-31). "Core 16 for ATI released; also note on NVIDIA GPU support for older boards". http://folding.typepad.com/news/2011/03/core-16-for-ati-released-also-note-on-nvidia-gpu-support-for-older-boards.html. Retrieved 2011-09-07.
- ^ a b Pande lab (2011-09-23). "Windows (FAH V7) Install Guide". Stanford University. http://folding.stanford.edu/English/WinGuide. Retrieved 2011-10-09.
- ^ a b Pande lab (2011). "Client FAQ" (FAQ). Stanford University. https://fah-web.stanford.edu/projects/FAHClient/wiki/ClientFAQ. Retrieved 2011-08-14.
- ^ a b Joseph Coffland (2011-09-19). "FAHClient V7.1.33 released (3rd Open-Beta)". http://foldingforum.org/viewtopic.php?f=67&t=19648#p195686. Retrieved 2011-09-19.
- ^ a b c Vijay Pande (2007-09-22). "How FAH works: Markov State Models". http://folding.typepad.com/news/2007/09/how-fah-works-m.html. Retrieved 2011-10-14.
- ^ Dan Ensign. "Runs, Clones, and Generations". http://fahwiki.net/index.php/Runs,_Clones_and_Gens. Retrieved 2011-08-10.
- ^ Vijay Pande (2011-04-05). "More transparency in testing". http://folding.typepad.com/news/2011/04/more-transparency-in-testing.html. Retrieved 2011-10-14.
- ^ Bruce Borden (bruce) (2011-08-07). "Re: Gromacs Cannot Continue Further". http://foldingforum.org/viewtopic.php?f=59&t=19315#p192836. Retrieved 2011-08-07.
- ^ Bruce Borden (bruce) (2011-08-09). "Re: Project: 6053 (Run 1, Clone 194, Gen 357)". http://foldingforum.org/viewtopic.php?f=19&t=19325&p=192927&hilit=8am#p192927. Retrieved 2011-08-09.
- ^ PantherX (2011-10-01). "Re: Project 6803: (Run 4, Clone 66, Gen 255)". http://foldingforum.org/viewtopic.php?f=19&t=19725#p196444. Retrieved 2011-10-09.
- ^ PantherX (2010-10-31). "Troubleshooting Bad WUs". http://foldingforum.org/viewtopic.php?f=19&t=16526. Retrieved 2011-08-07.
- ^ Pande lab. "Folding@home project descriptions - Project 10513". http://fah-web.stanford.edu/cgi-bin/fahproject.overusingIPswillbebanned?p=10513. Retrieved 2011-08-15.
- ^ Bojan Zagrovic, Christopher D. Snow, Michael R. Shirts, and Vijay S. Pande. (2002). "Simulation of Folding of a Small Alpha-helical Protein in Atomistic Detail using Worldwide distributed Computing". Journal of Molecular Biology 323 (5): 927–937. doi:10.1016/S0022-2836(02)00997-X. PMID 12417204.
- ^ Christopher D. Snow, Houbi Ngyen, Vijay S. Pande, and Martin Gruebele (2002). "Absolute comparison of simulated and experimental protein-folding dynamics". Nature 420 (6911): 102–106. doi:10.1038/nature01160. PMID 12422224.
- ^ a b c "Cores - FaHWiki" (FAQ). http://fahwiki.net/index.php/Cores. Retrieved 2007-11-06.
- ^ Pande lab (2005-10-16). "Folding@home with QMD core FAQ" (FAQ). Stanford University. http://folding.stanford.edu/QMD.html. Retrieved 2006-12-03.
- ^ a b Vijay Pande (2007-09-26). "How FAH works: Molecular dynamics". http://folding.typepad.com/news/2007/09/how-fah-works-1.html. Retrieved 2011-10-14.
- ^ Roux B, Simonson T (April 1999). "Implicit solvent models". Biophys. Chem. 78 (1–2): 1–20. doi:10.1016/S0301-4622(98)00226-9. ISSN 0301-4622. PMID 17030302.
- ^ Pande lab (2010-02-03). "Folding@home Open Source FAQ" (FAQ). Stanford University. http://folding.stanford.edu/English/FAQ-OpenSource. Retrieved 2011-09-11.
- ^ Vijay Pande (2011-09-06). "Re: Utilizing this resource". http://foldingforum.org/viewtopic.php?f=17&t=19545&p=194838#p194838. Retrieved 2011-09-11.
- ^ Pande lab. "Folding@home Rules and Policies" (FAQ). Stanford University. http://folding.stanford.edu/English/FAQ-Policies. Retrieved 2011-09-11.
- ^ Gregory R. Bowman and Vijay S. Pande (2009). "Simulated tempering yields insight into the low-resolution Rosetta scoring function". Proteins: Structure, Function, and Bioinformatics 74 (3): 777–88. doi:10.1002/prot.22210. PMID 18767152.
- ^ Vijay Pande (2006-03-05). "Rosetta and Folding". http://web.archive.org/web/20070927193033/http://forum.folding-community.org/viewtopic.php?p=125338. Retrieved 2011-11-09.
- ^ a b TJ Lane (Pande lab member) (2011-06-09). "Re: Course grained Protein folding in under 10 minutes". http://foldingforum.org/viewtopic.php?p=188496#p188496. Retrieved 2011-10-15.
- ^ Bruce Borden (bruce) (2011-09-24). "Re: New Invention Unravels Mystery of Protein Folding". http://foldingforum.org/viewtopic.php?f=17&t=19671#p195977. Retrieved 2011-10-29.
- ^ Bojan Zagrovic, Christopher D. Snow, Siraj Khaliq, Michael R. Shirts, and Vijay S. Pande (2002). "Native-like Mean Structure in the Unfolded Ensemble of Small Proteins". Journal of Molecular Biology 323 (1): 153–164. doi:10.1016/S0022-2836(02)00888-4. PMID 12368107.
- ^ G. R. Bowman and V. S. Pande (2009). Hofmann, Andreas. ed. "The Roles of Entropy and Kinetics in Structure Prediction". PLoS One 4 (6): e5840. Bibcode 2009PLoSO...4.5840B. doi:10.1371/journal.pone.0005840. PMC 2688754. PMID 19513117. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2688754.
- ^ Vijay Pande (2008-04-26). "Re: collaborating with competition". http://foldingforum.org/viewtopic.php?f=15&t=2322&p=20638#p20642. Retrieved 2011-11-09.
- ^ a b c d Thomas J. Lane, Gregory R. Bowman, Kyle A Beauchamp, Vincent Alvin Voelz, and Vijay S. Pande (2011). "Markov State Model Reveals Folding and Functional Dynamics in Ultra-Long MD Trajectories". Journal of the American Chemical Society. doi:10.1021/ja207470h.
- ^ a b Vijay Pande (2011-10-13). "Comparison between FAH and Anton's approaches". http://folding.typepad.com/news/2011/10/comparison-between-fah-and-antons-approaches.html. Retrieved 2011-10-13.
- ^ Heidi Ledford (2010-10-14). "Supercomputer sets protein-folding record". Nature News. doi:10.1038/news.2010.541. http://www.nature.com/news/2010/101014/full/news.2010.541.html. Retrieved 2011-10-15.
- ^ David E. Shaw, Paul Maragakis, Kresten Lindorff-Larsen, Stefano Piana, Ron O. Dror, Michael P. Eastwood, Joseph A. Bank, John M. Jumper, John K. Salmon, Yibing Shan, and Willy Wriggers (2010). "Atomic-Level Characterization of the Structural Dynamics of Proteins". Science 330 (6002): 341–346. doi:10.1126/science.1187409. PMID 20947758.
- ^ Pande lab (2011). "Project 7610 Description". http://fah-web.stanford.edu/cgi-bin/fahproject.overusingIPswillbebanned?p=7610. Retrieved 2011-10-14.
- ^ Vijay Pande (2008-11-09). "Re: ATI and NVIDIA stats vs. PPD numbers". http://foldingforum.org/viewtopic.php?p=67416#p67416. Retrieved 2011-09-22.
PlayStation 3 Accessories Software Network Games Retail games · Downloadable games (PSone Classics · TurboGrafx-16 games · Minis) · PlayStation Move games · Classics HD · 3D games · Backwards compatible PlayStation and PlayStation 2 gamesMedia PlayStation.Blog · PlayStation: The Official Magazine · PlayStation Official Magazine – UK · PlayStation Official Magazine – AustraliaOther PlayStation 3 hardware · PlayStation 3 launch · PlayStation Network outage · Zego · Folding@home · PlayStation 3 cluster · Cell (microprocessor) · RSX 'Reality Synthesizer' · PlayStation JailbreakCategories:- Distributed computing projects
- Molecular dynamics software
- Molecular modelling software
- Molecular modelling
- Simulation software
- Protein folds
- Protein structure
- Computational biology
- Mathematical and theoretical biology
- Bioinformatics
- Computational chemistry
- Hidden Markov models
- Data mining and machine learning software
- Proprietary cross-platform software
- Cross-platform software
- Windows software
- Linux science software
- Mac OS X software
- PlayStation 3 software
- 2000 introductions
Wikimedia Foundation. 2010.