- DNA nanotechnology
-
 DNA nanotechnology seeks to make artificial, designed nanostructures out of nucleic acids, such as this DNA tetrahedron.[1] Each edge of the tetrahedron is a 20 base pair DNA double helix, and each vertex is a three-arm junction.
DNA nanotechnology seeks to make artificial, designed nanostructures out of nucleic acids, such as this DNA tetrahedron.[1] Each edge of the tetrahedron is a 20 base pair DNA double helix, and each vertex is a three-arm junction.
DNA nanotechnology is a branch of nanotechnology which uses the molecular recognition properties of DNA and other nucleic acids to create designed, artificial structures out of DNA for technological purposes. In this field, DNA is used as a structural material rather than as a carrier of genetic information, making it an example of bionanotechnology. DNA nanotechnology has applications in molecular self-assembly and in DNA computing.
Although DNA is usually considered in the context of molecular biology as the carrier of genetic information in living cells, DNA nanotechnology considers DNA solely as a chemical and as a material, and is usually pursued outside of any biological context. DNA nanotechnology makes use of the fact that, due to the specificity of Watson-Crick base pairing, only portions of the strands which are complementary to each other will bind to each other to form duplex DNA. DNA nanotechnology attempts to rationally design sets of DNA strands so that desired portions of each strand will assemble in the correct positions for some desired target structure, a process called nucleic acid design.
Although the field is usually called DNA nanotechnology, its principles apply equally well to other nucleic acids such as RNA and PNA, and structures incorporating these have been made. For this reason the field is occasionally referred to as nucleic acid nanotechnology.
Contents
Fundamental concepts
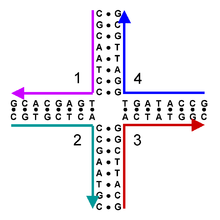 These four strands associate into a DNA four-arm junction because this structure maximizes the number of correct base pairs, with A's matched to T's and C's matched to G's.[2] See this image for a more realistic model of the four-arm junction showing its tertiary structure.
These four strands associate into a DNA four-arm junction because this structure maximizes the number of correct base pairs, with A's matched to T's and C's matched to G's.[2] See this image for a more realistic model of the four-arm junction showing its tertiary structure.
 A double-crossover (DX) molecule. This molecule consists of five DNA single strands which form two double-helical domains, on the left and the right in this image. There are two crossover points where the strands cross from one domain into the other.[2]
A double-crossover (DX) molecule. This molecule consists of five DNA single strands which form two double-helical domains, on the left and the right in this image. There are two crossover points where the strands cross from one domain into the other.[2]
DNA nanotechnology creates complex structures out of nucleic acids by making use of the specificity of base pairing in nucleic acid molecules. The structure of a nucleic acid molecule consists of a sequence of nucleotides, distinguished by which nucleobase they contain. In DNA, the four bases used are adenine (A), cytosine (C), guanine (G), and thymine (T). Nucleic acids have the property that two molecules will bind to each other to form a double helix only if the two sequences are complementary, meaning that they form matching sequences of base pairs, with A's only binding to T's, and C's only to G's. Because the formation of correctly matched base pairs is energetically favorable, nucleic acid strands are expected in most cases to bind to each other in the conformation that maximizes the number of correctly paired bases. This property, that the sequence determines the pattern of binding and the overall structure, is used by the field of DNA nanotechnology in that sequences are artificially designed so that a desired structure is favored to form.[3]
Nearly all structures in DNA nanotechnology make use of branched DNA structures containing junctions, as opposed to most biological DNA which exists in a linear double helix form. One of the simplest branched structures, and the first made, is a four-arm junction which can be made using four individual DNA strands which are complementary to each other in the correct pattern. Unlike in natural Holliday junctions, in the artificial immobile four-arm junction shown below, the base sequence of each arm is different, meaning that the junction point is fixed in a certain position.[3]
Junctions can be used in more complex molecules. One of the more widely used of these is the "double-crossover" or DX motif. A DX molecule can be thought of as two DNA duplexes positioned parallel to each other, with two crossover points where strands cross from one duplex into the other. Each junction point is itself topologically a four-arm junction. This molecule has the advantage that the junction points are now constrained to a single orientation as opposed to being flexible as in the four-arm junction. This makes the DX motif suitable as a structural building block for larger DNA complexes.[3]
Design
DNA nanostructures must be rationally designed so that the individual nucleic acid strands will assemble into the desired structures. The design process of such nanostructures usually begins with the specification of a desired target structure and/or functionality. Then, the overall secondary structure of the target molecule is designed, meaning the arrangement of nucleic acid strands within the structure, and which portions of those strands should be bound to each other. The last step is the primary structure design, the specification of the actual base sequences of each nucleic acid strand.[4]
Structural design
The first step in designing a nucleic acid nanostructure is to decide how a given structure should be represented by a specific arrangement of nucleic acid strands. This design step thus determines the secondary structure, or the series of base pairs which hold the individual strands together in the desired shape. There are several approaches which have been demonstrated:
- Tile-based structures. This approach breaks the target structure into smaller units with strong binding between the strands contained in each unit, and relatively weaker interactions between the units. It is often used to make periodic lattices, but can also be used to implement algorithmic self-assembly, making them one platform for DNA computing.[5]
- Folding structures. An alternative to the tile-based approach, folding approaches make the nanostructure out of a single long strand. This long strand can either have a designed sequence which folds due to its interactions with itself, or it can be folded into the desired shape by using shorter, "staple" strands. This latter method is called DNA origami, which allows the creation of two- and three-dimensional shapes at the nanoscale using DNA (see #Arbitrary shapes below).[6]
- Kinetic assembly. Recently, there has been interest in controlling the kinetics of DNA self-assembly, so that transient dynamics can also be programmed into the assembly. Such a method also has the advantage of proceeding isothermally and thus not requiring a thermal annealing step required by solely thermodynamic approaches.[7]
Sequence design
Main article: Nucleic acid designAfter any of the above approaches are used to design the secondary structure of a target molecule, an actual sequence of nucleotides must be devised which will form into the desired structure. Nucleic acid design is the process of assigning a specific nucleic acid base sequence to each strand so that they will associate into a desired conformation. Nucleic acid design is central to the field of DNA nanotechnology. Most methods seek to designing sequences so that the target structure is a thermodynamic minimum, and mis-assembled structures have higher energies and are thus disfavored. This is done either through heuristic methods such as sequence symmetry minimization and coding theory based approaches, or by explicitly using a full nearest-neighbor thermodynamic model. Geometric models are also used to examine tertiary structure of the nanostructures and ensure that the complexes are not overly strained.[4][8]
Nucleic acid design has similar goals to protein design: in both, the sequence of monomers is designed to favor the desired folded or associated structure and to disfavor alternate structures. Nucleic acid design has the advantage of being a much computationally simpler problem, since the simplicity of Watson-Crick base pairing rules leads to simple heuristic methods which yield experimentally robust designs. However, nucleic acid structures are less versatile than proteins in their functionality.[8]
Structural DNA nanotechnology
Structural DNA nanotechnology, sometimes abbreviated as SDN, focuses on synthesizing and characterizing nucleic acid complexes and materials with various nanoscale structures. Structural DNA nanotechnology is largely based on the fact that the three-dimensional structure of DNA—the nucleic acid double helix— has a robust, defined geometry which makes it possible to predict and design the structures of more complex DNA molecules. Many such structures have been created, including two- and three-dimensional structures; and periodic, aperiodic, and discrete structures.[9]
Periodic lattices
 Assembly of a DX array. Left, schematic diagram. Each bar represents a double-helical domain of DNA, with the shapes representing complimentary sticky ends. The DX molecule at top will combine into the two-dimensional DNA array shown at bottom.[2] Right, an atomic force microscope image of the assembled array. The individual DX tiles are clearly visible within the assembled structure. The field is 150 nm across.
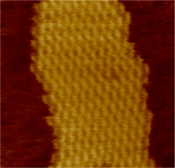
Assembly of a DX array. Left, schematic diagram. Each bar represents a double-helical domain of DNA, with the shapes representing complimentary sticky ends. The DX molecule at top will combine into the two-dimensional DNA array shown at bottom.[2] Right, an atomic force microscope image of the assembled array. The individual DX tiles are clearly visible within the assembled structure. The field is 150 nm across.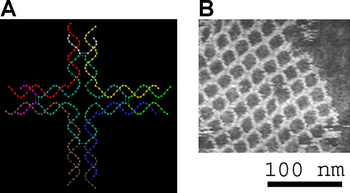 Left, a model of a DNA tile used to make another two-dimensional periodic lattice. Right, an atomic force micrograph of the assembled lattice.[10]
Left, a model of a DNA tile used to make another two-dimensional periodic lattice. Right, an atomic force micrograph of the assembled lattice.[10]
Smaller nucleic acid assemblies can be equipped with sticky ends in order to combine them into a two-dimensional periodic lattice. The earliest example of this was the array of DX, or double-crossover, molecules. Each DX molecule can be designed with four sticky ends, one at each end of the two double-helical domains, and these sticky ends can be designed with sequences that cause the DX units to combine into a specific tessellated pattern. They thus form extended flat sheets which are essentially rigid two-dimensional crystals of DNA.[11][12]
Two-dimensional arrays have been made out of other motifs as well, including the Holliday junction rhombus array[13] as well as various DX-based arrays making use of a double-cohesion scheme.[14][15]
Creating three-dimensional lattices out of DNA was the earliest goal of DNA nanotechnology, but proved to be one of the most difficult to realize. Success in constructing three-dimensional DNA lattices was finally reported in 2009 using a motif based on the concept of tensegrity, a balance between tension and compression forces.[16]
Nanotubes
In addition to flat sheets, DX arrays have been made to form hollow nanotubes of 4–20 nm diameter. These DNA nanotubes are somewhat similar in size and shape to carbon nanotubes, but the carbon nanotubes are stronger and better conductors, whereas the DNA nanotubes are more easily modified and connected to other structures. There have been multiple schemes for constructing DNA nanotubes, one of which uses the inherent curvature of DX tiles to form a DX lattice to curl around itself and close into a tube.[17] An alternative design uses single-stranded "tiles" for which the rigidity of the tube is an emergent property. This method also has the benefit of being able to determine the circumference of the nanotube in a simple, modular fashion.[18]
Polyhedra
A number of three-dimensional DNA molecules have been made which have the connectivity of a polyhedron such as an octahedron or cube. In other words, the DNA duplexes trace the edges of a polyhedron with a DNA junction at each vertex. The earliest demonstrations of DNA polyhedra involved multiple ligations and solid-phase synthesis steps to create catenated polyhedra.[19] More recent work has yielded polyhedra whose synthesis is much easier. These include a DNA octahedron made from a long single strand designed to fold into the correct conformation,[20] as well as a tetrahedron which can be produced from four DNA strands in a single step, pictured at the top of this article.[1]
Arbitrary shapes
See also: DNA origamiNanostructures of arbitrary shapes are usually made using the DNA origami method. DNA origami makes use of a long natural virus strand as a "scaffold" strand, and computationally designs shorter "staple" strands which bind to portions of the scaffold strand and force it to fold into the desired shape. This method has the advantage of being easy to design, as the base sequence is predetermined by the scaffold strand sequence, and it also does not require high strand purity and accurate stoichiometry, as most other DNA nanotechnology methods do. DNA origami was first demonstrated for two-dimensional shapes; demonstrated designs included the smiley face and a coarse map of North America.[6] This was later extended to solid three-dimensional shapes.[21]
In addition, structures have been constructed with two-dimensional faces which fold into an overall three-dimensional shape, akin to a cardboard box. These can be programmed to open and release their cargo in response to a stimulus, making them potentially useful as programmable molecular cages.[22][23]
Functional nucleic acid nanostructures
DNA nanotechnology focuses on creating molecules with designed functionalities as well as structures. These include both dynamic functionality within the nucleic acid structure itself, for example with computation and mechanical motion, as well as by including other components such as small molecules or nanoparticles which have their own functionalities. Many classes of functional systems have been demonstrated.
Nanoarchitecture
The idea of using DNA arrays to template the assembly of other functional molecules was first suggested by Nadrian Seeman in 1987,[24] but only recently has progress been made in reducing these kinds of schemes to practice. In 2006, researchers covalently attached gold nanoparticles to a DX-based tile and showed that self-assembly of the DNA structures also assembled the nanoparticles hosted on them.[25] Also that year, Dwyer and LaBean demonstrated the letters "D" "N" and "A" created on a 4x4 DX array using streptavidin,[26] and a hierarchical assembly based on this approach was also demonstrated that scales to larger arrays (8X8 and 8.96 MD).[27] A non-covalent hosting scheme was shown in 2007, using Dervan polyamides on a DX array to arrange streptavidin proteins on specific kinds of tiles on the DNA array. [28]
There has also been interest in using DNA nanotechnology to assemble molecular electronics devices. To this end, DNA has been used to assemble single walled carbon nanotubes into field-effect transistors.[29]
Algorithmic self-assembly
 DNA arrays that display a representation of the Sierpinski gasket on their surfaces. Click the image for further details.[30]
DNA arrays that display a representation of the Sierpinski gasket on their surfaces. Click the image for further details.[30] See also: DNA computing
See also: DNA computingDNA nanotechnology has been applied to the related field of DNA computing. The DX tiles can have their sticky end sequences chosen so that they act as Wang tiles, allowing them to perform computation. A DX array has been demonstrated whose assembly encodes an XOR operation; this allows the DNA array to implement a cellular automaton which generates a fractal called the Sierpinski gasket.[30] Another system has the function of a binary counter, displaying a representation of increasing binary numbers as it grows. These results show that computation can be incorporated into the assembly of DNA arrays, increasing its scope beyond simple periodic arrays.[31]
Note that DNA computing overlaps with, but is distinct from, DNA nanotechnology. The latter uses the specificity of Watson-Crick basepairing to make novel structures out of DNA. These structures can be used for DNA computing, but they do not have to be. Additionally, DNA computing can be realized without using the types of molecules made possible by DNA nanotechnology.
Nanomechanical devices
Main article: DNA machineDNA complexes have been made which change their conformation upon some stimulus. These are intended to have applications in nanorobotics. DNA machines have also been made which show a twisting motion. The first such device made use of the transition between the B-DNA and Z-DNA forms to respond to a change in buffer conditions.[32] This reliance on buffer conditions, however, caused all devices to change state at the same time. A subsequent system, called "molecular tweezers," changes from an open to a closed state based upon the presence of control strands, allowing multiple devices to be individually operated in solution.[33] This was followed up by another system which relies on the presence of control strands to switch from a paranemic-crossover (PX) conformation to a double-junction (JX2) conformation.[34]
Nucleic acid nanomachines have been made which exhibit directional motion along a linear track, called DNA walkers. A large number of schemes have been demonstrated.[35] One strategy is to control the motion of the walker along the track using control strands which need to be manually added in sequence.[36][37] Another approach is to make use of restriction enzymes or deoxyribozymes to cleave the strands and cause the walker to move forward, which has the advantage of running autonomously.[38][39] A later system extended the concept of DNA walkers to walk upon a two-dimensional surface rather than a linear track, and demonstrated the ability to selectively pick up and move molecular cargo.[40] Additionally, a linear walker has been demonstrated which performs DNA-templated synthesis as the walker advances along the track, allowing autonomous multistep chemical synthesis directed by the walker.[41]
Applications
DNA nanotechnology provides one of the only ways to form designed, complex structures with precise control over nanoscale features. The field is beginning to see application to solve basic science problems in structural biology and biophysics. One such application, the earliest envisaged for the field, is in crystallography, where molecules which are hard to crystallize by themselves could be arranged and oriented within a three-dimensional nucleic acid lattice, thus allowing determination of their structure. DNA origami rods have also been used to replace liquid crystals in residual dipolar coupling experiments in protein NMR spectroscopy; using DNA origami is advantageous because, unlike liquid crystals, they are tolerant of the detergents needed to suspend membrane proteins in solution. It has been demonstrated that DNA walkers can be used as nanoscale assembly lines to move nanoparticles and direct chemical synthesis. Furthermore, DNA origami structures have aided in the biophysical studies of enzyme function and protein folding.[9][42]
DNA nanotechnology is also moving towards potential real-world applications. It has been suggested that the ability of nucleic acid arrays to arrange other molecules has potential applications in molecular scale electronics, with the assembly of a nucleic acid lattice templating the assembly of molecular electronic elements such as molecular wires.[9] DNA nanotechnology has also been called a form of programmable matter because of the coupling of computation to its material properties.[43]
There are also potential applications for DNA nanotechnology in nanomedicine, making use of its ability to perform computation in a biocompatible format to make "smart drugs" for targeted drug delivery. One such system being investigated uses a hollow DNA box containing proteins which induce apoptosis, or cell death, which will only open when in proximity to a cancer cell.[42][44]
Materials and methods
The sequences of the individual DNA strands which make up the target structure are designed computationally, using molecular modeling and thermodynamic modeling software.[4][8] Once the sequences have been designed, the nucleic acid molecules themselves are synthesized through standard oligonucleotide synthesis methods. This process is usually automated by using a machine called an oligonucleotide synthesizer, and nucleic acids of custom sequence are commercially available from many vendors.[45] For methods which require pure strands of known concentration, the nucleic acid strands can be purified using denaturing gel electrophoresis,[46] and concentrations are determined by one of several nucleic acid quantitation methods using ultraviolet absorbance spectroscopy.[47]
The fully formed target structures are usually characterized by native gel electrophoresis, which provides information about the size and shape of DNA molecules. An electrophoretic mobility shift assay can indicate whether a stucture incorporates all the individual desired strands.[48] Fluorescent labeling and Förster resonance energy transfer (FRET) are also used to characterize the structure of the molecules.[49]
Nucleic acid structures can be directly imaged by atomic force microscopy, which is well-suited to extended two-dimensional structures, but is less useful for discrete three-dimensional structures due to the interaction of the microscope tip with the fragile nucleic acid structure. For these latter structures transmission electron microscopy and cryo-electron microscopy are important methods. Extended three-dimensional lattices are analyzed by X-ray crystallography.[50][51]
History
The concept of DNA nanotechnology was invented by Nadrian Seeman in the early 1980s.[52] Seeman was originally concerned with using a three-dimensional DNA lattice to orient target molecules, which would simplify their crystallographic study by eliminating the difficult process of obtaining pure crystals. This idea had reportedly come to him in fall 1980, after realizing the similarity between the M. C. Escher woodcut Depth and an array of DNA six-arm junctions.[3][53]
To this end, Seeman's laboratory in 1991 published the synthesis of a cube made of DNA, the first three-dimensional nanoscale object, for which he received the 1995 Feynman Prize in Nanotechnology, which was followed by a DNA truncated octahedron. However, it soon became clear that these molecules, polygonal shapes with flexible junctions as their vertices, were not rigid enough to form extended three-dimensional lattices. Seeman developed the more rigid double-crossover (DX) motif, and in collaboration with Erik Winfree, in 1998 published the creation of two-dimensional lattices of DX tiles.[3][52][54] The synthesis of a three-dimensional lattice was finally published by Seeman in 2009, nearly thirty years after he had set out to do so.[42]
These tile-based structures had the advantage that they provided the capability to implement DNA computing, which was demonstrated by Winfree and Paul Rothemund in their 2004 paper on the algorithmic self-assembly of a Sierpinski gasket structure, and for which they shared the 2006 Feynman Prize in Nanotechnology. Winfree's key insight was that the DX tiles could be used as Wang tiles, meaning that their assembly was capable of performing computation.[52]
The first DNA nanomachine—a motif which changes its structure in response to an input—was demonstrated in 1999 by Seeman. An improved system was demonstrated by Bernard Yurke the following year, which was also the first nucleic acid device to make use of toehold-mediated strand displacement. The next advance was to translate this into mechanical motion, and in 2004 and 2005, a number of DNA walker systems were demonstrated by the groups of Seeman, Niles Pierce, Andrew Turberfield, and Chengde Mao.[35]
In 2006, Rothemund first demonstrated the new DNA origami technique for easily and robustly creating folded DNA molecules of any shape. Rothemund had conceived of this method as being conceptually intermediate between Seeman's DX lattices, which used many short strands, and William Shih's DNA octahedron, which consisted mostly of one very long strand; Rothemund's DNA origami contains a long strand whose folding is assisted by a number of short strands. This method allowed the creation of much larger structures than were previously possible, which are less technically demanding to design and synthesize.[54] DNA origami was the cover story of Nature on March 15, 2006.[6] Rothemund's research demonstrating two-dimensional DNA origami structures was followed by the demonstration of solid three-dimensional DNA origami by Shih in 2009, while the labs of Jørgen Kjems and Hao Yan demonstrated hollow three-dimensional structures made out of two-dimensional faces.[42]
DNA nanotechnology as a field was initially met with some skepticism due to the unusual non-biological use of nucleic acids as a material for building structures and doing computation, as well as the preponderance of proof of principle experiments which extended the capabilities of the field but were far from actual applications. Seeman's seminal 1991 paper on the synthesis of the DNA cube was rejected by the journal Science after one reviewer praised its originality while another criticized it for its lack of biological relevance. By the early 2010s, however, the field was considered to have increased its capabilities to the point that applications for basic science research were beginning to be realized, and practical applications in medicine and other fields were beginning to be considered feasible.[42][55] In addition, the field had grown from very few active laboratories in 2001, to at least 60 in 2010, which increased the talent pool and thus the number of advancements in the field during that decade.[56]
See also
- Nucleic acid structure
- Molecular models of DNA
- List of nucleic acid simulation software
References
- ^ a b DNA polyhedra: Goodman, R.P.; Schaap, I.A.T.; Tardin, C.F.; Erben, C.M.; Berry, R.M.; Schmidt, C.F.; Turberfield, A.J. (9 December 2005). "Rapid chiral assembly of rigid DNA building blocks for molecular nanofabrication". Science 310 (5754): 1661–1665. Bibcode 2005Sci...310.1661G. doi:10.1126/science.1120367. PMID 16339440.
- ^ a b c Overview: Mao, Chengde (December 2004). "The Emergence of Complexity: Lessons from DNA". PLoS Biology 2 (12): 2036–2038. doi:10.1371/journal.pbio.0020431. PMC 535573. PMID 15597116. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=535573.
- ^ a b c d e Overview: Seeman, Nadrian C. (June 2004). "Nanotechnology and the double helix". Scientific American 290 (6): 64–75. doi:10.1038/scientificamerican0604-64. PMID 15195395. http://www.scientificamerican.com/article.cfm?id=nanotechnology-and-the-do.
- ^ a b c Sequence design: Brenneman, Arwen; Condon, Anne (2002). "Strand design for biomolecular computation". Theoretical Computer Science 287: 39. doi:10.1016/S0304-3975(02)00135-4.
- ^ Overview: Lin, Chenxiang; Liu, Yan; Rinker, Sherri; Yan, Hao (2006). "DNA Tile Based Self-Assembly: Building Complex Nanoarchitectures". ChemPhysChem 7 (8): 1641–7. doi:10.1002/cphc.200600260. PMID 16832805.
- ^ a b c DNA origami: Rothemund, Paul W. K. (2006). "Folding DNA to create nanoscale shapes and patterns". Nature 440 (7082): 297–302. Bibcode 2006Natur.440..297R. doi:10.1038/nature04586. PMID 16541064.
- ^ Kinetic assembly: Yin, Peng; Choi, Harry M. T.; Calvert, Colby R.; Pierce, Niles A. (2008). "Programming biomolecular self-assembly pathways". Nature 451 (7176): 318–22. Bibcode 2008Natur.451..318Y. doi:10.1038/nature06451. PMID 18202654.
- ^ a b c Sequence design: Dirks, Robert M.; Lin, Milo; Winfree, Erik & Pierce, Niles A. (2004). "Paradigms for computational nucleic acid design". Nucleic Acids Research 32 (4): 1392–1403. doi:10.1093/nar/gkh291. PMC 390280. PMID 14990744. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=390280.
- ^ a b c Structural DNA nanotechnology: Seeman, Nadrian C. (2007). "An Overview of Structural DNA Nanotechnology". Molecular Biotechnology 37 (3): 246–57. doi:10.1007/s12033-007-0059-4. PMID 17952671.
- ^ Other arrays: Strong, Michael (2004). "Protein Nanomachines". PLoS Biology 2 (3): e73. doi:10.1371/journal.pbio.0020073. PMC 368168. PMID 15024422. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=368168.
- ^ DX arrays: Winfree, Erik; Liu, Furong; Wenzler, Lisa A. & Seeman, Nadrian C. (6 August 1998). "Design and self-assembly of two-dimensional DNA crystals". Nature 394 (6693): 529–544. Bibcode 1998Natur.394..539W. doi:10.1038/28998. PMID 9707114.
- ^ DX arrays: Liu, Furong; Sha, Ruojie & Seeman, Nadrian C. (10 February 1999). "Modifying the Surface Features of Two-Dimensional DNA Crystals". Journal of the American Chemical Society 121 (5): 917–922. doi:10.1021/ja982824a.
- ^ Other arrays: Mao, Chengde; Sun, Weiqiong & Seeman, Nadrian C. (16 June 1999). "Designed Two-Dimensional DNA Holliday Junction Arrays Visualized by Atomic Force Microscopy". Journal of the American Chemical Society 121 (23): 5437–5443. doi:10.1021/ja9900398.
- ^ Other arrays: Constantinou, Pamela E.; Wang, Tong; Kopatsch, Jens; Israel, Lisa B.; Zhang, Xiaoping; Ding, Baoquan; Sherman, William B.; Wang, Xing; Zheng, Jianping; Sha, Ruojie & Seeman, Nadrian C. (2006). "Double cohesion in structural DNA nanotechnology". Organic and Biomolecular Chemistry 4 (18): 3414–3419. doi:10.1039/b605212f. PMID 17036134.
- ^ Other arrays: Mathieu, Frederick; Liao, Shiping; Kopatsch, Jens; Wang, Tong; Mao, Chengde & Seeman, Nadrian C. (April 2005). "Six-Helix Bundles Designed from DNA". Nano Letters 5 (4): 661–665. Bibcode 2005NanoL...5..661M. doi:10.1021/nl050084f. PMID 15826105.
- ^ Three-dimensional arrays: Zheng, Jianping; Birktoft, Jens J.; Chen, Yi; Wang, Tong; Sha, Ruojie; Constantinou, Pamela E.; Ginell, Stephan L.; Mao, Chengde et al. (2009). "From Molecular to Macroscopic via the Rational Design of a Self-Assembled 3D DNA Crystal". Nature 461 (7260): 74–7. Bibcode 2009Natur.461...74Z. doi:10.1038/nature08274. PMC 2764300. PMID 19727196. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2764300.
- ^ DNA nanotubes: Rothemund, Paul W. K.; Ekani-Nkodo, Axel; Papadakis, Nick; Kumar, Ashish; Fygenson, Deborah Kuchnir & Winfree, Erik (22 December 2004). "Design and Characterization of Programmable DNA Nanotubes". Journal of the American Chemical Society 126 (50): 16344–16352. doi:10.1021/ja044319l. PMID 15600335.
- ^ DNA nanotubes: Yin, P.; Hariadi, R. F.; Sahu, S.; Choi, H. M. T.; Park, S. H.; Labean, T. H.; Reif, J. H. (2008). "Programming DNA Tube Circumferences". Science 321 (5890): 824–826. Bibcode 2008Sci...321..824Y. doi:10.1126/science.1157312. PMID 18687961.
- ^ DNA polyhedra: Zhang, Yuwen; Seeman, Nadrian C. (1994). "Construction of a DNA-truncated octahedron". Journal of the American Chemical Society 116 (5): 1661–1669. doi:10.1021/ja00084a006.
- ^ DNA polyhedra: Shih, William M.; Quispe, Joel D.; Joyce, Gerald F. (12 February 2004). "A 1.7-kilobase single-stranded DNA that folds into a nanoscale octahedron". Nature 427 (6975): 618–621. doi:10.1038/nature02307. PMID 14961116.
- ^ DNA origami: Douglas, Shawn M.; Dietz, Hendrik; Liedl, Tim; Högberg, Björn; Graf, Franziska; Shih, William M. (2009). "Self-assembly of DNA into nanoscale three-dimensional shapes". Nature 459 (7245): 414–418. Bibcode 2009Natur.459..414D. doi:10.1038/nature08016. PMC 2688462. PMID 19458720. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2688462.
- ^ DNA boxes: Andersen, Ebbe S.; Dong, Mingdong; Nielsen, Morten M.; Jahn, Kasper; Subramani, Ramesh; Mamdouh, Wael; Golas, Monika M.; Sander, Bjoern et al. (2009). "Self-assembly of a nanoscale DNA box with a controllable lid". Nature 459 (7243): 73–6. Bibcode 2009Natur.459...73A. doi:10.1038/nature07971. PMID 19424153.
- ^ DNA boxes: Ke, Yonggang; Sharma, Jaswinder; Liu, Minghui; Jahn, Kasper; Liu, Yan; Yan, Hao (2009). "Scaffolded DNA Origami of a DNA Tetrahedron Molecular Container". Nano Letters 9 (6): 2445–7. Bibcode 2009NanoL...9.2445K. doi:10.1021/nl901165f. PMID 19419184.
- ^ Nanoarchitecture: Robinson, Bruche H.; Seeman, Nadrian C. (August 1987). "The Design of a Biochip: A Self-Assembling Molecular-Scale Memory Device". Protein Engineering 1 (4): 295–300. doi:10.1093/protein/1.4.295. PMID 3508280.
- ^ Nanoarchitecture: Zheng, Jiwen; Constantinou, Pamela E.; Micheel, Christine; Alivisatos, A. Paul; Kiehl, Richard A. & Seeman Nadrian C. (2006). "2D Nanoparticle Arrays Show the Organizational Power of Robust DNA Motifs". Nano Letters 6 (7): 1502–1504. Bibcode 2006NanoL...6.1502Z. doi:10.1021/nl060994c. PMID 16834438.
- ^ Nanoarchitecture: Park, Sung Ha; Sung Ha Park, Constantin Pistol, Sang Jung Ahn, John H. Reif, Alvin R. Lebeck, Chris Dwyer, Thomas H. LaBean (October 2006). "Finite-Size, Fully Addressable DNA Tile Lattices Formed by Hierarchical Assembly Procedures". Angewandte Chemie 118 (40): 749–753. doi:10.1002/ange.200690141.
- ^ Nanoarchitecture: Pistol, Constantin; Constantin Pistol, Chris Dwyer (March 2007). "Scalable, Low-cost, Hierarchical Assembly of Programmable DNA Nanostructures". Nanotechnology 18 (12): 125305–9. Bibcode 2007Nanot..18l5305P. doi:10.1088/0957-4484/18/12/125305.
- ^ Nanoarchitecture: Cohen, Justin D.; Sadowski, John P.; Dervan, Peter B. (2007). "Addressing Single Molecules on DNA Nanostructures". Angewandte Chemie 46 (42): 7956–7959. doi:10.1002/anie.200702767. PMID 17763481.
- ^ Nanoarchitecture: Maune, Hareem T.; Han, Si-Ping; Barish, Robert D.; Bockrath, Marc; Iii, William A. Goddard; Rothemund, Paul W. K.; Winfree, Erik (2009). "Self-assembly of carbon nanotubes into two-dimensional geometries using DNA origami templates". Nature Nanotechnology 5 (1): 61–6. Bibcode 2010NatNa...5...61M. doi:10.1038/nnano.2009.311. PMID 19898497.
- ^ a b Algorithmic self-assembly: Rothemund, Paul W. K.; Papadakis, Nick & Winfree, Erik (December 2004). "Algorithmic Self-Assembly of DNA Sierpinski Triangles". PLoS Biology 2 (12): 2041–2053. doi:10.1371/journal.pbio.0020424. PMC 534809. PMID 15583715. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=534809.
- ^ Algorithmic self-assembly: Barish, Robert D.; Rothemund, Paul W. K.; Winfree, Erik (2005). "Two Computational Primitives for Algorithmic Self-Assembly: Copying and Counting". Nano Letters 5 (12): 2586–2592. Bibcode 2005NanoL...5.2586B. doi:10.1021/nl052038l. PMID 16351220.
- ^ DNA machines: Mao, Chengde; Sun, Weiqiong; Shen, Zhiyong & Seeman, Nadrian C. (14 January 1999). "A DNA Nanomechanical Device Based on the B-Z Transition". Nature 397 (6715): 144–146. Bibcode 1999Natur.397..144M. doi:10.1038/16437. PMID 9923675.
- ^ DNA machines: Yurke, Bernard; Turberfield, Andrew J.; Mills, Allen P., Jr; Simmel, Friedrich C. & Neumann, Jennifer L. (10 August 2000). "A DNA-fuelled molecular machine made of DNA". Nature 406 (6796): 605–609. Bibcode 2000Natur.406..605Y. doi:10.1038/35020524. PMID 10949296.
- ^ DNA machines: Yan, Hao; Zhang, Xiaoping; Shen, Zhiyong & Seeman, Nadrian C. (3 January 2002). "A robust DNA mechanical device controlled by hybridization topology". Nature 415 (6867): 62–65. Bibcode 2002Natur.415...62Y. doi:10.1038/415062a. PMID 11780115.
- ^ a b DNA machines: Bath, Jonathan; Turberfield, Andrew J. (2007). "DNA nanomachines". Nature Nanotechnology 2 (5): 275–284. Bibcode 2007NatNa...2..275B. doi:10.1038/nnano.2007.104. PMID 18654284.
- ^ DNA walkers: Shin, JS; Pierce, NA (2004). "A synthetic DNA walker for molecular transport". Journal of the American Chemical Society 126 (35): 10834–5. doi:10.1021/ja047543j. PMID 15339155.
- ^ DNA walkers: Sherman, William B.; Seeman, Nadrian C. (2004). "A Precisely Controlled DNA Biped Walking Device". Nano Letters 4 (7): 1203–1207. Bibcode 2004NanoL...4.1203S. doi:10.1021/nl049527q.
- ^ DNA walkers: Tian, Ye; He, Yu; Chen, Yi; Yin, Peng; Mao, Chengde (2005). "A DNAzyme That Walks Processively and Autonomously along a One-Dimensional Track". Angewandte Chemie 117 (28): 4429–4432. doi:10.1002/ange.200500703.
- ^ DNA walkers: Bath, Jonathan; Green, Simon J.; Turberfield, Andrew J. (2005). "A Free-Running DNA Motor Powered by a Nicking Enzyme". Angewandte Chemie International Edition 44 (28): 4358–4361. doi:10.1002/anie.200501262.
- ^ Functional DNA walkers: Lund, Kyle; Manzo, Anthony J.; Dabby, Nadine; Michelotti, Nicole; Johnson-Buck, Alexander; Nangreave, Jeanette; Taylor, Steven; Pei, Renjun et al. (2010). "Molecular Robots Guided by Prescriptive Landscapes". Nature 465 (7295): 206–210. Bibcode 2010Natur.465..206L. doi:10.1038/nature09012. PMC 2907518. PMID 20463735. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2907518.
- ^ Functional DNA walkers: He, Yu; Liu, David R. (2010). "Autonomous Multistep Organic Synthesis in a Single Isothermal Solution Mediated by a DNA Walker". Nature Nanotechnology 5 (11): 778–782. Bibcode 2010NatNa...5..778H. doi:10.1038/nnano.2010.190. PMC 2974042. PMID 20935654. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2974042.
- ^ a b c d e History/applications: Service, Robert F. (3 June 2011). "DNA Nanotechnology Grows Up". Science 332 (6034): 1140–1143. doi:10.1126/science.332.6034.1140.
- ^ Applications: Rietman, Edward A. (2001). Molecular engineering of nanosystems. Springer. pp. 209–212. ISBN 978-0-387-98988-4. http://books.google.com/books?id=ga2DKYCm7xMC&pg=PA209. Retrieved 17 April 2011.
- ^ Applications: Jungmann, R; Renner, S; Simmel, FC (2008). "From DNA nanotechnology to synthetic biology". HFSP journal 2 (2): 99–109. doi:10.2976/1.2896331. PMC 2645571. PMID 19404476. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2645571.
- ^ Methods: Ellington, A.; Pollard, J. D. (2001). "Synthesis and Purification of Oligonucleotides". Current Protocols in Molecular Biology. doi:10.1002/0471142727.mb0211s42. ISBN 0471142727.
- ^ Methods: Ellington, A.; Pollard, J. D. (2001). "Purification of Oligonucleotides Using Denaturing Polyacrylamide Gel Electrophoresis". Current Protocols in Molecular Biology. doi:10.1002/0471142727.mb0212s42. ISBN 0471142727.
- ^ Methods: Gallagher, S. R.; Desjardins, P. (2011). "Quantitation of Nucleic Acids and Proteins". Current Protocols Essential Laboratory Techniques. doi:10.1002/9780470089941.et0202s5. ISBN 0470089938.
- ^ Methods: Chory, J.; Pollard, J. D. (2001). "Separation of Small DNA Fragments by Conventional Gel Electrophoresis". Current Protocols in Molecular Biology. doi:10.1002/0471142727.mb0207s47. ISBN 0471142727.
- ^ Methods: Walter, N. G. (2003). "Probing RNA Structural Dynamics and Function by Fluorescence Resonance Energy Transfer (FRET)". Current Protocols in Nucleic Acid Chemistry. doi:10.1002/0471142700.nc1110s11. ISBN 0471142700.
- ^ Methods: Lin, C.; Ke, Y.; Chhabra, R.; Sharma, J.; Liu, Y.; Yan, H. (2011). "Synthesis and Characterization of Self-Assembled DNA Nanostructures". In Zuccheri, G. and Samorì, B. DNA Nanotechnology: Methods and Protocols. Methods in Molecular Biology. 749. pp. 1–11. doi:10.1007/978-1-61779-142-0_1. ISBN 978-1-61779-141-3.
- ^ Methods: Bloomfield, Victor A.; Crothers, Donald M., Tinoco, Jr., Ignacio (2000). Nucleic acids: structures, properties, and functions. Sausalito, Calif: University Science Books. pp. 84–86, 396–407. ISBN 0935702490.
- ^ a b c History: Pelesko, John A. (2007). Self-assembly: the science of things that pu t thems elves together. New York: Chapman & Hall/CRC. pp. 201, 242, 259. ISBN 978 1 58488 687 7.
- ^ History: See Nadrian Seeman's homepage, Current crystallization protocol for a statement of the problem, and Nadrian Seeman's homepage, DNA cages containing oriented guests for the proposed solution.
- ^ a b DNA origami: Rothemund, Paul W. K. (2006). "Scaffolded DNA Origami: from Generalized Multicrossovers to Polygonal Networks". In Chen, Junghuei; Jonoska, Natasha; Rozenberg, Grzegorz. Nanotechnology: Science and Computation. Natural Computing Series. New York: Springer. pp. 3–21. doi:10.1007/3-540-30296-4_1. ISBN 978 3 540 30295 7.
- ^ History: Hopkin, Karen (August 2011). "Profile: 3-D Seer". The Scientist. http://the-scientist.com/2011/08/01/3-d-seer/. Retrieved 8 August 2011.
- ^ History: Seeman, Nadrian (2010). "Structural DNA Nanotechnology: Growing Along with NanoLetters". Nano Letters 10 (6): 1971–1978. Bibcode 2010NanoL..10.1971S. doi:10.1021/nl101262u. PMC 2901229. PMID 20486672. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2901229.
Further reading
Following is a list of review articles and books which can further introduce the reader to the field of DNA nanotechnology.
- Seeman, Nadrian C. (June 2004). "Nanotechnology and the double helix". Scientific American 290 (6): 64–75. doi:10.1038/scientificamerican0604-64. PMID 15195395. http://www.scientificamerican.com/article.cfm?id=nanotechnology-and-the-do.—An article written for laypeople by the founder of the field
- Pelesko, John A. (2007). "Chapter 8: DNA Self-Assembly". Self-assembly: the science of things that put themselves together. New York: Chapman & Hall/CRC. ISBN 978 1 58488 687 7.—An overview of the results of the field from the perspective of self-assembly
- Seeman, Nadrian C. (2007). "An Overview of Structural DNA Nanotechnology". Molecular Biotechnology 37 (3): 246–57. doi:10.1007/s12033-007-0059-4. PMID 17952671.—A review including some older results in the field
- Seeman, Nadrian (2010). "Structural DNA Nanotechnology: Growing Along with NanoLetters". Nano Letters 10 (6): 1971–1978. Bibcode 2010NanoL..10.1971S. doi:10.1021/nl101262u. PMC 2901229. PMID 20486672. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=2901229.—A review of more recent results in the period 2001–2010
- Feldkamp, Udo; Niemeyer, Christof M. (2006). "Rational Design of DNA Nanoarchitectures". Angewandte Chemie International Edition 45 (12): 1856–76. doi:10.1002/anie.200502358. PMID 16470892.—A review coming from the viewpoint of secondary structure design
- Lin, Chenxiang; Liu, Yan; Rinker, Sherri; Yan, Hao (2006). "DNA Tile Based Self-Assembly: Building Complex Nanoarchitectures". ChemPhysChem 7 (8): 1641–7. doi:10.1002/cphc.200600260. PMID 16832805.—A minireview specifically focusing on tile-based assembly
- Bath, Jonathan; Turberfield, Andrew J. (2007). "DNA nanomachines". Nature Nanotechnology 2 (5): 275–284. Bibcode 2007NatNa...2..275B. doi:10.1038/nnano.2007.104. PMID 18654284.—A review of nucleic acid nanomechanical devices
- Zhang, David Yu; Seelig, Georg (2011). "Dynamic DNA nanotechnology using strand-displacement reactions". Nature Chemistry 3 (2): 103–113. Bibcode 2011NatCh...3..103Z. doi:10.1038/nchem.957. PMID 21258382.—A review of DNA systems making use of strand displacement mechanisms
External links
- International Society for Nanoscale Science, Computation and Engineering
- What is Bionanotechnology?—a video introduction to DNA nanotechnology
Categories:
Wikimedia Foundation. 2010.


