- Pseudomonas
Taxobox
color = lightgrey
name = "Pseudomonas"
image_width = 280px
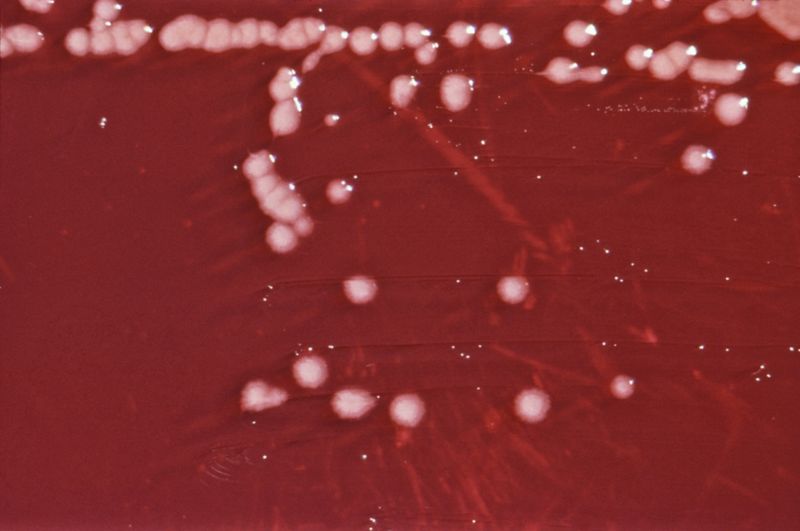
image_caption = "P. aeruginosa" colonies on anagar plate .
regnum = Bacteria
phylum =Proteobacteria
classis = Gamma Proteobacteria
ordo =Pseudomonadales
familia =Pseudomonadaceae
genus = "Pseudomonas"
genus_authority = Migula 1894
type_species = "Pseudomonas aeruginosa "
subdivision_ranks = Species
subdivision ="P. aeruginosa" group:"P. aeruginosa":"P. alcaligenes":"P. anguilliseptica":"P. argentinensis":"P. borbori":"P. citronellolis":"P. flavescens":"P. mendocina":"P. nitroreducens":"P. oleovorans":"P. pseudoalcaligenes":"P. resinovorans":"P. straminea""P. chlororaphis" group:"P. aurantiaca":"P. aureofaciens":"P. chlororaphis":"P. fragi":"P. lundensis":"P. taetrolens""P. fluorescens" group:"P. antarctica":"P. azotoformans":"'P. blatchfordae' ":"P. brassicacearum":"P. brenneri":"P. cedrina":"P. corrugata":"P. fluorescens":"P. gessardii":"P. libanensis":"P. mandelii":"P. marginalis":"P. mediterranea":"P. meridiana:"P. migulae":"P. mucidolens":"P. orientalis":"P. panacis":"P. proteolytica":"P. rhodesiae":"P. synxantha":"P. thivervalensis":"P. tolaasii":"P. veronii""P. pertucinogena" group:"P. denitrificans":"P. pertucinogena""P. putida" group:"P. cremoricolorata":"P. fulva":"P. monteilii":"P. mosselii":"P. oryzihabitans":"P. parafulva":"P. plecoglossicida":"P. putida""P. stutzeri" group:"P. balearica":"P. luteola":"P. stutzeri""P. syringae" group:"P. amygdali":"P. avellanae":"P. caricapapayae":"P. cichorii":"P. coronafaciens":"P. ficuserectae":"'P. helianthi'" :"P. meliae":"P. savastanoi":"P. syringae":"'P. tomato'":"P. viridiflava"incertae sedis :"P. abietaniphila":"P. acidophila":"P. agarici":"P. alcaliphila":"P. alkanolytica":"P. amyloderamosa":"P. asplenii":"P. azotifigens":"P. cannabina":"P. coenobios":"P. congelans":"P. costantinii":"P. cruciviae":"P. delhiensis":"P. excibis":"P. extremorientalis":"P. frederiksbergensis":"P. fuscovaginae":"P. gelidicola":"P. grimontii":"P. indica":"P. jessenii":"P. jinjuensis":"P. kilonensis":"P. knackmussii":"P. koreensis":"P. lini":"P. lutea":"P. moraviensis":"P. otitidis":"P. pachastrellae":"P. palleroniana":"P. papaveris":"P. peli":"P. perolens":"P. poae":"P. pohangensis":"P. psychrophila":"P. psychrotolerans":"P. rathonis:"P. reptilivora":"P. resiniphila":"P. rhizosphaerae":"P. rubescens":"P. salomonii":"P. segitis":"P. septica":"P. simiae":"P. suis":"P. thermotolerans":"P. tremae":"P. trivialis":"P. turbinellae":"P. tuticorinensis":"P. umsongensis":"P. vancouverensis":"P. vranovensis":"P. xanthomarina""Pseudomonas" is a
genus of gammaproteobacteria , belonging to the larger family ofpseudomonad s.Recently, 16S rRNA sequence analysis has redefined the taxonomy of many bacterial species.cite journal |author=Anzai Y, Kim H, Park, JY, Wakabayashi H |title=Phylogenetic affiliation of the pseudomonads based on 16S rRNA sequence |journal=Int J Syst Evol Microbiol |volume=50 |issue= |pages=1563–89 |year=2000 |pmid=10939664] As a result the genus "Pseudomonas" includes strains formerly classified in the genera "Chryseomonas" and "Flavimonas". [cite journal |title=The phylogeny of the genera "Chryseomonas", "Flavimonas", and "Pseudomonas" supports synonymy of these three genera |journal=Int J Syst Bacteriol |volume=47 |issue=2 |pages=249–51 |year=1997 |pmid=9103607 |unused_data=Anzai, "et al." ] Other strains previously classified in the genus "Pseudomonas" are now classified in the genera "
Burkholderia " and "Ralstonia ".History
"Pseudomonad" literally means 'false unit', being derived from the Greek "pseudo" ("ψευδο" 'false') and "monas" ("μονάς / μονάδα" 'a single unit'). The term "monad" was used in the early history of microbiology to denote single-celled organisms.
Because of their widespread occurrence in water, the
pseudomonad s were observed early in the history ofmicrobiology . The generic name "Pseudomonas" created for these organisms was defined in rather vague terms in 1894 as a genus of Gram-negative, rod-shaped and polar-flagella bacteria. Soon afterwards, a very large number of species was assigned to thegenus . Pseudomonads were isolated from many natural niches and a large number of species names was originally assigned to the genus. New methodology and the inclusion of approaches based on the studies of conservative macromolecules have reclassified many strains. cite book | author = Cornelis P (editor). | title = Pseudomonas: Genomics and Molecular Biology | edition = 1st ed. | publisher = Caister Academic Press | year = 2008 | url=http://www.horizonpress.com/pseudo | id = [http://www.horizonpress.com/pseudo ISBN 978-1-904455-19-6] ]"Pseudomonas aeruginosa" is increasingly recognized as an emerging
opportunistic pathogen of clinical relevance. Several different epidemiological studies indicate thatantibiotic resistance is increasing in clinical isolates [J. van Eldere. " [Multicentre surveillance of Pseudomonas aeruginosa susceptibility patterns in nosocomial infections http://jac.oxfordjournals.org/cgi/content/abstract/51/2/347] " J. Antimicrobial Chemotherapy (2003) 51 347-352] .In the year 2000, the complete
genome sequence of a "Pseudomonas" species was determined; more recently the sequence of other strains have been determined including "P. aeruginosa" strains PAO1 (2000), "P. putida" KT2440 (2002), "P. fluorescens" Pf-5 (2005), "P. syringae" pathovar tomato DC3000 (2003), "P. syringae" pathovar syringae B728a (2005), "P. syringae" pathovar phaseolica 1448A (2005), "P. fluorescens" PfO-1 and "P. entomophila" L48.cite book | author = Cornelis P (editor). | title = Pseudomonas: Genomics and Molecular Biology | edition = 1st ed. | publisher = Caister Academic Press | year = 2008 | url=http://www.horizonpress.com/pseudo | id = [http://www.horizonpress.com/pseudo ISBN 978-1-904455-19-6] ]An article published in the journal "Science" in 2008 showed that "Pseudomonas" may be the most common nucleator of ice crystals in clouds, thereby being of utmost importance to the formation of snow and rain around the world. [ [http://www.efluxmedia.com/news_Bacteria_The_Main_Ingredient_in_Snowflakes_Scientists_Say_14620.html Bacteria – The Main Ingredient in Snowflakes, Scientists Say ] ]
Characteristics
Members of the genus display the following defining characteristics:cite book | last = Krieg | first = Noel | title = Bergey's Manual of Systematic Bacteriology, Volume 1 | publisher = Williams & Wilkins | location = Baltimore | year = 1984 | isbn = 0683041088 ]
* Rod shaped
*Gram-negative
* One or more polarflagella , providingmotility
* Aerobic, although some species have been found to be facultative anaerobes (e.g. "P. aeruginosa")
* Non–spore forming
* Positivecatalase testOther characteristics which tend to be associated with "Pseudomonas" species (with some exceptions) include secretion of pyoverdin (
fluorescein ), a fluorescent yellow-greensiderophore cite journal | author = Meyer JM, Geoffroy VA, Baida N, "et al" | title = Siderophore typing, a powerful tool for the identification of fluorescent and nonfluorescent pseudomonads | journal = Appl. Environ. Microbiol. | volume = 68 | issue = 6 | pages = 2745–53 | year = 2002 | pmid = 12039729 | doi = 10.1128/AEM.68.6.2745-2753.2002| accessdate = 2007-05-27] under iron-limiting conditions. Certain "Pseudomonas" species may also produce additional types of siderophore, such as pyocyanin by "Pseudomonas aeruginosa "cite journal | author = Lau GW, Hassett DJ, Ran H, Kong F | title = The role of pyocyanin in Pseudomonas aeruginosa infection | journal = Trends in molecular medicine | volume = 10 | issue = 12 | pages = 599–606 | year = 2004 | pmid = 15567330 | doi = 10.1016/j.molmed.2004.10.002 | accessdate = 2007-05-27] and thioquinolobactin by "Pseudomonas fluorescens ",cite journal | author = Matthijs S, Tehrani KA, Laus G, Jackson RW, Cooper RM, Cornelis P | title = Thioquinolobactin, a Pseudomonas siderophore with antifungal and anti-Pythium activity | journal = Environ. Microbiol. | volume = 9 | issue = 2 | pages = 425–34 | year = 2007 | pmid = 17222140 | doi = 10.1111/j.1462-2920.2006.01154.x | accessdate = 2007-05-27] . "Pseudomonas" species also typically give a positive result to the oxidase test, the absence of gas formation from glucose, glucose is oxidised in oxidation/fermentation test using Hugh and Leifson O/F test, betahemolytic (on blood agar), indole negative,methyl red negative,Voges Proskauer test negative, citrate positive.The genus demonstrates a great deal of metabolic diversity, and consequently are able to colonise a wide range of nichescite book | author = Madigan M; Martinko J (editors). | title = Brock Biology of Microorganisms | edition = 11th ed. | publisher = Prentice Hall | year = 2005 | id = ISBN 0131443291 ] . Their ease of culture "
in vitro " and availability of an increasing number of "Pseudomonas" straingenome sequences has made the genus an excellent focus for scientific research; the best studied species include "P. aeruginosa" in its role as an opportunistic human pathogen, the plant pathogen "P. syringae", the soil bacterium "P. putida", and the plant growth promoting "P. fluorescens ".Biofilm formation
All
species and strains of "Pseudomonas" areGram-negative rods, and have historically been classified as strict aerobes. Exceptions to this classification have recently been discovered in "Pseudomonas" biofilmscite journal |author=Hassett D, Cuppoletti J, Trapnell B, Lymar S, Rowe J, Yoon S, Hilliard G, Parvatiyar K, Kamani M, Wozniak D, Hwang S, McDermott T, Ochsner U |title=Anaerobic metabolism and quorum sensing by "Pseudomonas aeruginosa" biofilms in chronically infected cystic fibrosis airways: rethinking antibiotic treatment strategies and drug targets |journal=Adv Drug Deliv Rev |volume=54 |issue=11 |pages=1425–43 |year=2002 |pmid=12458153 |doi=10.1016/S0169-409X(02)00152-7] . A significant number can produce exopolysaccharides that are known asslime layers . Secretion of exopolysaccharide makes it difficult for Pseudomonads to bephagocytose d by mammalianwhite blood cells .cite book | author = Ryan KJ; Ray CG (editors) | title = Sherris Medical Microbiology | edition = 4th ed. | publisher = McGraw Hill | year = 2004 | id = ISBN 0838585299 ] Slime production also contributes to surface-colonisingbiofilm s which are difficult to remove from food preparation surfaces. Growth of Pseudomonads on spoiling foods can generate a "fruity" odor."Pseudomonas" have the ability to metabolise a variety of diverse nutrients. Combined with the ability to form biofilms, they are thus able to survive in a variety of unexpected places. For example, they have been found in areas where pharmaceuticals are prepared. A simple carbon source, such as
soap residue or cap liner-adhesives is a suitable place for the Pseudomonads to thrive. Other unlikely places where they have been found includeantiseptic s such as quaternaryammonium compounds and bottledmineral water .Antibiotic resistance
Being Gram-negative bacteria, most "Pseudomonas spp." are naturally resistant to
penicillin and the majority of relatedbeta-lactam antibiotics , but a number are sensitive topiperacillin ,imipenem ,tobramycin , orciprofloxacin .This ability to thrive in harsh conditions is a result of their hardy
cell wall that contains porins. Their resistance to most antibiotics is attributed toefflux pumps calledABC transporters , which pump out some antibiotics before they are able to act."Pseudomonas aeruginosa" is a highly relevant
opportunistic pathogen . One of the most worrisome characteristics of "P. aeruginosa" consists in its lowantibiotic susceptibility. This low susceptibility is attributable to a concerted action ofmultidrug efflux pump s with chromosomally-encodedantibiotic resistance genes and the low permeability of the bacterial cellular envelopes. Besides intrinsic resistance, "P. aeruginosa" easily develop acquired resistance either bymutation in chromosomally-encoded genes, or by the horizontal gene transfer of antibiotic resistance determinants. Development ofmultidrug resistance by "P. aeruginosa" isolates requires several different genetic events that include acquisition of different mutations and/or horizontal transfer ofantibiotic resistance gene s. Hypermutation favours the selection of mutation-driven antibiotic resistance in "P. aeruginosa" strains producing chronic infections, whereas the clustering of several different antibiotic resistance genes inintegron s favours the concerted acquisition of antibiotic resistance determinants. Some recent studies have shown that phenotypic resistance associated tobiofilm formation or to the emergence of small-colony-variants may be important in the response of "P. aeruginosa" populations toantibiotic s treatment.cite book | author = Cornelis P (editor). | title = Pseudomonas: Genomics and Molecular Biology | edition = 1st ed. | publisher = Caister Academic Press | year = 2008 | url=http://www.horizonpress.com/pseudo | id = [http://www.horizonpress.com/pseudo ISBN 978-1-904455-19-6] ]Taxonomy
The studies on the
taxonomy of this complicatedgenus groped their way in the dark while following the classical procedures developed for the description and identification of theorganism s involved in sanitarybacteriology during the first decades of the twentieth century. This situation sharply changed with the proposal to introduce as the central criterion the similarities in the composition and sequences ofmacromolecule s components of theribosomal RNA . The new methodology clearly showed that thegenus "Pseudomonas", as classically defined, consisted in fact of a conglomerate of genera that could clearly be separated into five so-calledrRNA homology groups. Moreover, the taxonomic studies suggested an approach that might proved useful in taxonomic studies of all otherprokaryotic groups. A few decades after the proposal of the new genus "Pseudomonas" by Migula in 1894, the accumulation ofspecies names assigned to the genus reached alarming proportions. At the present moment, the number of species in the current list has contracted more than tenfold. In fact, this approximated reduction may be even more dramatic if one considers that the present list contains many new names, i.e., relatively few names of the original list survived in the process. The new methodology and the inclusion of approaches based on the studies of conservative macromolecules other thanrRNA components, constitutes an effective prescription that helped to reduce "Pseudomonas" nomenclatural hypertrophy to a manageable size.cite book | author = Cornelis P (editor). | title = Pseudomonas: Genomics and Molecular Biology | edition = 1st ed. | publisher = Caister Academic Press | year = 2008 | url=http://www.horizonpress.com/pseudo | id = [http://www.horizonpress.com/pseudo ISBN 978-1-904455-19-6] ]Pathogenicity
Animal pathogens
"P. aeruginosa" is an opportunistic human pathogen, most commonly affecting
immunocompromised patients, such as those withcystic fibrosis cite journal | author = Elkin S, Geddes D | title = Pseudomonal infection in cystic fibrosis: the battle continues | journal = Expert review of anti-infective therapy | volume = 1 | issue = 4 | pages = 609–18 | year = 2003 | pmid = 15482158 | doi = 10.1586/14787210.1.4.609| accessdate = 2007-05-27] orAIDS .cite journal | author = Shanson DC | title = Septicaemia in patients with AIDS | journal = Trans. R. Soc. Trop. Med. Hyg. | volume = 84 Suppl 1 | issue = | pages = 14–6 | year = 1990 | pmid = 2201108 | doi = | accessdate = 2007-05-27] Infection can affect many different parts of the body, but infections typically target the respiratory tract (e.g. patients with CF or those onmechanical ventilation ), causing bacterialpneumonia . Treatment of such infections can be difficult due to multiple antibiotic resistance.cite journal | author = McGowan JE | title = Resistance in nonfermenting gram-negative bacteria: multidrug resistance to the maximum | journal = Am. J. Med. | volume = 119 | issue = 6 Suppl 1 | pages = S29–36; discussion S62–70 | year = 2006 | pmid = 16735148 | doi = 10.1016/j.amjmed.2006.03.014 | accessdate = 2007-05-27]"P. oryzihabitans" can also be a human pathogen, although infections are rare. It can cause
peritonitis ,cite journal | author = Levitski-Heikkila TV, Ullian ME | title = Peritonitis with multiple rare environmental bacteria in a patient receiving long-term peritoneal dialysis | journal = Am. J. Kidney Dis. | volume = 46 | issue = 6 | pages = e119–24 | year = 2005 | pmid = 16310563 | doi = 10.1053/j.ajkd.2005.08.021 | accessdate = 2007-05-27]endophthalmitis ,cite journal | author = Yu EN, Foster CS | title = Chronic postoperative endophthalmitis due to Pseudomonas oryzihabitans | journal = Am. J. Ophthalmol. | volume = 134 | issue = 4 | pages = 613–4 | year = 2002 | pmid = 12383826 | doi = 10.1016/S0002-9394(02)01586-6| accessdate = 2007-05-27]septicemia andbacteremia . Similar symptoms although also very rare can be seen by infections of "P. luteola".cite journal | author = Kodama K, Kimura Nm Komagata K | title = Two new species of "Pseudomonas": "P. oryzihabitans" isolated from rice paddy and clinical specimens and "P. luteola" isolated from clinical specimens | journal = Int J Syst Bacteriol | volume = 35 | pages = 467–74 | year = 1985 ]"P. plecoglossicida" is a fish pathogenic species, causing hemorrhagic
ascites in theayu ("Plecoglossus altivelis").cite journal | author = Nishimori E, Kita-Tsukamoto K, Wakabayashi H | title = Pseudomonas plecoglossicida sp. nov., the causative agent of bacterial haemorrhagic ascites of ayu, Plecoglossus altivelis | journal = Int. J. Syst. Evol. Microbiol. | volume = 50 Pt 1 | issue = | pages = 83–9 | year = 2000 | pmid = 10826790 | doi = | accessdate = 2007-05-27] "P. anguilliseptica" is also a fish pathogen.cite journal | author = López-Romalde S, Magariños B, Ravelo C, Toranzo AE, Romalde JL | title = Existence of two O-serotypes in the fish pathogen Pseudomonas anguilliseptica | journal = Vet. Microbiol. | volume = 94 | issue = 4 | pages = 325–33 | year = 2003 | pmid = 12829386 | doi = 10.1016/S0378-1135(03)00124-X| accessdate = 2007-05-27]Due to their hemolytic activity, even non-pathogenic species of "Pseudomonas" can occasionally become a problem in clinical settings, where they have been known to infect blood transfusions.cite journal | author = Khabbaz RF, Arnow PM, Highsmith AK, "et al" | title = Pseudomonas fluorescens bacteremia from blood transfusion | journal = Am. J. Med. | volume = 76 | issue = 1 | pages = 62–8 | year = 1984 | pmid = 6419604 | doi = 10.1016/0002-9343(84)90751-4| accessdate = 2007-05-27]
Plant pathogens
"P. syringae" is a prolific
plant pathogen . It exists as over 50 different pathovars, many of which demonstrate a high degree of host plant specificity. There are numerous other "Pseudomonas" species that can act as plant pathogens, notably all of the other members of the "P. syringae" subgroup, but "P. syringae" is the most widespread and best studied.Although not strictly a plant pathogen, "P. tolaasii" can be a major agricultural problem, as it can cause bacterial blotch of cultivated
mushrooms .cite journal | author = Brodey CL, Rainey PB, Tester M, Johnstone K | title = Bacterial blotch disease of the cultivated mushroom is caused by an ion channel forming lipodepsipeptide toxin | journal = Molecular Plant–Microbe Interaction | year = 1991 | volume = 1 | pages = 407–11 | url = ] . Similarly, "P. agarici" can cause drippy gill in cultivated mushrooms.cite journal | author = Young JM | title = Drippy gill: a bacterial disease of cultivated mushrooms caused by "Pseudomonas agarici" n. sp | journal = NZ J Agric Res | year = 1970 | volume = 13 | pages = 977–90 | url = ] .Use as biocontrol agents
Since the mid 1980s, certain members of the "Pseudomonas" genus have been applied to cereal seeds or applied directly to soils as a way of preventing the growth or establishment of crop pathogens. This practice is generically referred to as
biocontrol . The biocontrol properties of "P. fluorescens" strains (CHA0 or Pf-5 for example) are currently best understood, although it is not clear exactly how the plant growth promoting properties of "P. fluorescens" are achieved. Theories include: that the bacteria might induce systemic resistance in the host plant, so it can better resist attack by a true pathogen; the bacteria might out compete other (pathogenic) soil microbes, e.g. bysiderophore s giving a competitive advantage at scavenging for iron; the bacteria might produce compounds antagonistic to other soil microbes, such asphenazine -type antibiotics orhydrogen cyanide . There is experimental evidence to support all of these theories, in certain conditions; a good review of the topic is written by Haas and Defago [cite journal |author=Haas D, Defago G |title=Biological control of soil-borne pathogens by fluorescent pseudomonads |journal=Nature Reviews in Microbiology |volume=3 |issue=4 |pages=307–19 |year=2005 |pmid=15759041 |doi=10.1038/nrmicro1129] .Other notable "Pseudomonas" species with biocontrol properties include "P. chlororaphis" which produces a
phenazine typeantibiotic active agent against certain fungal plant pathogens [cite journal |author=Chin-A-Woeng TF, "et al." |title=Root colonization by phenazine-1-carboxamide-producing bacterium "Pseudomonas chlororaphis" PCL1391 is essential for biocontrol of tomato foot and root rot |journal=Mol Plant Microbe Interact |volume=13 |issue=12 |pages=1340–5 |year=2000 |pmid=11106026 |doi=10.1094/MPMI.2000.13.12.1340] , and the closely related species "P. aurantiaca" which produces di-2,4-diacetylfluoroglucylmethan, a compoundantibiotic ally active againstGram-positive organisms [cite journal |author=Esipov, "et al." |title=New antibiotically active fluoroglucide from "Pseudomonas aurantiaca" |journal=Antibiotiki |volume=20 |issue=12 |pages=1077–81 |year=1975 |pmid=1225181 ] .Use as bioremediation agents
Some members of the genus "Pseudomonas" are able to metabolise chemical pollutants in the environment, and as a result can be used for
bioremediation . Notable species demonstrated as suitable for use as bioremediation agents include:* "P. alcaligenes", which can degrade
polycyclic aromatic hydrocarbon s.cite journal | author = O'Mahony MM, Dobson AD, Barnes JD, Singleton I | title = The use of ozone in the remediation of polycyclic aromatic hydrocarbon contaminated soil | journal = Chemosphere | volume = 63 | issue = 2 | pages = 307–14 | year = 2006 | pmid = 16153687 | doi = 10.1016/j.chemosphere.2005.07.018 | accessdate = 2007-05-27]
* "P. mendocina", which is able to degradetoluene .cite journal | author = Yen KM, Karl MR, Blatt LM, "et al" | title = Cloning and characterization of a Pseudomonas mendocina KR1 gene cluster encoding toluene-4-monooxygenase | journal = J. Bacteriol. | volume = 173 | issue = 17 | pages = 5315–27 | year = 1991 | pmid = 1885512 | doi = | accessdate = 2007-05-27]
* "P. pseudoalcaligenes" is able to usecyanide as anitrogen source.cite journal | author = Huertas MJ, Luque-Almagro VM, Martínez-Luque M, "et al" | title = Cyanide metabolism of Pseudomonas pseudoalcaligenes CECT5344: role of siderophores | journal = Biochem. Soc. Trans. | volume = 34 | issue = Pt 1 | pages = 152–5 | year = 2006 | pmid = 16417508 | doi = 10.1042/BST0340152 | accessdate = 2007-05-27]
* "P. resinovorans" can degradecarbazole .cite journal | author = Nojiri H, Maeda K, Sekiguchi H, "et al" | title = Organization and transcriptional characterization of catechol degradation genes involved in carbazole degradation by Pseudomonas resinovorans strain CA10 | journal = Biosci. Biotechnol. Biochem. | volume = 66 | issue = 4 | pages = 897–901 | year = 2002 | pmid = 12036072 | doi = 10.1271/bbb.66.897| accessdate = 2007-05-27]
* "P. veronii" has been shown to degrade a variety of simplearomatic organic compound s. [cite journal |author=Nam, "et al." |title=A novel catabolic activity of "Pseudomonas veronii" in biotransformation of pentachlorophenol |journal=Applied Microbiology and Biotechnology |volume=62 |pages=284–90 |year=2003 |pmid=12883877 |doi=10.1007/s00253-003-1255-1] [cite journal |author=Onaca, "et al." |title=Degradation of alkyl methyl ketones by "Pseudomonas veronii" |journal=Journal of Bacteriology |year=2007 Mar 9 |pmid=17351032 |unused_data=| [Epub ahead of print] ]
* "P. putida" has the ability to degrade organic solvents such astoluene .cite journal | author = Marqués S, Ramos JL | title = Transcriptional control of the Pseudomonas putida TOL plasmid catabolic pathways | journal = Mol. Microbiol. | volume = 9 | issue = 5 | pages = 923–9 | year = 1993 | pmid = 7934920 | doi = 10.1111/j.1365-2958.1993.tb01222.x| accessdate = 2007-05-27] At least one strain of this bacterium is able to convertmorphine in aqueous solution into the stronger and somewhat expensive to manufacture drughydromorphone (Dilaudid).
* Strain KC of "P. stutzeri" is able to degradecarbon tetrachloride . [cite journal |author=Sepulveda-Torres, "et al." |title=Generation and initial characterization of "Pseudomonas stutzeri" KC mutants with impaired ability to degrade carbon tetrachloride |journal=Arch Microbiol |volume=171 |issue=6 |pages=424–9 |year=1999 |pmid=10369898 |doi=10.1007/s002030050729]Food spoilage agents
As a result of their metabolic diversity, ability to grow at low temperatures and ubiquitous nature, many "Pseudomonas" can cause food spoilage. Notable examples include dairy spoilage by "P. fragi", [cite journal |author=Pereira, JN, and Morgan, ME |title=Nutrition and physiology of "Pseudomonas fragi" |journal=J Bacteriol |volume=74 |issue=6 |pages=710–3 |year=1957 Dec |pmid=13502296 ] mustiness in eggs caused by "P. taetrolens" and "P. mudicolens", [cite journal |author=Levine, M, and Anderson, DQ |title=Two New Species of Bacteria Causing Mustiness in Eggs |journal=J Bacteriol |volume=23 |issue=4 |pages=337–47 |year=1932 Apr |pmid=16559557 ] and "P. lundensis", which causes spoilage of
milk ,cheese ,meat , and fish. [cite journal |author=Gennari, M, and Dragotto, F |title=A study of the incidence of different fluorescent "Pseudomonas" species and biovars in the microflora of fresh and spoiled meat and fish, raw milk, cheese, soil and water |journal=J Appl Bacteriol |volume=72 |issue=4 |pages=281–8 |year=1992 Apr |pmid=1517169 ]pecies previously classified in the genus
Recently, 16S rRNA sequence analysis redefined the taxonomy of many bacterial species previously classified as being in the "Pseudomonas" genus.cite journal |author=Anzai Y, Kim H, Park, JY, Wakabayashi H |title=Phylogenetic affiliation of the pseudomonads based on 16S rRNA sequence |journal=Int J Syst Evol Microbiol |volume=50 |issue= |pages=1563–89 |year=2000 |pmid=10939664] Species which moved from the "Pseudomonas" genus are listed below; clicking on a species will show its new classification. Note that the term 'Pseudomonad' does not apply strictly to just the "Pseudomonas" genus, and can be used to also include previous members such as the genera "
Burkholderia " and "Ralstonia ".α proteobacteria: "P. abikonensis", "P. aminovorans", "P. azotocolligans", "P. carboxydohydrogena", "P. carboxidovorans", "P. compransoris", "P. diminuta", "P. echinoides", "P. extorquens", "P. lindneri", "P. mesophilica", "P. paucimobilis", "P. radiora", "P. rhodos", "P. riboflavina", "P. rosea", "P. vesicularis".
β proteobacteria: "P. acidovorans", "P. alliicola", "P. antimicrobica", "P. avenae", "P. butanovorae", "P. caryophylli", "P. cattleyae", "P. cepacia", "P. cocovenenans", "P. delafieldii", "P. facilis", "P. flava", "P. gladioli", "P. glathei", "P. glumae", "P. graminis", "P. huttiensis", "P. indigofera", "P. lanceolata", "P. lemoignei", "P. mallei", "P. mephitica", "P. mixta", "P. palleronii", "P. phenazinium", "P. pickettii", "P. plantarii", "P. pseudoflava", "P. pseudomallei", "P. pyrrocinia", "P. rubrilineans", "P. rubrisubalbicans", "P. saccharophila", "P. solanacearum", "P. spinosa", "P. syzygii", "P. taeniospiralis", "P. terrigena", "P. testosteroni".
γ-β proteobacteria: "P. beteli", "P. boreopolis", "P. cissicola", "P. geniculata", "P. hibiscicola", "P. maltophilia", "P. pictorum".
γ proteobacteria: "P. beijerinckii", "P. diminuta", "P. doudoroffii", "P. elongata", "P. flectens", "P. halodurans", "P. halophila", "P. iners", "P. marina", "P. nautica", "P. nigrifaciens", "P. pavonacea", "P. piscicida", "P. stanieri".
δ proteobacteria: "P. formicans"...
References
ee also
*
Pseudomonas phage Φ6 External links
General
* [http://www.efluxmedia.com/news_Bacteria_The_Main_Ingredient_in_Snowflakes_Scientists_Say_14620.html Pseudomonas at origin of world's rain and snow]
* [http://www.time.com/time/magazine/article/0,9171,894282,00.html?promoid=googlep Pseudomonas survive in nuclear reactor]
* [http://www.pseudomonas.com "Pseudomonas" genome database]
* [http://www.horizonpress.com/gateway/pseudomonas.html "Pseudomonas"]
* Fluorescent Pseudomonas [http://www.tgw1916.net/movies.html video]
Wikimedia Foundation. 2010.
