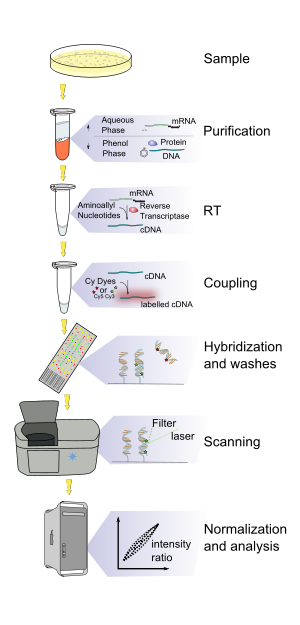
- DNA microarray experiment
-
For DNA microarrays in general, see DNA microarray.
This is an example of a DNA microarray experiment, detailing a particular case to better explain DNA microarray experiments, while enumerating possible alternatives.
- The two samples to be compared (pairwise comparison) are grown/acquired. In this example treated sample (case) and untreated sample (control).
- The nucleic acid of interest is purified: this can be all RNA for expression profiling, DNA for comparative hybridization, or DNA/RNA bound to a particular protein which is immunoprecipitated (ChIP-on-chip) for epigenetic or regulation studies. In this example total RNA is isolated (total as it is nuclear and cytoplasmic) by Guanidinium thiocyanate-phenol-chloroform extraction (e.g. Trizol) which isolates most RNA (whereas column methods have a cut off of 200 nucleotides) and if done correctly has a better purity.
- The purified RNA is analysed for quality (by capillary electrophoresis) and quantity (by using a spectrophotometer like NanoPhotometerTM or nanodrop): if enough material (>1μg) is present the experiment can continue.
- The labelled product is generated via reverse transcription and sometimes with an optional PCR amplification. The RNA is reverse transcribed with either polyT primers which amplify only mRNA or random primers which amplify all RNA which is mostly rRNA, miRNA microarray ligate an oligonucleotide to the purified small RNA (isolated with a fractionator) and then RT and amplified. The label is added either in the RT step or in an additional step after amplification if present. The sense that is labelled depends on the microarray, which means that if the label is added with the RT mix, the cDNA is on the template strand while the probe is on the sense strand (unless they are negative controls). The label is typically fluorescent; only one machine uses radiolabels. The labelling can be direct (not used) or indirect which requires a coupling stage. The coupling stage can occur before hybridization (two-channel arrays) using aminoallyl-UTP and NHS amino-reactive dyes (like cyanine dyes) or after (single-channel arrays) using biotin and labelled streptavin. The modified nucleotides (typically a 1 aaUTP: 4 TTP mix) are added enzymatically at a lower rate compared to normal nucleotides, typically resulting in 1 every 60 bases. The aaDNA is then purified with a column (using solution containing phosphate buffer as Tris contains amine groups). After purification, labelling efficiency (Frequency of Incorporation (FOI)) of each sample has to be measured with a small volume photometer (like the NanoPhotometerTM[1]) to ensure equal amounts of dye incorporation to the samples to be compared. The aminoallyl group is an amine group on a long linker attached to the nucleobase, which reacts with a reactive dye. A dye flip is a type of replicate done to remove any dye effects in two-channel dyes, in one slide one same is labeled with Cy3 the other with Cy5, this is reversed in a different slide. In this example, in the presence of aminoallyl-UTP added in the RT mix.
- The labeled samples are then mixed with a propriety hybridization solution which may contain SDS, SSC, dextran sulfate, a blocking agent (such as COT1 DNA, salmon sperm DNA, calf thymus DNA, PolyA or PolyT), Denhardt's solution and formamine.
- This mix is denatured and added to a pin hole in a microarray, which can be a gene chip (holes in the back) or a glass microarray which is bound by a cover, called a mixer, containing two pinholes and sealed with the slide at the perimeter.
- The holes are sealed and the microarray hybridized, either in a hyb oven, where the microarray is mixed by rotation, or in a mixer, where the microarray is mixed by alternating pressure at the pinholes.
- After an overnight hybridization, all nonspecific binding is washed off (SDS and SSC).
- The microarray is dried and scanned in a special machine where a laser excites the dye and a detector measures its emission.
- The image is gridded with a template and the intensities of the features (several pixels make a feature) are quantified.
- The raw data is normalized, the simplest way is to subtract the background intensity and then divide the intensities making either the total intensity of the features on each channel equal or the intensities of a reference gene and then the t-value for all the intensities is calculated. More sophisticated methods include z-ratio, loess and lowess regression and RMA (robust multichip analysis) for Affymetrix chips (single-channel, silicon chip, in situ synthesised short oligonucleotides).
See also
- DNA microarray
- Microfluidics or lab-on-chip
- Quantification of nucleic acids
- Cyanine dyes, such as Cy3 and Cy5
- Protocol (natural sciences)
- sample replication
- Molecular biology
- Serial analysis of gene expression
External links
Molecular biology Computational biology • Developmental biology • Functional Biology Overview Element(Genetic • Heredity)
Promoter (Pribnow box, TATA box) • Operon (gal operon, lac operon, trp operon) • Intron • Exon • Terminator • Enhancer • Repressor (lac repressor, trp repressor) • Silencer • Histone methylationLinked LifeEngineering Conceptmitosis • cell signalling • Post-transcriptional modification and Post-translational modification • Dry Lab/Wet labTechniqueCell culture • model organisms (such as C57BL/6 mice) • method (Nucleic acid • Protein) • Fluorescence, Pigment & Radioactivity
- High-throughput Technique (-omics): DNA microarray • Mass spectrometry • Lab-on-a-chip
Glossary References
- Gibson and Muse, A primer of genome science etc ISBN 0-87893-232-1
- Chomczynski, P. & Sacchi, N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction:Twenty-something years on. Nature Prot. 1, 581–585 (2006).
- Sambrook and Russell (2001). Molecular Cloning: A Laboratory Manual, 3rd edition, Cold Spring Harbor Laboratory Press.
Lab protocols found on microarray labs: [1][2] [3][4]
- ^ Kartha, R. Spectrophotometric Quantification of Nano- and Standard-Volume Samples, (2008, October 7), American Biotechnology Laboratory, http://www.iscpubs.com/Media/PublishingTitles/b0608kar.pdf
Categories:- Molecular biology
- Gene expression
- Bioinformatics
- Glass coating and surface modification
- DNA
- Microarrays
Wikimedia Foundation. 2010.