- Dictyostelium discoideum
-
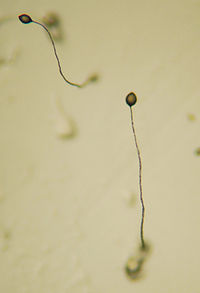
Dictyostelium discoideum Fruiting bodies of D. discoideum Scientific classification Domain: Eukarya Kingdom: Amoebozoa Superphylum: Conosa Phylum: Mycetozoa Class: Dictyostelia Order: Dictyosteliida Family: Dictyosteliidae Genus: Dictyostelium Species: D. discoideum Binomial name Dictyostelium discoideum
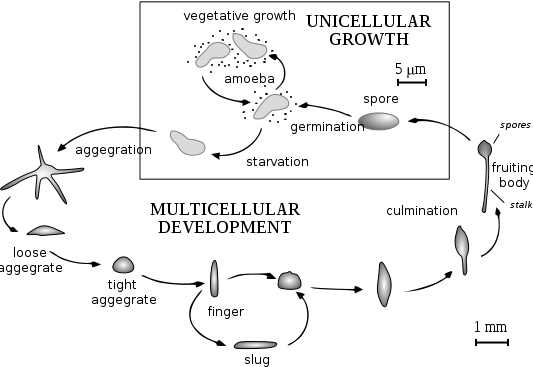
Raper, 1935[1]Dictyostelium discoideum is a species of soil-living amoeba belonging to the phylum Mycetozoa. D. discoideum, commonly referred to as slime mold, is a eukaryote that transitions from a collection of unicellular amoebae into a multicellular slug and then into a fruiting body within its lifetime. D. discoideum has a unique asexual lifecycle that consists of four stages: vegetative, aggregation, migration, and culmination. The life cycle of D. discoideum is relatively short, which allows for timely viewing of all life stages. The cells involved in the life cycle undergo movement, chemical signaling, and development, which are applicable to human cancer research. The simplicity of its life cycle makes D. discoideum a valuable model organism to study genetic, cellular, and biochemical processes in other organisms.
Contents
Natural habitat and diet
In the wild, D. discoideum can be found in soil and moist leaf litter. The primary diet of D. discoideum consists of bacteria, such as Escherichia coli, that are found in the soil and decaying organic matter. Uninucleate amoebae of D. discoideum consumes bacteria contiguous to its natural habitat, which includes deciduous forest soil and decaying leaves.[2]
Life cycle and reproduction
The life cycle of D. discoideum begins as spores are released from a mature sorocarp (fruiting body). Myxamoebae hatch from the spores under warm and moist conditions. During their vegetative stage, the myxamoebae divide by mitosis as they feed on bacteria. The bacteria secrete folic acid, attracting the myxamoebae. When the supply of bacteria is depleted, the myxamoebae enter the aggregation stage.
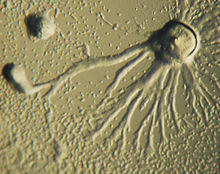
During aggregation, starvation initiates the creation of a biochemical machinery which includes glycoproteins and adenylyl cyclase.[3] The glycoproteins allow for cell-cell adhesion and adenylyl cyclase creates cyclic AMP. Cyclic AMP is secreted by the amoebas to attract neighboring cells to a central location. As they move towards the signal, they bump into each other and stick together by the use of glycoprotein adhesion molecules.
The migration stage begins once the amoebas have formed a tight aggregate and the elongated mound of cells tip over to lie flat on the ground. The amoebas work together as a motile pseudoplasmodium, also known as a slug. The slug is approximately 2–4 mm long and is capable of movement by producing a cellulose sheath in its anterior cells through which the slug moves.[4] Part of this sheath is left behind as a slimy trail as it moves towards attractants such as light, heat, and humidity in a forward-only direction.[4] Cyclic AMP and a substance called differentiation-inducing factor (DIF), help to form different cell types.[4] The slug becomes differentiated into prestalk and prespore cells that move to the anterior and posterior ends, respectively. Once the slug has found a suitable environment, the anterior end of the slug will form the stalk of the fruiting body and the posterior end will form the spores of the fruiting body.[4] Anterior-like cells, which have only been recently discovered, are also dispersed throughout the posterior region of the slug. These anterior-like cells form the very bottom of the fruiting body and the caps of the spores.[4] After the slug settles into one spot, the posterior end spreads out with the anterior end raised in the air, forming what is called the "Mexican hat," and the culmination stage begins.
The prestalk cells and prespore cells switch positions in the culmination stage in order to form the mature fruiting body.[4] The anterior end of the Mexican hat forms a cellulose tube, which allows the more posterior cells to move up the outside of the tube to the top, and the prestalk cells move down.[4] This rearrangement forms the stalk of the fruiting body made up of the cells from the anterior end of the slug, and the cells from the posterior end of the slug are on the top and now form the spores of the fruiting body. At the end of this 8–10 hours process, the mature fruiting body is fully formed.[4] This fruiting body is 1–2 mm tall and is now able to start the entire cycle over again by releasing the mature spores that become myxamoebae.
Generally reproducing asexually, D. discoideum are still capable of sexual reproduction if certain conditions are met. If two amoebae of different mating types are present in a dark and wet environment, they can fuse during aggregation to form a giant cell. The giant cell will then engulf the other cells in the aggregate and encase the whole aggregate in a thick, cellulose wall to protect it. This is known as a macrocyst. Inside the macrocyst, the giant cell divides first through meiosis, then through mitosis to produce many haploid amoebae that will be released to feed as normal amoebae would. While sexual reproduction is possible, it is very rare to see successful germination of a D. discoideum macrocyst under laboratory conditions.
Use as a model organism
Because of the simple life cycle of D. discoideum, it is commonly used as a model organism. It can be observed at organismic, cellular, and molecular levels primarily because of their restricted number of cell types, behaviors, and their rapid growth.[4] It is used to study cell differentiation, chemotaxis and programmed cell death, which are all normal cellular processes. It is also used to study other aspects of development including cell sorting, pattern formation, phagocytosis, motility, and signal transduction.[5] These processes and aspects of development are either absent or too difficult to view in other model organisms. D. discoideum is closely related to higher metazoans. It carries similar genes and pathways making it a good candidate for gene knockout.[6]
Cell differentiation is the process that occurs when a cell becomes more specialized to develop into a multicellular organism. Changes in size, shape, metabolic activities, and responsiveness can occur as a result of adjustments in gene expression. Cell diversity and differentiation, in this species, involves decisions made from cell-cell interactions in pathways to either stalk cells or spore cells.[7] These cell fates depend on their environment and pattern formation. Therefore, the organism is an excellent model for studying cell differentiation.
Chemotaxis is defined as a passage of an organism toward or away from a chemical stimulus along a chemical concentration gradient. Certain organisms demonstrate chemotaxis when they move toward a supply of nutrients. In D. discoideum, the amoeba secretes the signal, cAMP, out of the cell attracting other amoebas to migrate toward the source. Every amoeba moves toward a central amoeba, the one dispensing the greatest amount of cAMP secretions. The secretion of the cAMP is then exhibited by all amoebas and is a call for the amoebas to begin aggregation. These chemical emissions and amoeba movement occur every six minutes. The amoebas move toward the concentration gradient for sixty seconds and stop until the next secretion is sent out. This behavior of individual cells tends to cause oscillations in a group of cells, and chemical waves of varying cAMP concentration propagate through the group in spirals.[8]
The use of cAMP as a chemotactic agent is not established in any other organism. In developmental biology, this is one of the comprehensible examples of chemotaxis.[4]
Thermotaxis is movement along a gradient of temperature. The slugs have been shown to migrate along extremely shallow gradients of only 0.05C/cm. But the direction chosen is complicated; it seems to be away from a temperature about 2C below the temperature to which they had been acclimated. This complicated behavior has been analyzed by computer modeling of the behavior and the periodic pattern of temperature changes in soil caused by daily changes in air temperature. The conclusion is that the behavior moves slugs a few centimeters below the soil surface up to the surface. This is an amazingly sophisticated behavior by a primitive organism with no sense of gravity.[9]
Programmed cell death (apoptosis) is a normal part of species development.[3] Apoptosis is necessary for the proper spacing and sculpting of complex organs. Around 20% of cells in D. discoideum altruistically sacrifice themselves in the formation of the mature fruiting body. During the pseudoplasmodium (slug or grex) stage of its life cycle, the organism has formed three main types of cells: prestalk, prespore, and anterior-like cells. During culmination, the prestalk cells secrete a cellulose coat and extend as a tube through the grex.[3] As they differentiate, they form vacuoles and enlarge lifting up the prespore cells. The stalk cells undergo apoptosis and die as the prespore cells are lifted high above the substrate. The prespore cells then become spore cells; each one becoming a new myxamoeba upon dispersal.[4] This is an example of how apoptosis is used in the formation of a reproductive organ, the mature fruiting body.
A recent major contribution from Dictyostelium research has come from new techniques allowing the activity of individual genes to be visualised in living cells.[10] This has shown that transcription occurs in "bursts" or "pulses" (see transcriptional bursting) rather than following simple probabilistic or continuous behaviour. Bursting transcription now appears to be conserved between bacteria and humans. Another remarkable feature of the organism is that it has sets of DNA repair enzymes found in human cells which are lacking from many other popular metazoan model systems.[11] Defects in DNA repair leads to devastating human cancers, so the ability to study human repair proteins in a simple tractable model will prove invaluable.
Lab cultivation
D. discoideum's ability to be easily cultivated in the lab[4] adds to its appeal as a model organism. While D. discoideum can be grown in liquid culture, it is usually grown in Petri dishes containing nutrient agar and the surfaces are kept moist. The cultures grow best at 22o-24oC (room temperature). D. discoideum primarily feed on E. coli, which is adequate for all stages of the life cycle. When the food supply is diminished, the myxamoebae will aggregate to form pseudoplasmodia. Soon, the dish will be covered with various stages of the life cycle. Checking the dish often will allow for detailed observations of development. The cells can be harvested at any stage of development and grown quickly.
While cultivating in a lab, it is important to take into account D. discoideum's behavioral responses. For example, they have an affinity toward light, higher temperatures, high humidity, low ionic concentrations, and the acidic side of the pH gradient. Experiments are often done to see how manipulations of these parameters hinder, stop, or accelerate development. Variations of these parameters can alter the rate and viability of culture growth. Also the fruiting bodies, being that this is the tallest stage of development, are very responsive to air currents and physical stimuli. It is unknown if there is a stimulus involved with spore release.
"Farming"
A 2011 report in Nature published findings that demonstrated a "primitive farming behaviour" in D. discoideum colonies.[12][13] Described as a "symbiosis" between D. discoideum and bacterial prey, approximately one third of wild-collected D. discoideum colonies engaged in the "husbandry" of the bacteria when the bacteria were included within the slime mold fruiting bodies.[13] The incorporation of the bacteria into the fruiting bodies allows the "seeding" of the food source at the location of the spore dispersal, which is particularly valuable if the new location is low in food resources.[13] Colonies produced from the "farming" spores typically also show the same behavior when sporulating; colonies. This incorporation has a cost associated with it: those colonies that do not consume all of the prey bacteria produce smaller spores that cannot disperse as widely. Additionally, there is much less benefit for bacteria-containing spores that land in a food-rich region. This balance of the costs and benefits of the behavior may contribute to the fact that a minority of D. discoideum colonies engage in this practice.[12][13] D.discoideum is known for eating gram positive as well as gram negative bacteria. But some of the phagocited bacteria, including some human pathogens[14], are able to live in the amoebae and exit without killing the cell. When do they enter the cell? Where do they reside? When do they get out of the cell? The research is not yet conclusive but, making a little puzzle, it is possible to draw a general life cycle of Dictyostelium discoideum adapted for farmer clones in order to better understand this symbiotic process .

In the picture we can see the different stages. First in the starvation stageº, bacteria are enclosed within D. discoideum[14], after entry into amoebae, in a phagosome the fusion with lysosomes is blocked and these unmatured phagosomes are surrounded by host cell organelles such as mitochondria, vesicles, and a multilayer membrane derived from the rough endoplasmic reticulum (RER) of amoeba. The role of the RER in the intracellular infection is not known, but the RER is not required as a source of proteins for the bacteria[15]. The bacteria resides within these phagosomes during the aggregation and the multicellular development stages. The amoebae preserve her individuality and each amoeba has their own bacteria. During the culmination stage, when the spores are produced, the bacteria pass from the cell to the sorus with the help of a cytoskeletal structure that prevents host cell destruction[16]. Some results suggest that bacteria exploit the exocytosis without killing the cell[16]. Free-living amoebae seem to play a crucial role for persistence and dispersal of some pathogens in the environment. Transient association with amoebae has been reported for a number of different bacteria including Legionella pneumophila, many Mycobacterium spp, Francisella tubarensis, and Escherichia coli, among others[15]. Agriculture seems to play a crucial role for pathogens survival as they can live and replicate inside Dyctiostelium discoideum making husbandry. Nature’s report has made an important advance in the knowledge of amoebas behavior, and the famous Spanish phrase “ you are more stupid than a amoebae” is losing the sense because amoebas are an excellent example of social behavior with an amazing coordination and sense of sacrifice for the benefit of the species.
Current applications in research
Several D. discoideum genes are homologous to human genes making them a useful model organism. The entire genome was sequenced in a model organism database called dictyBase. Using these sequences scientists are now able to run more complex experiments especially on human-diseases. These slime molds are used to test anti-cancer drugs, immune-cell diseases, and bacterial intracellular pathogenesis. Also, recently it was established that Dictyostelium use DNA methylation for regulating gene expression.
Individual cell behavior accounts for many phases of health and disease. This is portrayed in D. discoideum in many different ways. Cytokinesis acts as part of immune response, tissue maintenance, and cancer, in the form of cell proliferation. Chemotaxis deals with inflammation, arthritis, asthma, lymphocyte trafficking, and axon guidance. Phagocytosis is used in immune surveillance and antigen presentation, while cell-type determination, cell sorting, and pattern formation are basic features of embryogenesis.
Legionella is a genus of bacteria that includes species that can cause Legionnaire's disease in humans. D. discoideum is also a host for Legionella and is a suitable model for studying the infection process.[17] Specifically, D. discoideum shares with mammalian host cells a similar cytoskeleton and cellular processes relevant to Legionella infection, including phagocytosis, membrane trafficking, endocytosis, vesicle sorting, and chemotaxis.
Much research has been done to better understand the behavioral responses of D. discoideum specifically, the slug form that has an array of behavioral traits. Although it lacks sensory cells and organs it has a keen response to external stimuli, such as temperature and light. Over 100,000 individual amoebae are attracted to one another through chemotaxis. However, light is directly related to the cAMP cell-cell signaling in chemotaxis. Within the D. discoideum the light causes cAMP to be released from the slug tip thus speeding up and directionalizing movement toward the light source. D. discoideum also prefers a slightly acidic pH. When intracellular pH is increased, it results in increased locomotion during chemotaxis. The movement of the slug is also increased because the movement of each individual cell has increased.
Classification and phylogeny
Dictyostelium have maintained more of their ancestral genome diversity than plants and animals, although proteome-based phylogeny confirms that amoebozoa diverged from the animal–fungal lineage after the plant–animal split.[18] The aggregation of individual amoebae into a multicellular fruiting body was an important factor that related the acrasids and dictyostelid cellular slime molds into a larger Class Acrasiomycetes.[19] Subclass, Dictyosteliidae, Order Dictyosteliales is a monophyletic assemblage within the Mycetozoa, a group that includes the protostelid, dictyostelid, and myxogastrid slime molds. Elongation factor-1α (EF-1α) data analyses support Mycetozoa as a monophyletic group even though rRNA trees place it as a polyphyletic group. Further, these data support the idea that the dictyostelid and myxogastrid are more closely related to each other than they are the protostelid. EF-1α analysis also placed the Mycetozoa as the immediate outgroup for the “animal-fungal clade.[20] Though not strongly supported, there is evidence that Mycetozoa are crown eukaryotes, more closely related to animals and fungi than are the green plants. The multicellularity of the dictyostelid evidences its relationship to animals. Comparing specific amino acid sequences of eight proteins from D. discoideum to those of their homologs in bacteria, yeast, and other eukaryotes indicates that Dictyostelium diverged from the line leading to mammals about the same time as the plant and animal divergence.[21] Therefore, the comparison of these eight proteins shows evidence that mammals are more closely related to Dictyostelium than to yeast. The experiment that was conducted by Loomis and Smith in 1990 suggested that yeast diverged much earlier.
Genome
The D. discoideum genome sequencing project was completed and published in 2005 by an international collaboration of institutes. This was the first free-living protozoan genome to be fully sequenced. D. discoideum consists of a 34Mb haploid genome with a base composition of 77% [A+T] and contains six chromosomes that encode approximately 12,500 proteins.[2] Sequencing of the D. discoideum genome provides a more intricate study of its cellular and developmental biology.
Tandem repeats of trinucleotides are very abundant in this genome; one class of the genome is clustered leading researchers to believe it serves as centromeres. The repeats correspond to repeated sequences of amino acids and are thought to be expanded by nucleotide expansion.[2] Expansion of trinucleotide repeats also occurs in humans, generally leading to many diseases. Learning how D. discoideum cells endure these amino acid repeats may provide insight to allow humans to tolerate them.
Every genome that is sequenced plays an important role in identifying genes that have been gained or lost over time. Comparative genomic studies allows for comparison of eukaryotic genomes. A phylogeny based on the proteome showed that the amoebozoa deviated from the animal-fungal lineage after the plant-animal split.[2] The D. discoideum genome is noteworthy because its many encoded proteins are commonly found in fungi, plants, and animals.[2]
Notes
- ^ Raper, K.B. (1935). "Dictyostelium discoideum, a new species of slime mold from decaying forest leaves". J. Agr. Res 50: 135–147. http://preserve.nal.usda.gov/jag/v50/v50i02/500135/a500135.htm
- ^ a b c d e Eichinger L; Noegel, AA (2003). "Crawling in to a new era – the Dictyostelium genome project". EMBO Journal 22 (9): 1941–1946. doi:10.1093/emboj/cdg214. PMC 156086. PMID 12727861. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=156086.
- ^ a b c Gilbert S.F. 2006. Developmental Biology. 8th ed. Sunderland (MA):Sinauer p. 36-39.
- ^ a b c d e f g h i j k l Tyler M.S. 2000. Developmental Biology: A guide for experimental study. 2nd ed. Sunderland (MA): Sinauer. p. 31-34. ISBN 0878938435
- ^ Dictybase, About Dictyostelium. [Online] (1, May, 2009). http://dictybase.org/
- ^ Dilip K. Nag, Disruption of Four Kinesin Genes in Dictyostelium. [Online] (22, April, 2008). http://ukpmc.ac.uk/articlerender.cgi?artid=1529371
- ^ Kay R.R., Garrod D., Tilly R. (1978). "Requirements for cell differentiation in Dictyostelium discoideum". Nature 211 (5640): 58–60. doi:10.1038/271058a0.
- ^ Dusenbery, David B. (1996). “Life at Small Scale”, pp.174-175. Scientific American Library, New York. ISBN 0-7167-5060-0.
- ^ Dusenbery, David B. (1996). “Life at Small Scale”, pp.108-109. Scientific American Library, New York. ISBN 0-7167-5060-0.
- ^ Chubb, JR; Trcek, T; Shenoy, SM; Singer, RH (2006). "Transcriptional pulsing of a developmental gene". Current biology : CB 16 (10): 1018–25. doi:10.1016/j.cub.2006.03.092. PMID 16713960.
- ^ Hudson J. J., Hsu D. W., Guo K., Zhukovskaya N., Liu P. H., Williams J. G., Pears C. J., Lakin N. D. (2005). "DNA-PKcs-dependent signaling of DNA damage in Dictyostelium discoideum". Curr Biol 15 (20): 1880–5. doi:10.1016/j.cub.2005.09.039. PMID 16243037.
- ^ a b "Amoebas show primitive farming behaviour as they travel", BBC News, 19 January 2011
- ^ a b c d Brock DA, Douglas TE, Queller DC, Strassmann JE (20 January 2011 2011). "Primitive agriculture in a social amoeba". Nature 469 (7330): 393–396. doi:10.1038/nature09668. PMID 21248849.
- ^ a b "Recent insights into host-pathogen interactions from Dyctiostelium". Cellular Microbiology 12 (3): 283–291. 2010.
- ^ a b Molmeret M., Horn, M., Wagner, M., Abu Kwaik, Y (January 2005). "Primitive Amoebae as Training Grounds for Intracellular Bacterial Pathogens". Appl Environ Microbiol: 20–28. doi:10.1128/AEM.71.1.20-28.2005.
- ^ a b Grant P.Ottom Mary Y.Wu,Margaret Clarke, Hao Lu, O.Roger Anderson, Hubert Hilbi, Howard A. Shuman, Richard H. Kessin (11 November 2003). Macroautophagy is dispensable for intracellular replication of Legionella pneumophila in Dictyostelium discoideum. doi:10.1046/j.1365-2958.2003.03826.x.
- ^ Bruhn H. 2008. Dictyostelium, a tractable model host organism for Legionella. In: Heuner K, Swanson M, editors. Legionella: Molecular Microbiology. Norwich (UK): Caister Academic Press. ISBN 978-1-904455-26-4
- ^ Eichenger L. et al. (2005). "The genome of the social amoeba Dictyostelium discoideum". Nature 435 (7038): 34–57. doi:10.1038/nature03481. PMC 1352341. PMID 15875012. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1352341.
- ^ Cavender J.C., Spiegl F., Swanson A. (2002). "Taxonomy, slime molds, and the questions we ask". The Mycological Society of America 96 (6): 968–979. PMID 21156570.
- ^ Baldauf S.L., Doolittle W.F. (1997). "Origin and evolution of the slime molds (Mycetozoa)". PNAS 94 (22): 12007–12012. doi:10.1073/pnas.94.22.12007. PMC 23686. PMID 9342353. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=23686.
- ^ Loomis W., Smith D. (1990). "Molecular phylogeny of Dictyostelium discoideum by protein sequence comparison". PNAS 87 (23): 9093–9098. doi:10.1073/pnas.87.23.9093. PMC 55110. PMID 2251251. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=55110.
References
- Tyler M.S. 2000. Developmental Biology A Guide for Experimental Study. 2nd ed. Sunderland (MA): Sinauer. pp. 31–34. ISBN 0878938435.
- Gilbert S. F. 2006. Developmental Biology. 8th ed. Sunderland (MA): Sinauer. pp. 36–39. ISBN 087893250X.
External links
Categories:- Developmental biology
- Model organisms
- Mycetozoa
Wikimedia Foundation. 2010.