- DDT (gene)
-
D-dopachrome tautomerase 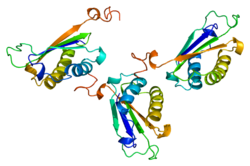
PDB rendering based on 1dpt.Available structures PDB 1dpt Identifiers Symbols DDT; DDCT; FLJ78842 External IDs OMIM: 602750 MGI: 1298381 HomoloGene: 1038 GeneCards: DDT Gene EC number 4.1.1.84 Gene Ontology Molecular function • dopachrome isomerase activity
• protein binding
• lyase activity
• D-dopachrome decarboxylase activityCellular component • cytoplasm Biological process • melanin biosynthetic process Sources: Amigo / QuickGO RNA expression pattern 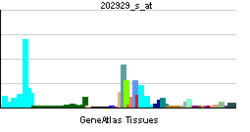
More reference expression data Orthologs Species Human Mouse Entrez 1652 13202 Ensembl ENSG00000099977 ENSMUSG00000001666 UniProt P30046 Q3UNI8 RefSeq (mRNA) NM_001084392.1 NM_010027.1 RefSeq (protein) NP_001077861.1 NP_034157.1 Location (UCSC) Chr 22:
24.31 – 24.32 MbChr 10:
75.23 – 75.24 MbPubMed search [1] [2] D-dopachrome decarboxylase is an enzyme that in humans is encoded by the DDT gene.[1][2][3]
D-dopachrome tautomerase converts D-dopachrome into 5,6-dihydroxyindole. The DDT gene is related to the macrophage migration inhibitory factor (MIF) in terms of sequence, enzyme activity, and gene structure. DDT and MIF are closely linked on chromosome 22.[3]
See also
- Macrophage migration inhibitory factor domain
References
- ^ Nishihira J, Fujinaga M, Kuriyama T, Suzuki M, Sugimoto H, Nakagawa A, Tanaka I, Sakai M (Mar 1998). "Molecular cloning of human D-dopachrome tautomerase cDNA: N-terminal proline is essential for enzyme activation". Biochem Biophys Res Commun 243 (2): 538–44. doi:10.1006/bbrc.1998.8123. PMID 9480844.
- ^ Coggan M, Whitbread L, Whittington A, Board P (Nov 1998). "Structure and organization of the human theta-class glutathione S-transferase and D-dopachrome tautomerase gene complex". Biochem J. 334 ( Pt 3): 617–23. PMC 1219731. PMID 9729470. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1219731.
- ^ a b "Entrez Gene: DDT D-dopachrome tautomerase". http://www.ncbi.nlm.nih.gov/sites/entrez?Db=gene&Cmd=ShowDetailView&TermToSearch=1652.
Further reading
- Hochstrasser DF, Frutiger S, Paquet N, et al. (1993). "Human liver protein map: a reference database established by microsequencing and gel comparison.". Electrophoresis 13 (12): 992–1001. doi:10.1002/elps.11501301201. PMID 1286669.
- Odh G, Hindemith A, Rosengren AM, et al. (1994). "Isolation of a new tautomerase monitored by the conversion of D-dopachrome to 5,6-dihydroxyindole.". Biochem. Biophys. Res. Commun. 197 (2): 619–24. doi:10.1006/bbrc.1993.2524. PMID 8267597.
- Esumi N, Budarf M, Ciccarelli L, et al. (1999). "Conserved gene structure and genomic linkage for D-dopachrome tautomerase (DDT) and MIF.". Mamm. Genome 9 (9): 753–7. doi:10.1007/s003359900858. PMID 9716662.
- Sugimoto H, Taniguchi M, Nakagawa A, et al. (1999). "Crystal structure of human D-dopachrome tautomerase, a homologue of macrophage migration inhibitory factor, at 1.54 A resolution.". Biochemistry 38 (11): 3268–79. doi:10.1021/bi982184o. PMID 10079069.
- Dunham I, Shimizu N, Roe BA, et al. (1999). "The DNA sequence of human chromosome 22.". Nature 402 (6761): 489–95. doi:10.1038/990031. PMID 10591208.
- Lubetsky JB, Dios A, Han J, et al. (2002). "The tautomerase active site of macrophage migration inhibitory factor is a potential target for discovery of novel anti-inflammatory agents.". J. Biol. Chem. 277 (28): 24976–82. doi:10.1074/jbc.M203220200. PMID 11997397.
- Strausberg RL, Feingold EA, Grouse LH, et al. (2003). "Generation and initial analysis of more than 15,000 full-length human and mouse cDNA sequences.". Proc. Natl. Acad. Sci. U.S.A. 99 (26): 16899–903. doi:10.1073/pnas.242603899. PMC 139241. PMID 12477932. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=139241.
- Gevaert K, Goethals M, Martens L, et al. (2004). "Exploring proteomes and analyzing protein processing by mass spectrometric identification of sorted N-terminal peptides.". Nat. Biotechnol. 21 (5): 566–9. doi:10.1038/nbt810. PMID 12665801.
- Sonesson B, Rosengren E, Hansson AS, Hansson C (2004). "UVB-induced inflammation gives increased d-dopachrome tautomerase activity in blister fluid which correlates with macrophage migration inhibitory factor.". Exp. Dermatol. 12 (3): 278–82. doi:10.1034/j.1600-0625.2003.120307.x. PMID 12823441.
- Collins JE, Wright CL, Edwards CA, et al. (2005). "A genome annotation-driven approach to cloning the human ORFeome.". Genome Biol. 5 (10): R84. doi:10.1186/gb-2004-5-10-r84. PMC 545604. PMID 15461802. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=545604.
- Gerhard DS, Wagner L, Feingold EA, et al. (2004). "The status, quality, and expansion of the NIH full-length cDNA project: the Mammalian Gene Collection (MGC).". Genome Res. 14 (10B): 2121–7. doi:10.1101/gr.2596504. PMC 528928. PMID 15489334. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=528928.
- Ewing RM, Chu P, Elisma F, et al. (2007). "Large-scale mapping of human protein-protein interactions by mass spectrometry.". Mol. Syst. Biol. 3 (1): 89. doi:10.1038/msb4100134. PMC 1847948. PMID 17353931. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1847948.
PDB gallery Categories:- Human proteins
- Chromosome 22 gene stubs
Wikimedia Foundation. 2010.

