- Cutinase
-
cutinase Identifiers EC number 3.1.1.74 Databases IntEnz IntEnz view BRENDA BRENDA entry ExPASy NiceZyme view KEGG KEGG entry MetaCyc metabolic pathway PRIAM profile PDB structures RCSB PDB PDBe PDBsum Gene Ontology AmiGO / EGO Search PMC articles PubMed articles Cutinase 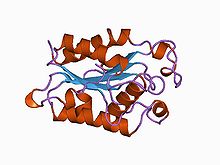
Structure of Fusarium solani cutinase.[1] Identifiers Symbol Cutinase Pfam PF01083 InterPro IPR000675 PROSITE PDOC00140 SCOP 1cex OPM family 135 OPM protein 1oxm Available protein structures: Pfam structures PDB RCSB PDB; PDBe PDBsum structure summary A cutinase (EC 3.1.1.74) is an enzyme that catalyzes the chemical reaction
- cutin + H2O
 cutin monomers
cutin monomers
Thus, the two substrates of this enzyme are cutin and H2O, whereas its product is cutin monomer.
This enzyme belongs to the family of hydrolases, specifically those acting on carboxylic ester bonds. The systematic name of this enzyme class is cutin hydrolase.
Aerial plant organs are protected by a cuticle composed of an insoluble polymeric structural compound, cutin, which is a polyester composed of hydroxy and hydroxyepoxy fatty acids[2]. Plant pathogenic fungi produce extracellular degradative enzymes[3] that play an important role in pathogenesis. They include cutinase, which hydrolyses cutin, facilitating fungus penetration through the cuticle. Inhibition of the enzyme can prevent fungal infection through intact cuticles. Cutin monomers released from the cuticle by small amounts of cutinase on fungal spore surfaces can greatly increase the amount of cutinase secreted by the spore, the mechanism for which process is as yet unknown[2][3].
Cutinase is a serine esterase containing the classical Ser, His, Asp triad of serine hydrolases[2]. The protein belongs to the alpha-beta class, with a central beta-sheet of 5 parallel strands covered by 5 helices on either side of the sheet. The active site cleft is partly covered by 2 thin bridges formed by amino acid side chains, by contrast with the hydrophobic lid possessed by other lipases[4]. The protein also contains 2 disulfide bridges, which are essential for activity, their cleavage resulting in complete loss of enzymatic activity[2]. Two cutinase-like proteins (MtCY39.35 and MtCY339.08c) have been found in the genome of the bacteria Mycobacterium tuberculosis.
References
- ^ Longhi S, Czjzek M, Lamzin V, Nicolas A, Cambillau C (May 1997). "Atomic resolution (1.0 A) crystal structure of Fusarium solani cutinase: stereochemical analysis". J. Mol. Biol. 268 (4): 779–99. doi:10.1006/jmbi.1997.1000. PMID 9175860.
- ^ a b c d Ettinger WF, Thukral SK, Kolattukudy PE (1987). "Structure of cutinase gene, cDNA, and the derived amino acid sequence from phytopathogenic fungi". Biochemistry 26 (24): 7883–7892. doi:10.1021/bi00398a052.
- ^ a b Sweigard JA, Chumley FG, Valent B (1992). "Cloning and analysis of CUT1, a cutinase gene from Magnaporthe grisea". Mol. Gen. Genet. 232 (2): 174–182. PMID 1557023.
- ^ Cambillau C, Martinez C, De Geus P, Lauwereys M, Matthyssens G (1992). "Fusarium solani cutinase is a lipolytic enzyme with a catalytic serine accessible to solvent". Nature 356 (6370): 615–618. doi:10.1038/356615a0. PMID 1560844.
- Garcia-Lepe R, Nuero OM, Reyes F, Santamaria F (1997). "Lipases in autolysed cultures of filamentous fungi". Lett. Appl. Microbiol. 25 (2): 127–30. doi:10.1046/j.1472-765X.1997.00187.x. PMID 9281862.
- Purdy RE, Kolattukudy PE (1975). "Hydrolysis of plant cuticle by plant pathogens. Purification, amino acid composition, and molecular weight of two isozymes of cutinase and a nonspecific esterase from Fusarium solani f. pisi". Biochemistry 14 (13): 2824–31. doi:10.1021/bi00684a006. PMID 1156575.
- Purdy RE, Kolattukudy PE (1975). "Hydrolysis of plant cuticle by plant pathogens. Properties of cutinase I, cutinase II, and a nonspecific esterase isolated from Fusarium solani pisi". Biochemistry 14 (13): 2832–40. doi:10.1021/bi00684a007. PMID 239740.
This article includes text from the public domain Pfam and InterPro IPR000675

This hydrolase article is a stub. You can help Wikipedia by expanding it. - cutin + H2O
