- DNA supercoil
-
DNA supercoiling refers to the over- or under-winding of a DNA strand, and is an expression of the strain on the polymer. Supercoiling is important in a number of biological processes, such as compacting DNA. Additionally, certain enzymes such as topoisomerases are able to change DNA topology to facilitate functions such as DNA replication or transcription. Mathematical expressions are used to describe supercoiling by comparing different coiled states to relaxed B-form DNA.
Contents
Role of supercoiling
In a "relaxed" double-helical segment of B-DNA, the two strands twist around the helical axis once every 10.4-10.5 base pairs of sequence. Adding or subtracting twists, as some enzymes can do, imposes strain. If a DNA segment under twist strain were closed into a circle by joining its two ends and then allowed to move freely, the circular DNA would contort into a new shape, such as a simple figure-eight. Such a contortion is a supercoil.
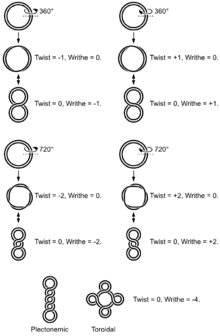
The simple figure eight is the simplest supercoil, and is the shape a circular DNA assumes to accommodate one too many or one too few helical twists. The two lobes of the figure eight will appear rotated either clockwise or counterclockwise with respect to one another, depending on whether the helix is over or underwound. For each additional helical twist being accommodated, the lobes will show one more rotation about their axis.
The noun form "supercoil" is rarely used in the context of DNA topology. Instead, global contortions of a circular DNA, such as the rotation of the figure-eight lobes above, are referred to as writhe. The above example illustrates that twist and writhe are interconvertible. "Supercoiling" is an abstract mathematical property representing the sum of twist and writhe. The twist is the number of helical turns in the DNA and the writhe is the number of times the double helix crosses over on itself (these are the supercoils).
Extra helical twists are positive and lead to positive supercoiling, while subtractive twisting causes negative supercoiling. Many topoisomerase enzymes sense supercoiling and either generate or dissipate it as they change DNA topology. DNA of most organisms is negatively supercoiled.
In part because chromosomes may be very large, segments in the middle may act as if their ends are anchored. As a result, they may be unable to distribute excess twist to the rest of the chromosome or to absorb twist to recover from underwinding—the segments may become supercoiled, in other words. In response to supercoiling, they will assume an amount of writhe, just as if their ends were joined.
Supercoiled DNA forms two structures; a plectoneme or a toroid, or a combination of both. A negatively supercoiled DNA molecule will produce either a one-start left-handed helix, the toroid, or a two-start right-handed helix with terminal loops, the plectoneme. Plectonemes are typically more common in nature, and this is the shape most bacterial plasmids will take. For larger molecules it is common for hybrid structures to form - a loop on a toroid can extend into a plectoneme. If all the loops on a toroid extend then it becomes a branch point in the plectonemic structure.
Occurrence of DNA supercoiling
DNA supercoiling is important for DNA packaging within all cells. Because the length of DNA can be thousands of times that of a cell, packaging this genetic material into the cell or nucleus (in eukaryotes) is a difficult feat. Supercoiling of DNA reduces the space and allows for much more DNA to be packaged. In prokaryotes, plectonemic supercoils are predominant, because of the circular chromosome and relatively small amount of genetic material. In eukaryotes, DNA supercoiling exists on many levels of both plectonemic and solenoidal supercoils, with the solenoidal supercoiling proving most effective in compacting the DNA. Solenoidal supercoiling is achieved with histones to form a 10 nm fiber. This fiber is further coiled into a 30 nm fiber, and further coiled upon itself numerous times more.
DNA packaging is greatly increased during nuclear division events such as mitosis or meiosis, where DNA must be compacted and segregated to daughter cells. Condensins and cohesins are Structural Maintenance of Chromosome proteins that aid in the condensation of sister chromatids and the linkage of the centromere in sister chromatids. These SMC proteins induce positive supercoils.
Supercoiling is also required for DNA/RNA synthesis. Because DNA must be unwound for DNA/RNA polymerase action, supercoils will result. The region ahead of the polymerase complex will be unwound; this stress is compensated with positive supercoils ahead of the complex. Behind the complex, DNA is rewound and there will be compensatory negative supercoils. It is important to note that topoisomerases such as DNA gyrase (Type II Topoisomerase) play a role in relieving some of the stress during DNA/RNA synthesis.[1]
Modeling using mathematics
DNA supercoiling can be described numerically by changes in the 'linking number' Lk. The linking number is the most descriptive property of supercoiled DNA. Lko, the number of turns in the relaxed (B type) DNA plasmid/molecule, is determined by dividing the total base pairs of the molecule by the relaxed bp/turn which, depending on reference is 10.4-10.5.
- Lko = bp / 10.4
Lk is merely the number of crosses a single strand makes across the other in a planar projection. The topology of the DNA is described by the equation below in which the linking number is equivalent to the sum of TW, which is the number of twists or turns of the double helix, and Wr which is the number of coils or 'writhes'. If there is a closed DNA molecule, the sum of TW and Wr, or the linking number, does not change. However, there may be complementary changes in TW and Wr without changing their sum.
- Lk = Tw + Wr
The change in the linking number, ΔLk, is the actual number of turns in the plasmid/molecule, Lk, minus the number of turns in the relaxed plasmid/molecule Lko.
- ΔLk = Lk − Lko
If the DNA is negatively supercoiled ΔLk < 0. The negative supercoiling implies that the DNA is underwound.
A standard expression independent of the molecule size is the "specific linking difference" or "superhelical density" denoted σ. σ represents the number of turns added or removed relative to the total number of turns in the relaxed molecule/plasmid, indicating the level of supercoiling.
- σ = ΔLk / Lko
The Gibbs free energy associated with the coiling is given by the equation below[2]
- ΔG / N = 10RTσ2
Examples
Since the linking number L of supercoiled DNA is the number of times the two strands are intertwined (and both strands remain covalently intact), L cannot change. The reference state (or parameter) L0 of a circular DNA duplex is its relaxed state. In this state, its writhe W = 0. Since L = T + W, in a relaxed state T = L. Thus, if we have a 400 bp relaxed circular DNA duplex, L ~ 40 (assuming ~10 bp per turn in B-DNA). Then T ~ 40.
- Positively supercoiling:
- T = 0, W = 0, then L = 0
- T = +3, W = 0, then L = +3
- T = +2, W = +1, then L = +3
- Negatively supercoiling:
- T = 0, W = 0, then L = 0
- T = -3, W = 0, then L = -3
- T = -2, W = -1, then L = -3
Negative supercoils favor local unwinding of the DNA, allowing processes such as transcription, DNA replication, and recombination. Negative supercoiling is also thought to favour the transition between B-DNA and Z-DNA, and moderate the interactions of DNA binding proteins involved in gene regulation.[3]
See also
- Mechanical properties of DNA
- Ribbon theory
References
- ^ Albert A-C, Spirito F, Figueroa-Bossi N, Bossi L, Rahmouni AR (1996). "Hyper-negative template DNA supercoiling during transcription of the tetracycline-resistance gene in topA mutants is largely constrained in vivo". Nucl Acids Res 24 (15): 3093–3099. doi:10.1093/nar/24.15.3093. PMC 146055. PMID 8760899. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=146055.
- ^ Vologodskii AV, Lukashin AV, Anshelevich VV et al. (1979). "Fluctuations in superhelical DNA". Nucleic Acids Res 6 (3): 967–682. doi:10.1093/nar/6.3.967. PMC 327745. PMID 155809. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=327745.
- ^ H. S. Chawla (2002). Introduction to Plant Biotechnology. Science Publishers. ISBN 1578082285.
Categories:- DNA
- Helices
- Molecular biology
Wikimedia Foundation. 2010.